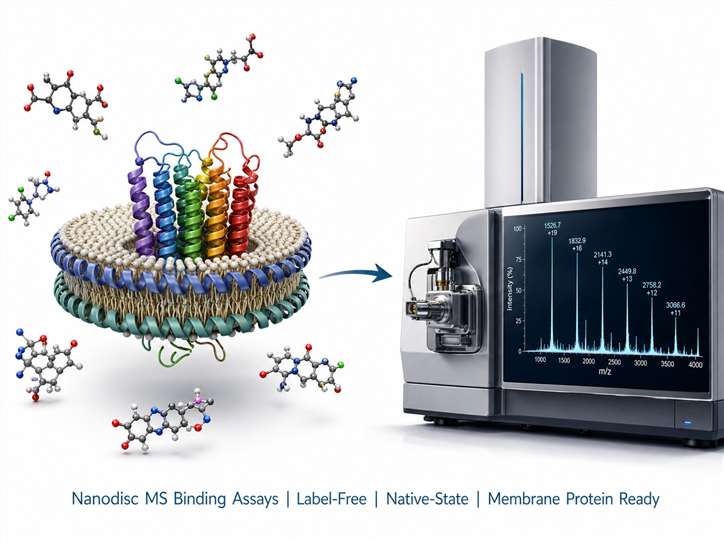
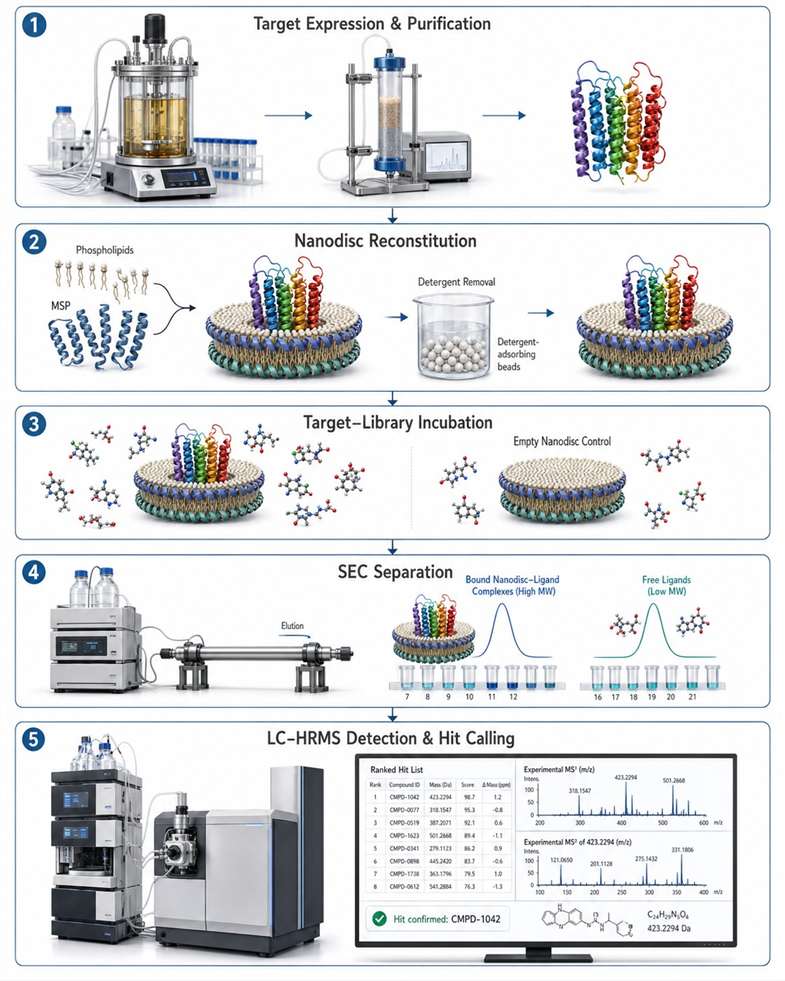
Nanodisc Reconstitution & MS Binding Assays for Membrane Protein Drug Discovery
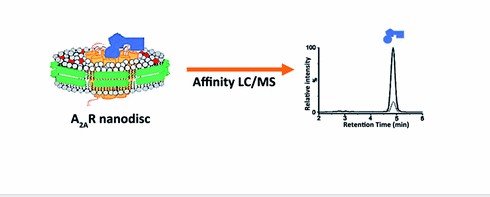
We combine nanodisc reconstitution with affinity selection mass spectrometry (ASMS) to deliver label-free, native-state binding data for membrane protein targets — GPCRs, ion channels, and transporters — without detergent denaturation or target immobilisation.
Key Advantages:
- Native lipid bilayer environment — Nanodiscs keep membrane proteins in a near-physiological state, avoiding the denaturation risks that come with detergent micelles.
- Label-free MS detection — Direct binding readout. No fluorescent tags, no immobilisation, no reporter assays.
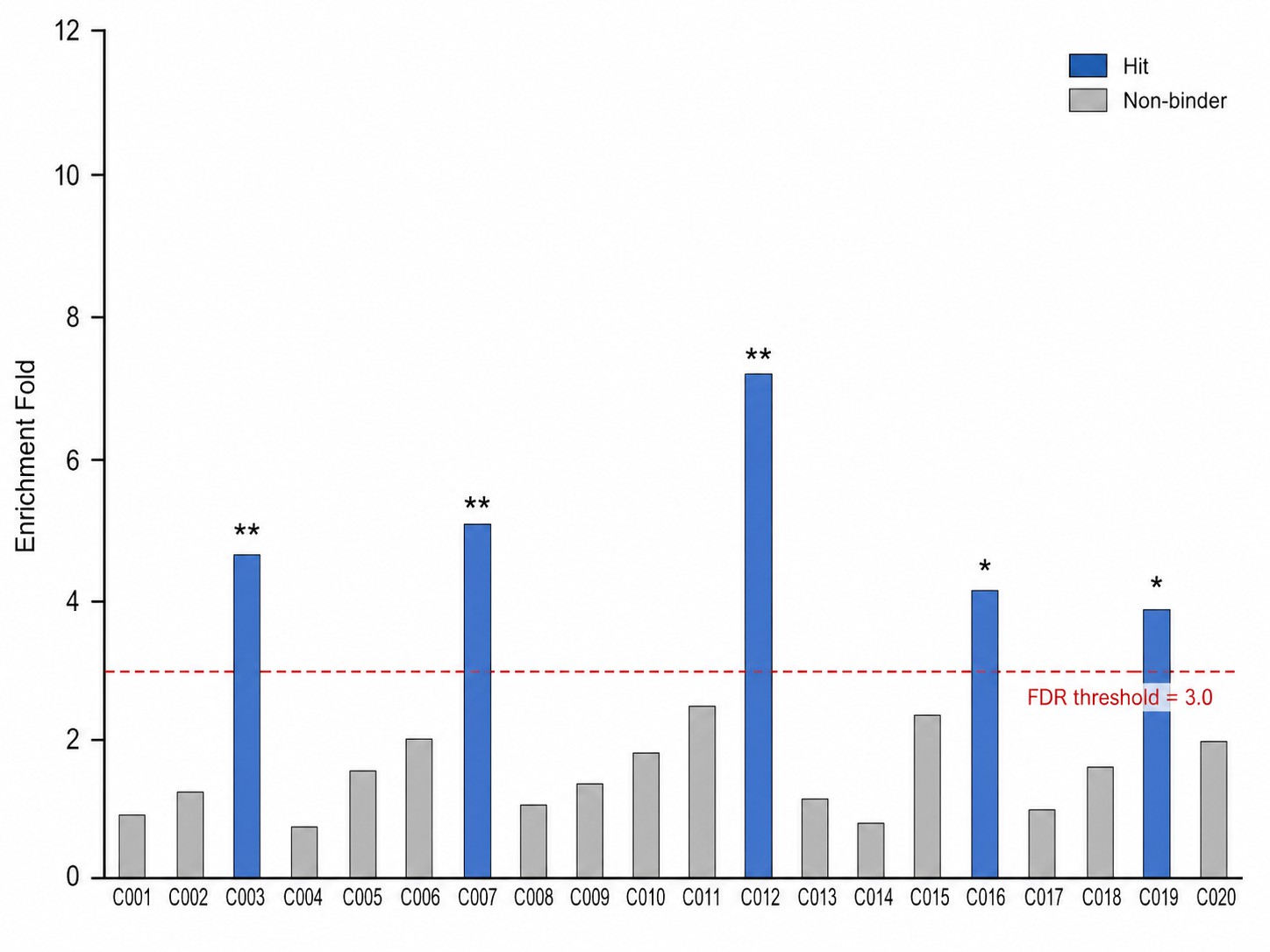
- Sensitive to weak and transient interactions — Our high-resolution MS captures low-affinity binding events that fragment-based and early-stage discovery campaigns depend on.
- Pooled library throughput — Screen up to 1,000 compounds per run in a single ASMS experiment, cutting assay development time and consumable costs.