Tissue Slice (PCLS) MS Drug Response
Comprehensive LC-MS drug response profiling in precision-cut tissue slices for translational ADME and toxicity research.
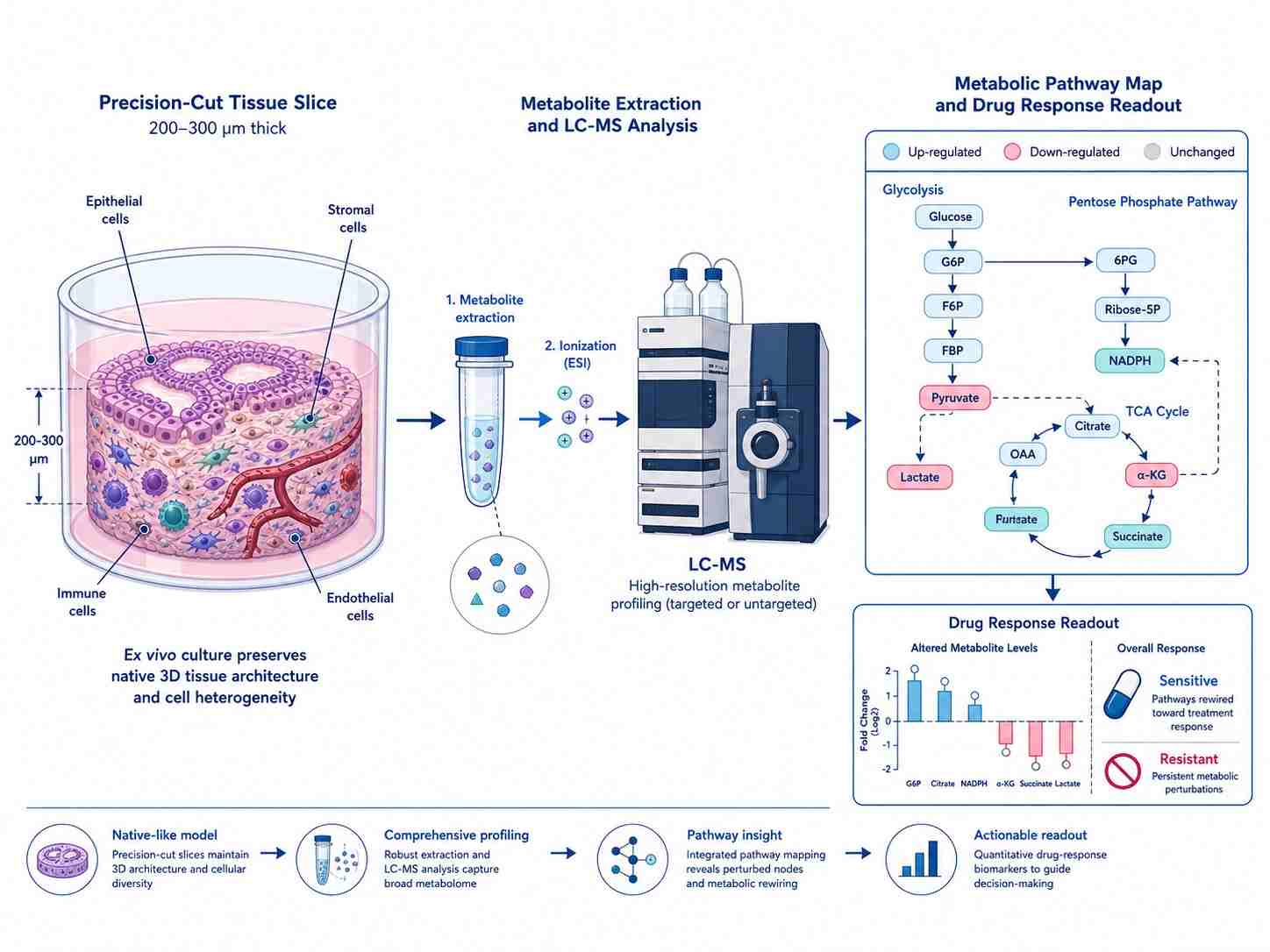
Precision-cut tissue slices (PCTS) — including precision-cut lung slices (PCLS), precision-cut liver slices, and precision-cut intestinal slices (PCIS) — represent the gold-standard ex vivo model for drug metabolism and toxicity research. By combining precision-cut tissue slice technology with LC-MS metabolomics, lipidomics, and targeted drug quantification, our Tissue Slice (PCLS) MS Drug Response service delivers a comprehensive readout of drug-tissue interactions in a physiologically relevant, human-relevant platform.
Unlike 2D cell cultures that rapidly lose CYP enzyme activity or animal models that suffer from species-specific metabolic differences, precision-cut tissue slices retain the native tissue architecture — including all resident cell types (epithelial, stromal, immune, endothelial), intact extracellular matrix, and metabolic zonation — making them the most translationally predictive ex vivo model for ADME, drug-induced toxicity, and pharmacometabolomics studies.
Key Capabilities:
- Multi-organ PCLS capability: lung (healthy and diseased), liver (healthy, fibrotic, steatotic), intestine (small and colon), and tumor tissue.
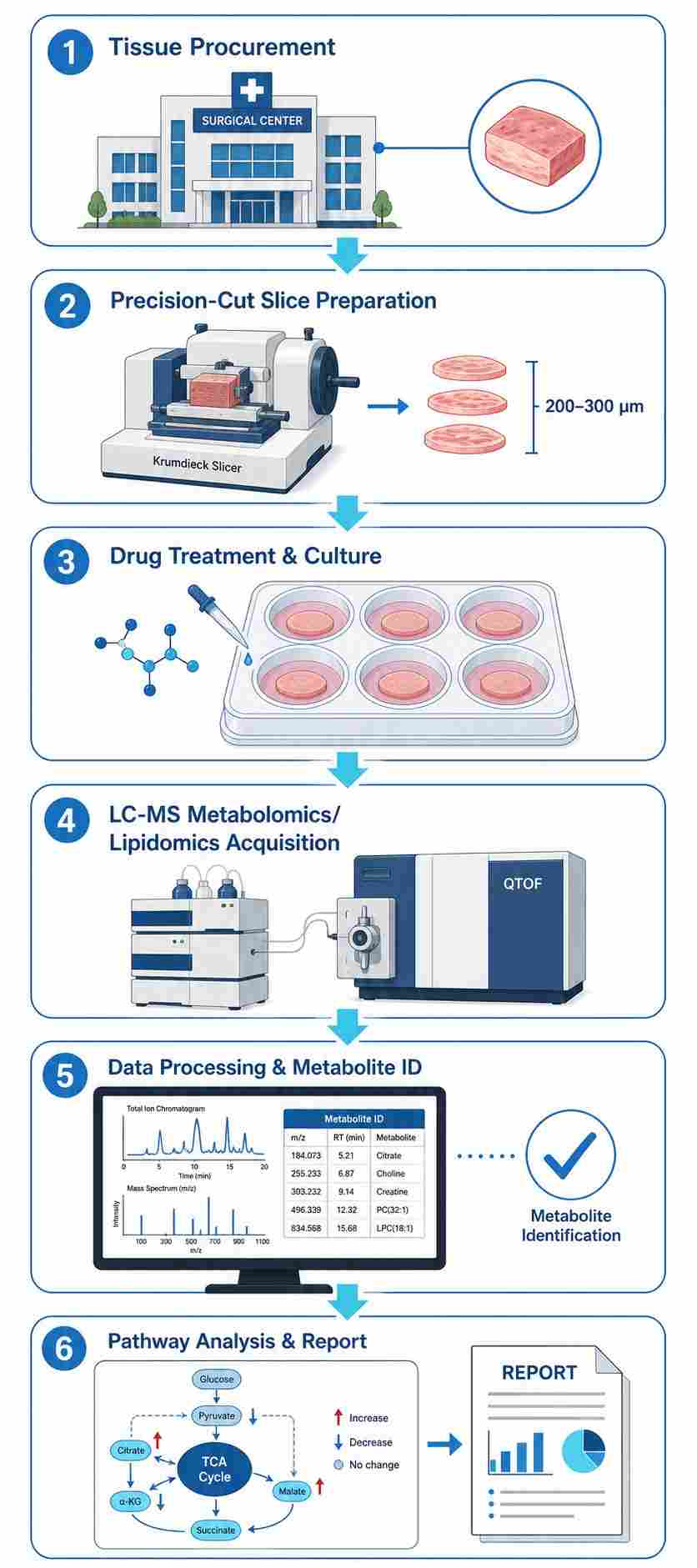
- Integrated LC-QTOF-MS untargeted metabolomics/lipidomics and LC-TQ-MS targeted drug quantification from the same tissue slices.
- Validated slice preparation protocol (Krumdieck slicer, 200-300 µm) with >80% viability post-cutting.
- Flexible service model: full-service (tissue-to-report), slice-only, or analysis-only options.
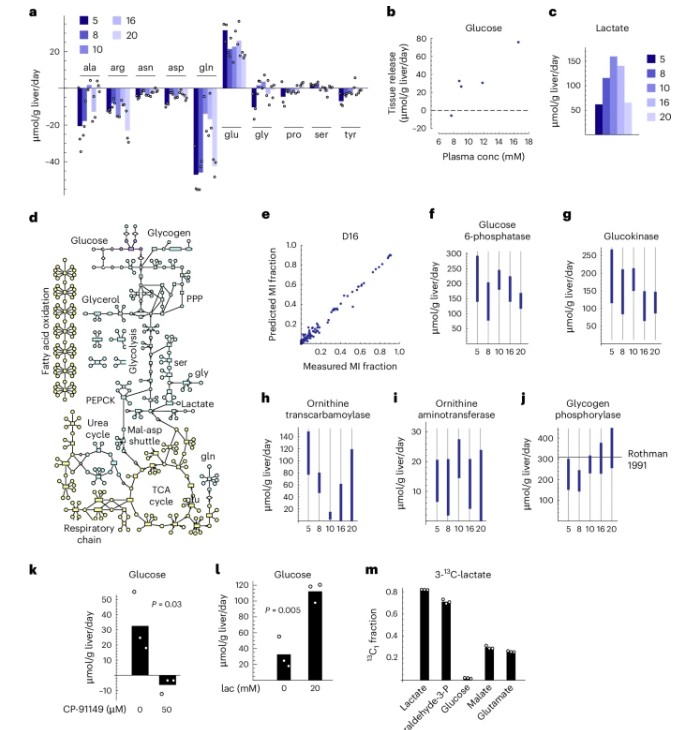
- Stable isotope tracing (¹³C) for metabolic flux analysis in drug-treated tissue slices.
- Comprehensive pathway enrichment analysis with KEGG, Reactome, and HMDB mapping.