Enzyme-Ligand Residence Time MS
Kinetics-driven drug discovery — directly measure how long your drug stays bound, not just how tightly.
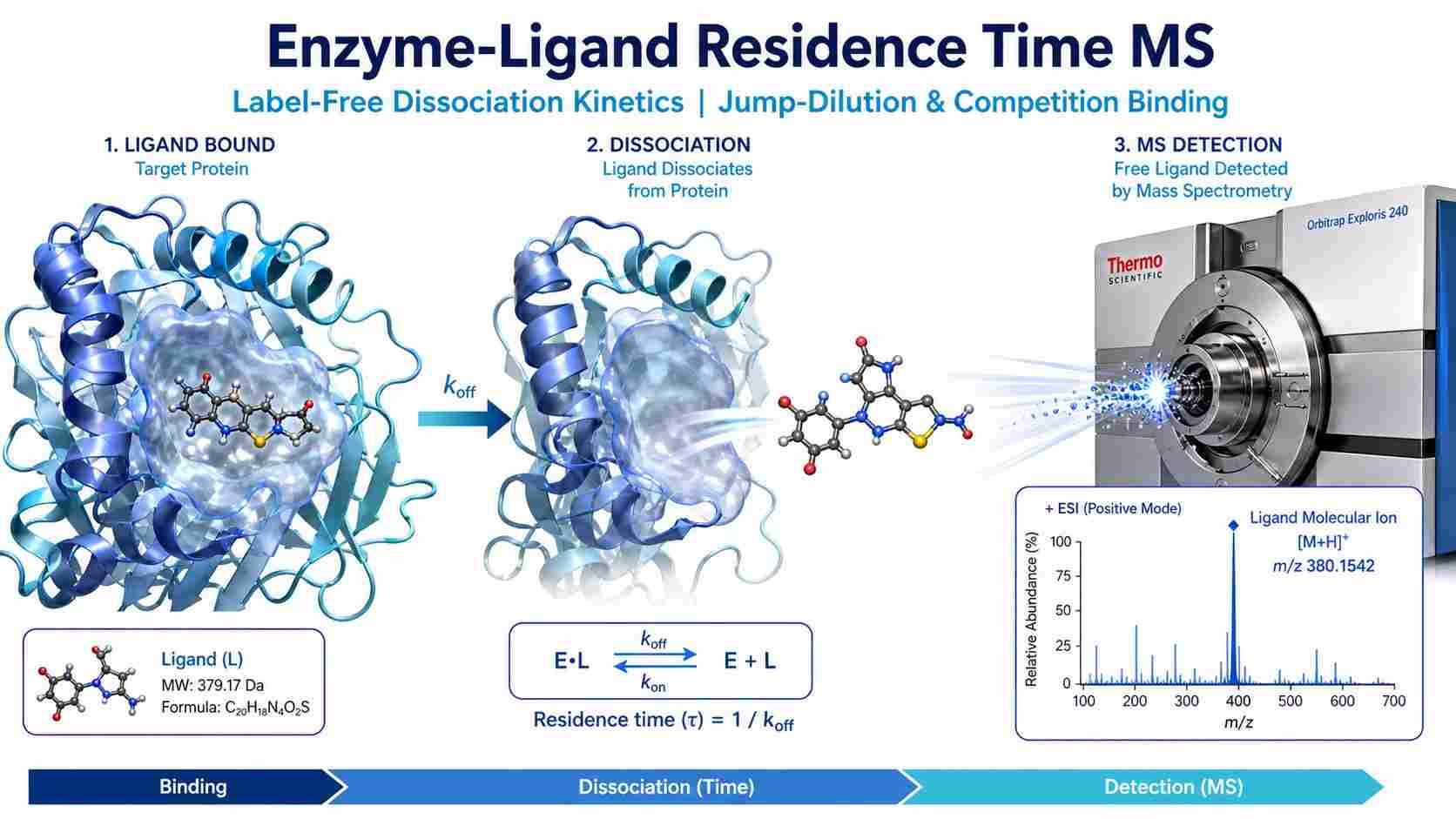
Drug-target residence time — the duration a ligand remains bound to its biological target — is increasingly recognized as a better predictor of in vivo efficacy than equilibrium binding affinity (KD) alone. Two compounds can have identical KD values yet exhibit dramatically different dissociation rates, leading to vastly different pharmacodynamic profiles.
At Creative Proteomics, our MassTarget™ Residence Time MS service uses label-free mass spectrometry to directly monitor dissociation kinetics of enzyme-ligand complexes under native conditions. By combining our RapidFire MS, SEC-MS, and pulsed ultrafiltration platforms with validated jump-dilution and competition binding protocols, we provide robust koff and t½ measurements without radioactive labels, fluorophores, or target immobilization.
Core Capabilities:
- Label-free dissociation kinetics — direct MS readout of free ligand reappearance
- No target immobilization — native-state binding, no surface artifacts
- Broad kinetic range — t½ from seconds to hours
- Multiple assay formats — jump-dilution MS, competition binding MS, pulsed ultrafiltration
- Low sample consumption — micrograms of target protein per determination