Transferase (HAT/HMT) MS Activity Assays
Label-free mass spectrometry-based activity assays for histone acetyltransferase and histone methyltransferase inhibitor screening.
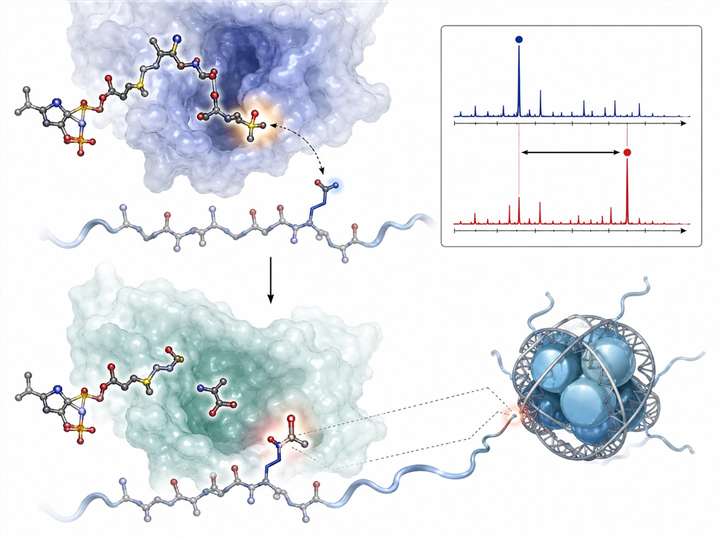
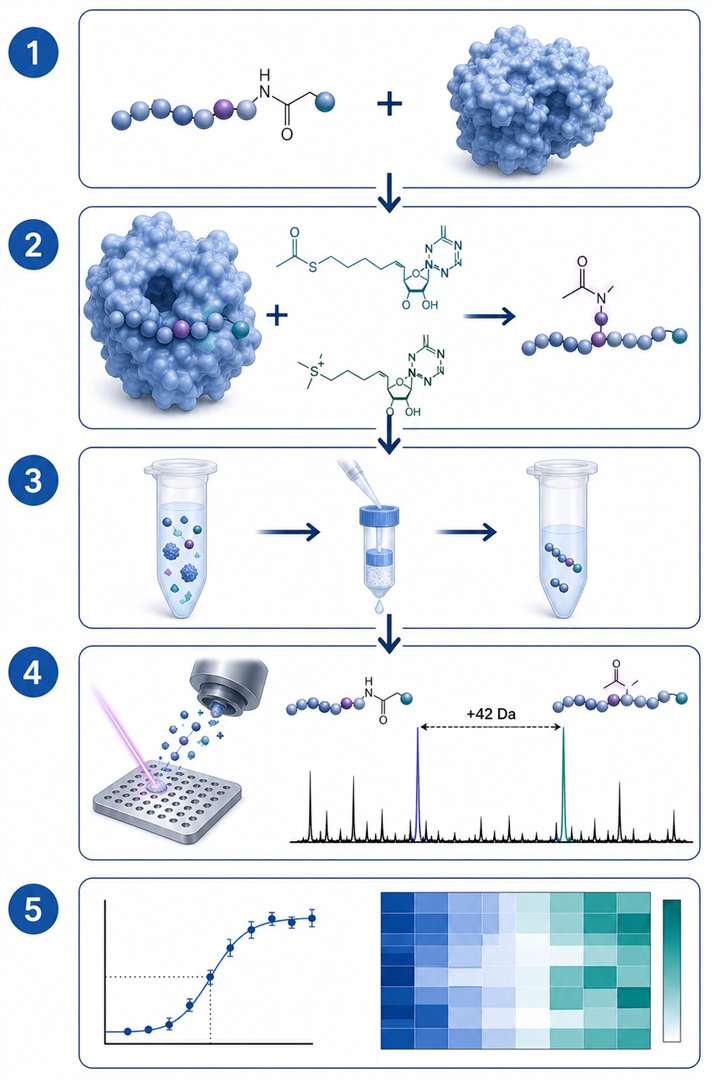
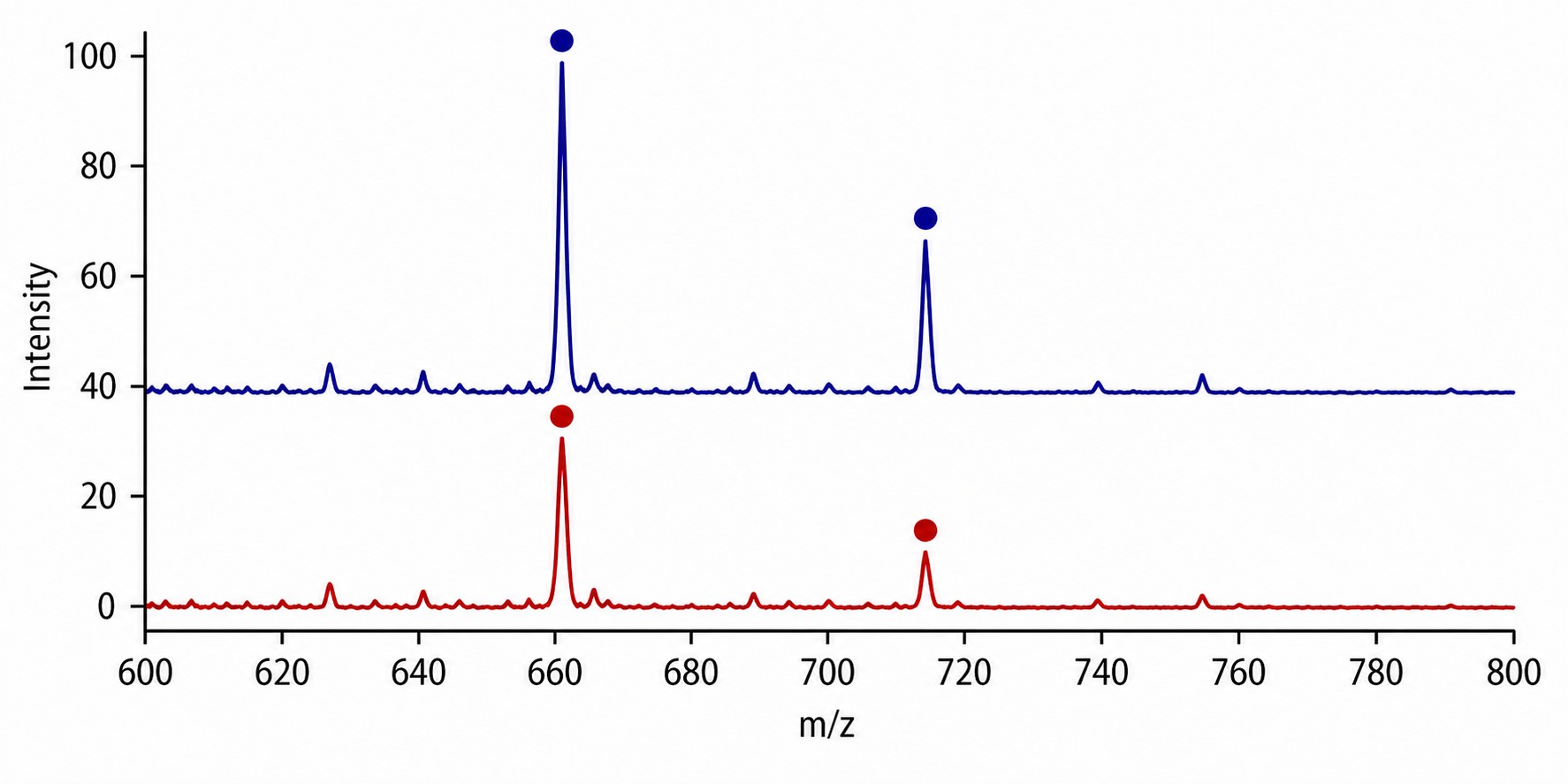
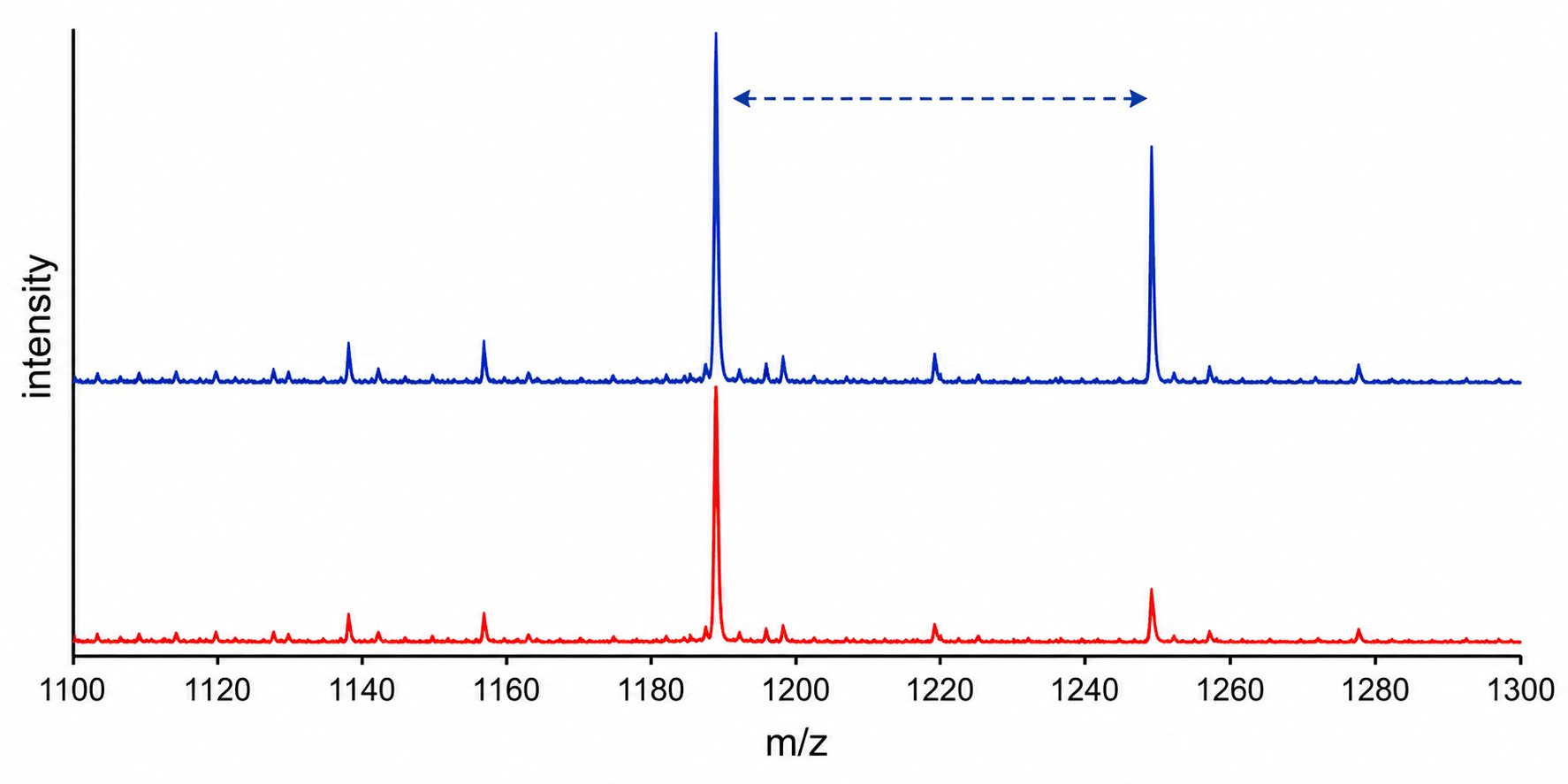
Transferase MS activity assays use label-free mass spectrometry to directly measure enzyme-catalysed group transfer reactions — acetylation by histone acetyltransferases (HATs) and methylation by histone methyltransferases (HMTs). Unlike fluorescence- or antibody-based methods, MS detects the mass shift of the substrate peptide directly (+42 Da for acetylation, +14/28/42 Da for mono-/di-/trimethylation), giving you an unambiguous, quantitative readout of enzyme activity without interference from compound autofluorescence or antibody cross-reactivity.
At Creative Proteomics, our Transferase MS service is designed to address the challenges of epigenetic drug discovery: compound interference in fluorescence-based assays, the need for site-specific resolution, and the demand for a single assay platform that works across both HAT and HMT targets.
Leveraging our MALDI-TOF and LC-MS platforms, we deliver high-throughput, label-free activity data for small-molecule inhibitors, fragments, and tool compounds targeting HATs and HMTs.
Key Advantages:
- Direct mass readout eliminates compound autofluorescence and quenching interference.
- Single assay format works for both HATs (p300/CBP, PCAF, GCN5, MYST) and HMTs (G9a, EZH2, DOT1L, SETD7, PRMT family).
- Site-specific resolution — identifies exactly which lysine residue is modified on multi-site substrates.
- MALDI-TOF HTS format with <5 s acquisition per sample in 384-well plates.
- Compatible with peptide and full-length recombinant nucleosome substrates.