Background
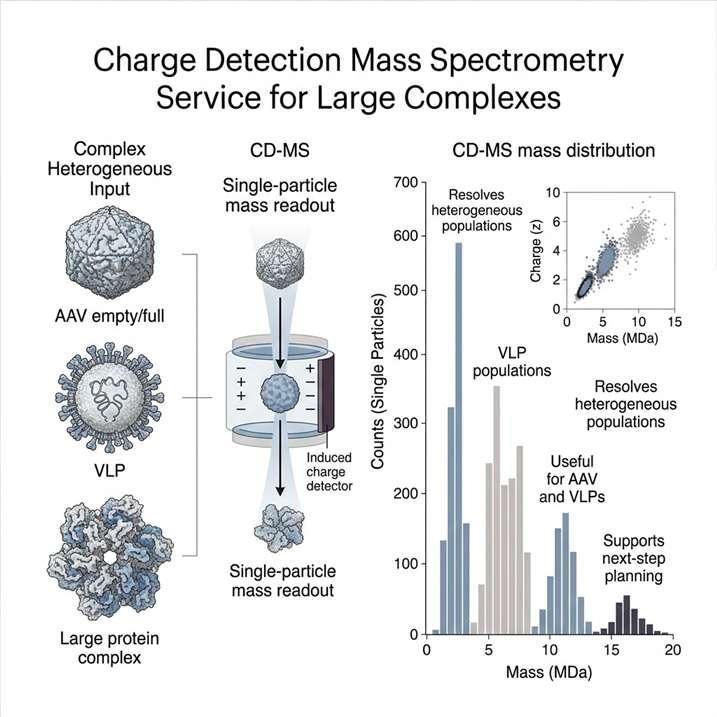
AAV manufacturing and purification workflows generate samples that can contain multiple particle populations, including target capsids, aggregates, and non-target material. For development-stage characterization, the key question is often not simply whether AAV is present, but whether the particle population is clean, reproducible, and analytically distinguishable across runs and conditions.
Methods
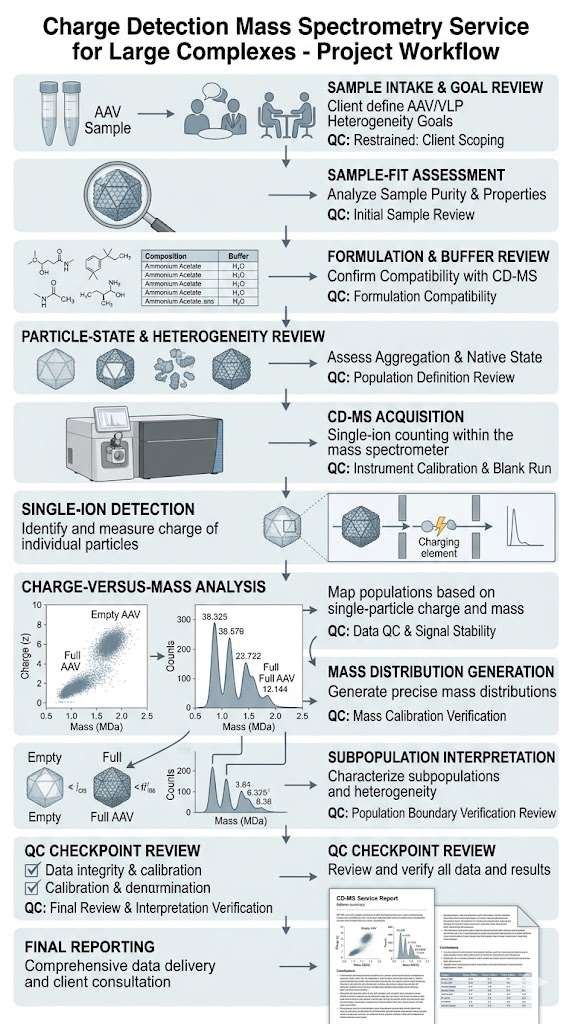
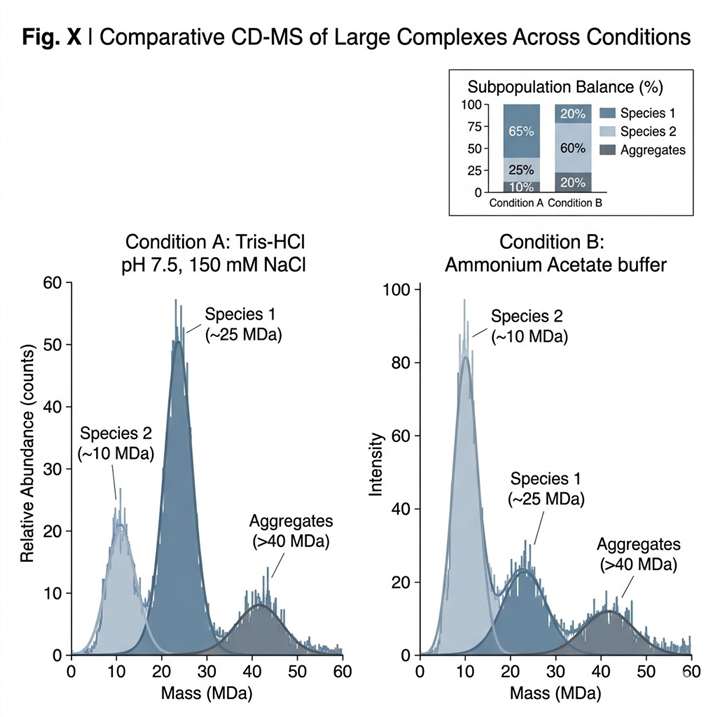
In the study, purified AAV9-1 material was analyzed by CD-MS in triplicate. The workflow included mass histograms and charge-versus-mass scatterplots so that individual ions could be evaluated across the characteristic charging regions observed in native electrospray. This approach allowed the authors to assess reproducibility and to distinguish typical AAV particle signals from lower-mass material, aggregates, and other non-target populations.
Results
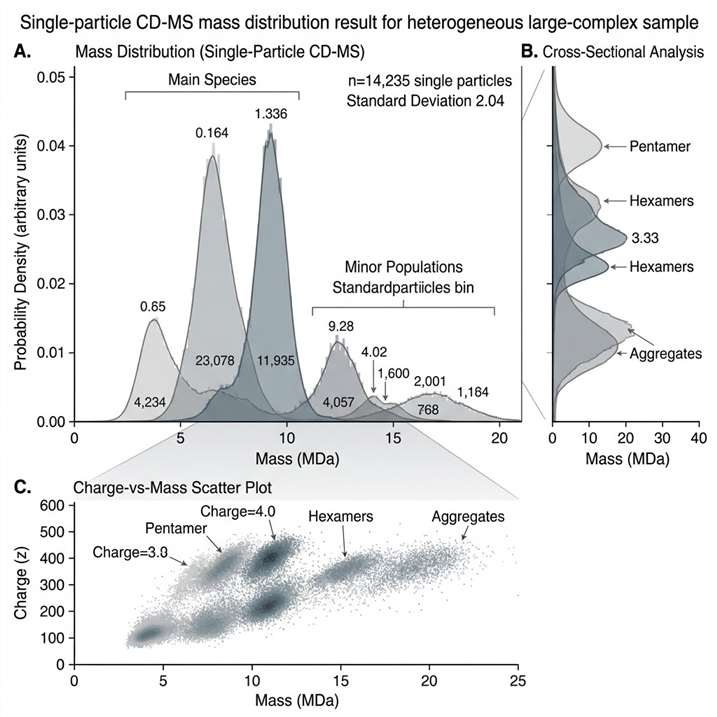
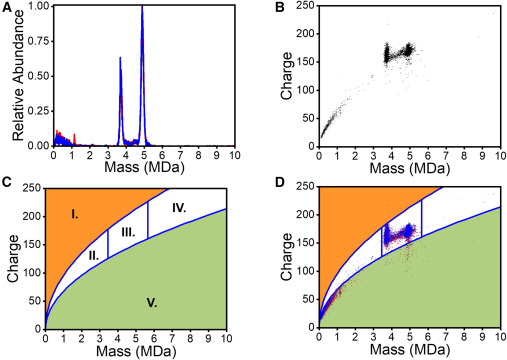
In Figure 1, the case is presented through four linked views. Figure 1A shows triplicate mass histograms for purified AAV9-1, demonstrating reproducible distribution patterns across replicate measurements. Figure 1B shows a charge-versus-mass scatterplot in which each dot represents a single AAV ion from one run. Figure 1C defines the typical charging regions observed in native electrospray, separating DNA and denatured proteins, low-molecular-weight species, AAV particles, AAV aggregates, and surfactant micelles or high-density aggregates. Figure 1D overlays triplicate charge-versus-mass scatterplots and shows that typical AAV ions fall within the defined blue-line boundaries. Together, these views show that CD-MS can provide both reproducible mass distributions and clear contextual separation between target AAV populations and non-target signal regions.
Conclusion
This figure is highly relevant for a CD-MS service page because it shows exactly the kind of result clients need to interpret: reproducibility across runs, direct single-ion distribution readout, and visually defined boundaries between intended particle populations and non-target material. For large-complex and viral-particle characterization projects, that kind of output supports sample assessment, subpopulation review, and more confident decisions about what to do next.
Source for verification: End-to-end characterization of AAV manufacturing process using charge detection mass spectrometry