Enzyme Kinetics via Continuous-Flow MS
Real-time, label-free kinetic profiling — native substrates, microgram-scale enzyme, full parameter determination in hours, not days.
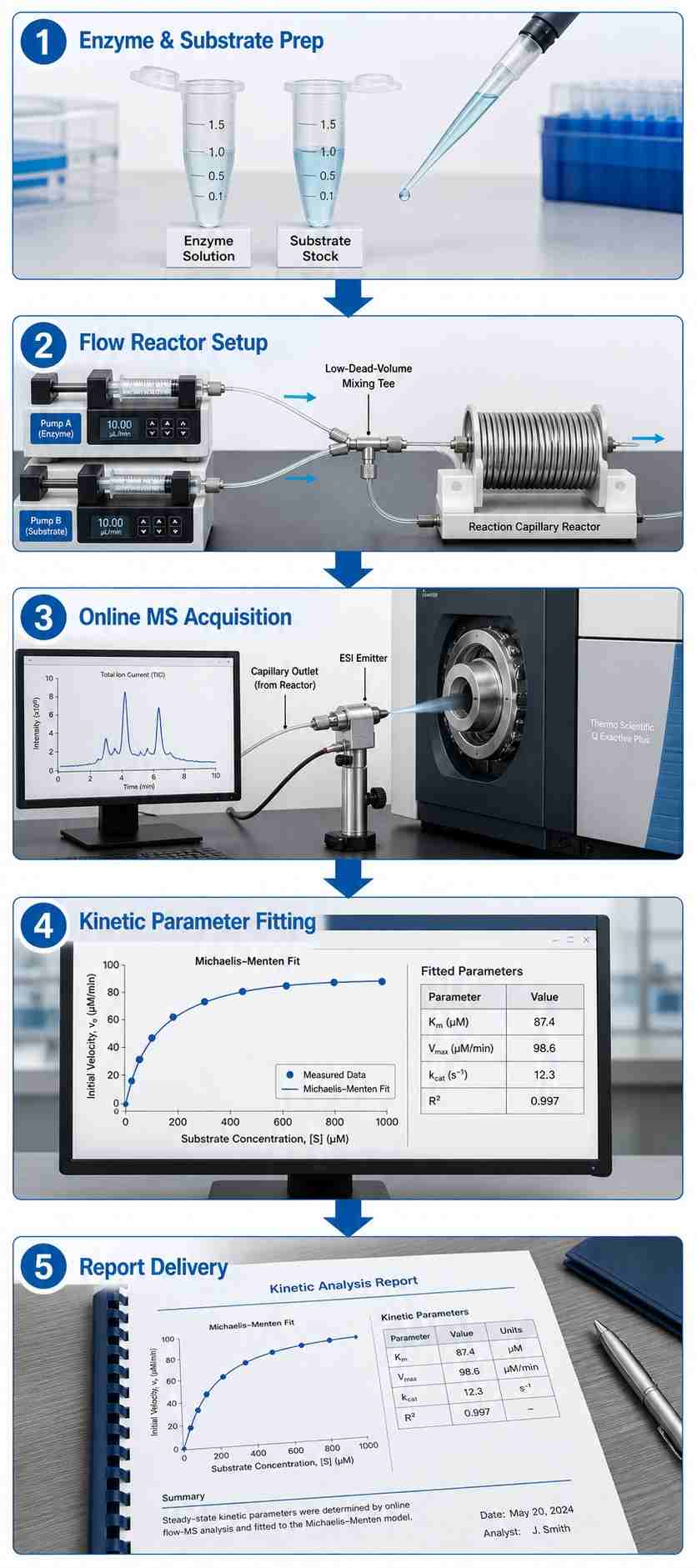
Continuous-flow mass spectrometry (CF-MS) kinetics directly couples a microfluidic flow reactor to a high-resolution mass spectrometer, capturing enzyme progress curves in continuous real time from native substrates. Unlike plate-reader fluorescence assays, there is no labeling requirement. Unlike discrete LC-MS, there is no fraction collection. The result: a direct, continuous readout of substrate consumption and product formation from which Km, Vmax, kcat, IC50, and Ki are determined in a single experiment.
At Creative Proteomics, our MassTarget™ CF-MS Kinetics platform integrates precision syringe pumps, temperature-controlled flow reactors, and high-resolution QToF mass spectrometry into a standardized service workflow covering kinases, P450s, proteases, transferases, hydrolases, and other enzyme classes.
Core Capabilities:
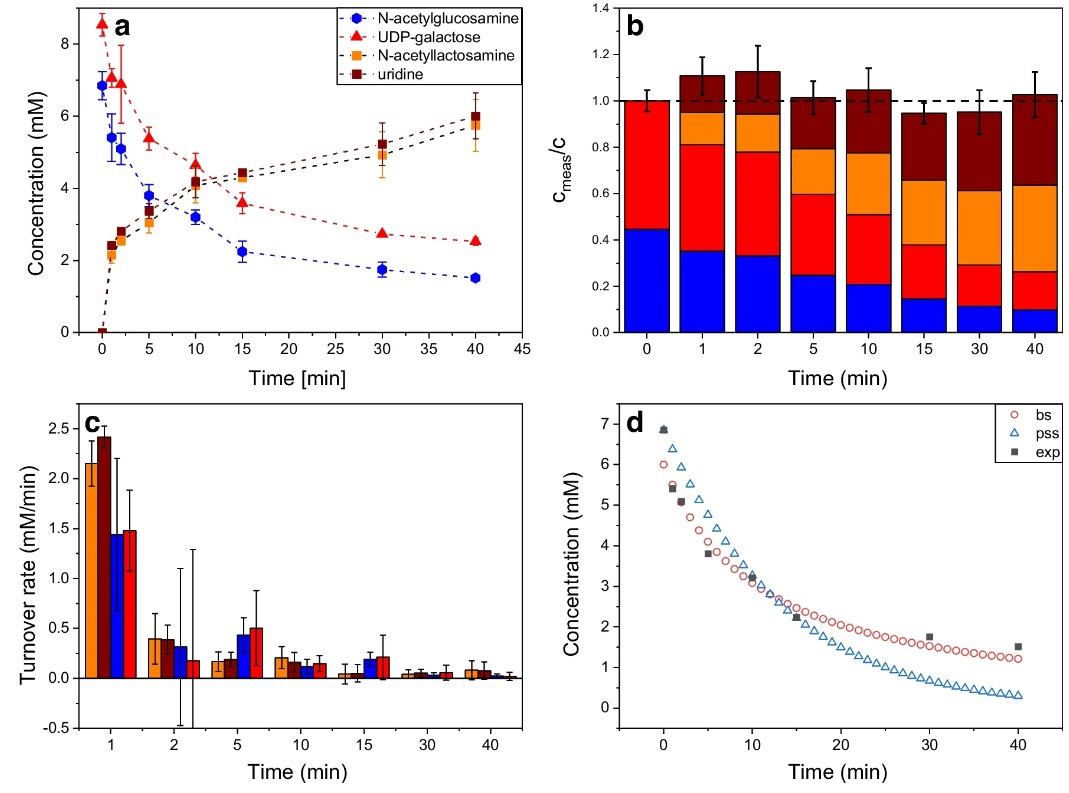
- Real-time progress curves — continuous ion intensity traces, not end-point snapshots
- Native substrates — any ionizable substrate, no fluorophore or chromophore required
- Microgram-scale consumption — 0.1–10 µg per Km curve
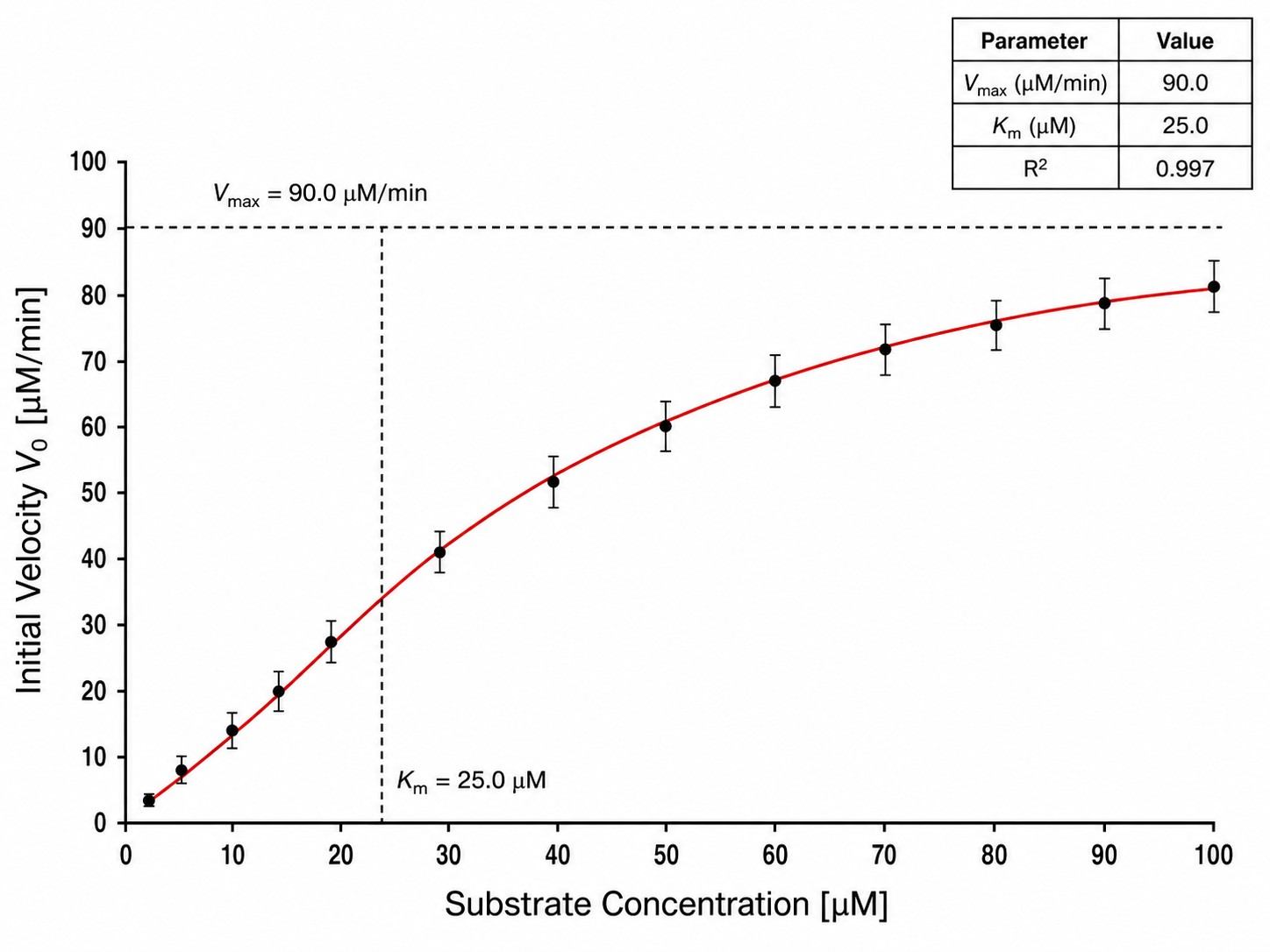
- Full kinetic parameters — Km, Vmax, kcat, IC50, Ki (competitive/non-competitive/mixed)
- Rapid deployment — no assay development if substrate/product are MS-detectable