Cellular Metabolomics Screening
Label-free, comprehensive profiling of drug-induced metabolic perturbations in living cells — untargeted discovery and targeted absolute quantification by mass spectrometry.
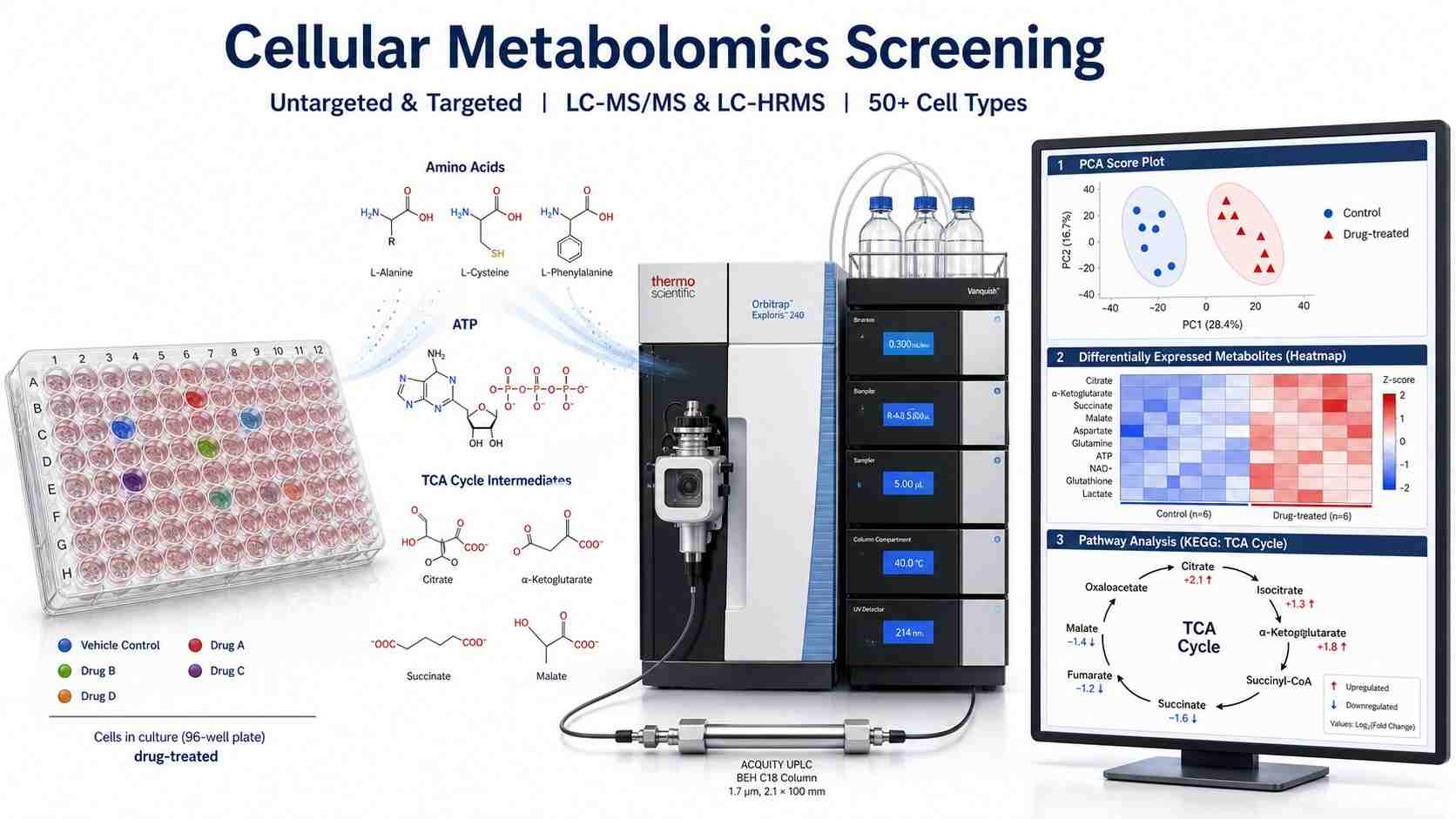
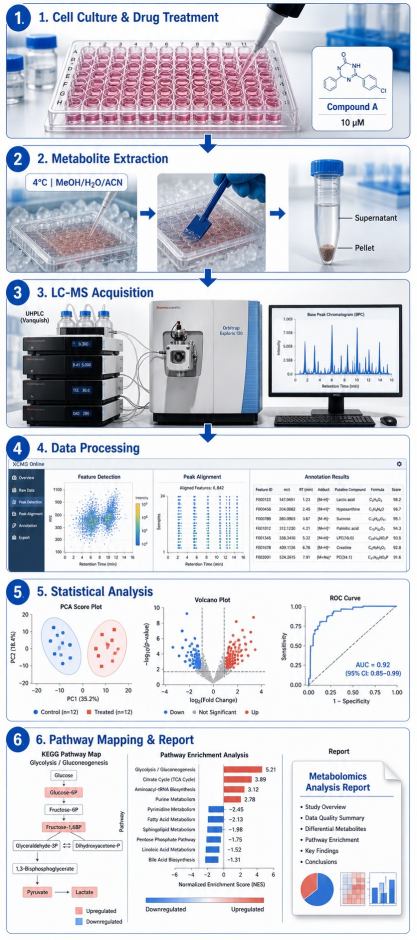
Cellular metabolomics screening by mass spectrometry captures the global metabolic state of living cells in response to drug treatment. Unlike endpoint biochemical assays that measure a single enzyme activity or reporter signal, cellular metabolomics simultaneously quantifies hundreds to thousands of intracellular metabolites — amino acids, organic acids, nucleotides, sugars, central carbon cycle intermediates, and signaling lipids — providing a systems-level view of drug-induced metabolic perturbations.
This approach is increasingly adopted in early-stage drug discovery for mechanism-of-action (MOA) elucidation, on-target vs off-target metabolic effect profiling, metabolic biomarker discovery, and drug resistance characterization. By measuring the actual metabolic endpoints of drug activity in a physiologically relevant cell context, cellular metabolomics bridges the gap between biochemical target engagement data and phenotypic outcomes.
Key Capabilities:
- Untargeted and targeted metabolomics workflows on a single platform
- 50+ validated cell types including adherent, suspension, primary, spheroids, and organoids
- Absolute quantification with stable isotope-labeled internal standards
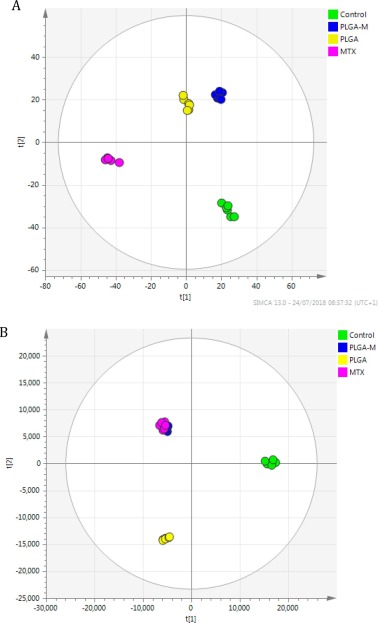
- PCA/PLS-DA/OPLS-DA multivariate analysis + KEGG pathway mapping
- Dedicated project management from cell culture to final report