MS-Based Apoptosis Markers Analysis
Targeted mass spectrometry for simultaneous, absolute quantification of key apoptosis-related proteins — from caspase activation to PARP cleavage — in a single LC-MS/MS run.
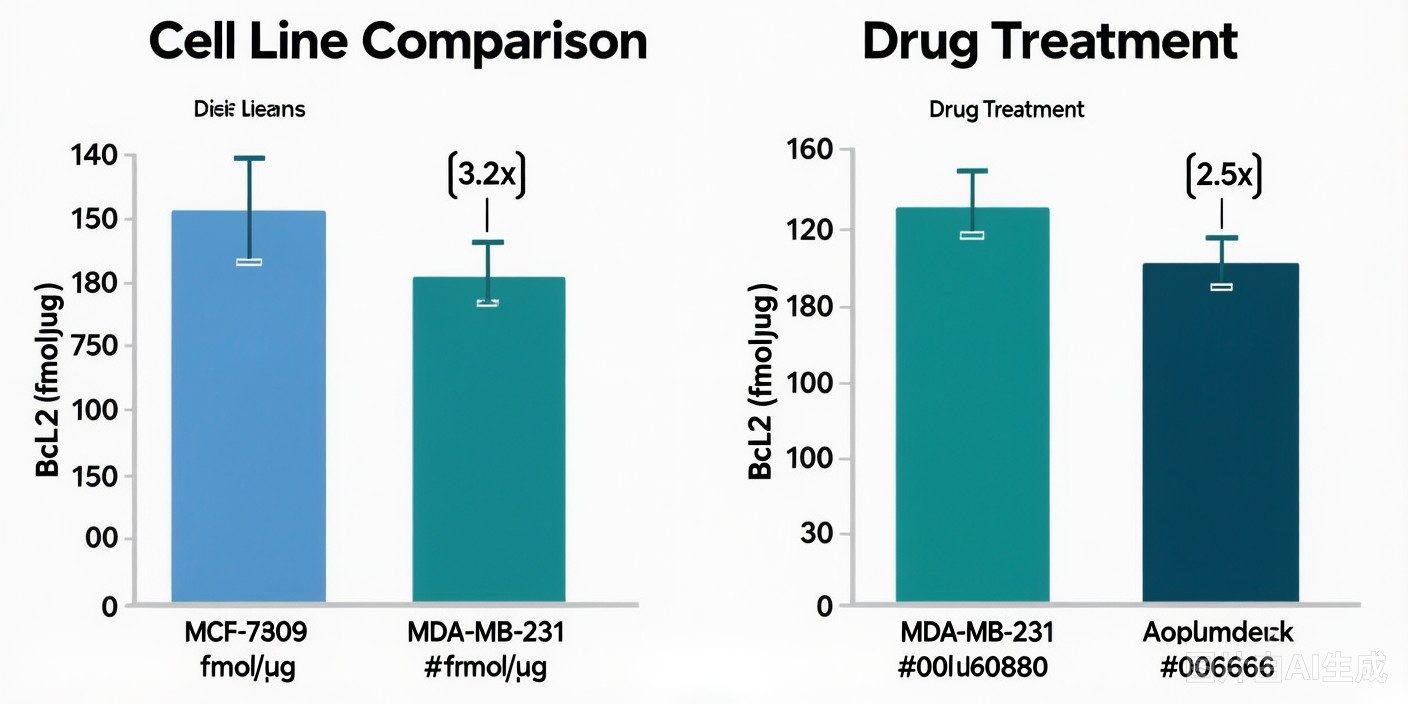
MS-based apoptosis markers analysis is a targeted mass spectrometry approach that enables simultaneous, absolute quantification of key apoptosis-related proteins — including caspases, cleaved PARP, cytochrome c, and Bcl-2 family members — from a single sample injection, providing comprehensive apoptosis pathway profiling with superior specificity and dynamic range compared to conventional antibody-based methods.
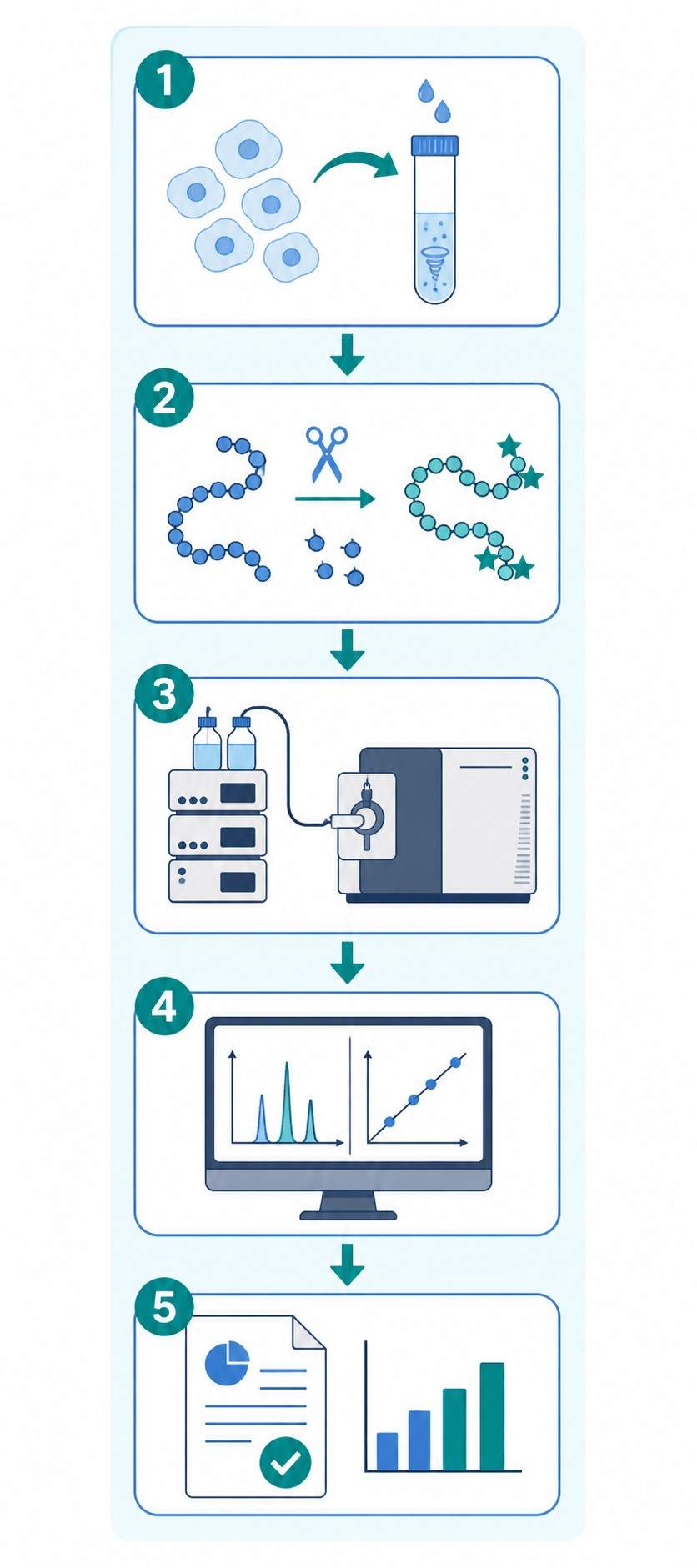
At Creative Proteomics, our MassTarget™ platform provides a fully integrated MS-based apoptosis marker analysis service, from method development through sample processing, data acquisition, and bioinformatics reporting, supporting preclinical drug discovery programs across oncology, neurology, and immuno-oncology.
Key Advantages:
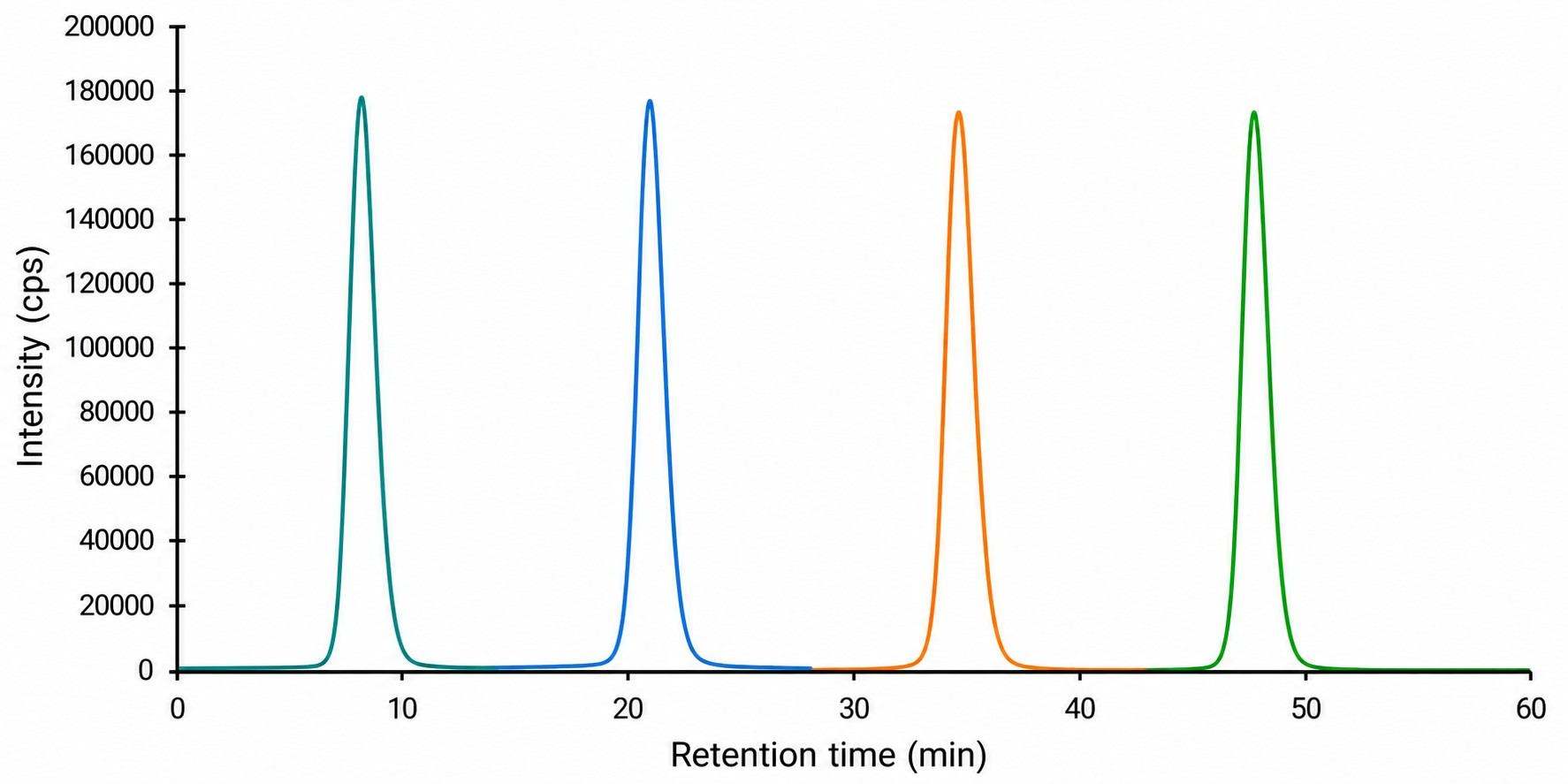
- Multiplex quantification of 10+ apoptosis markers in a single LC-MS/MS run.
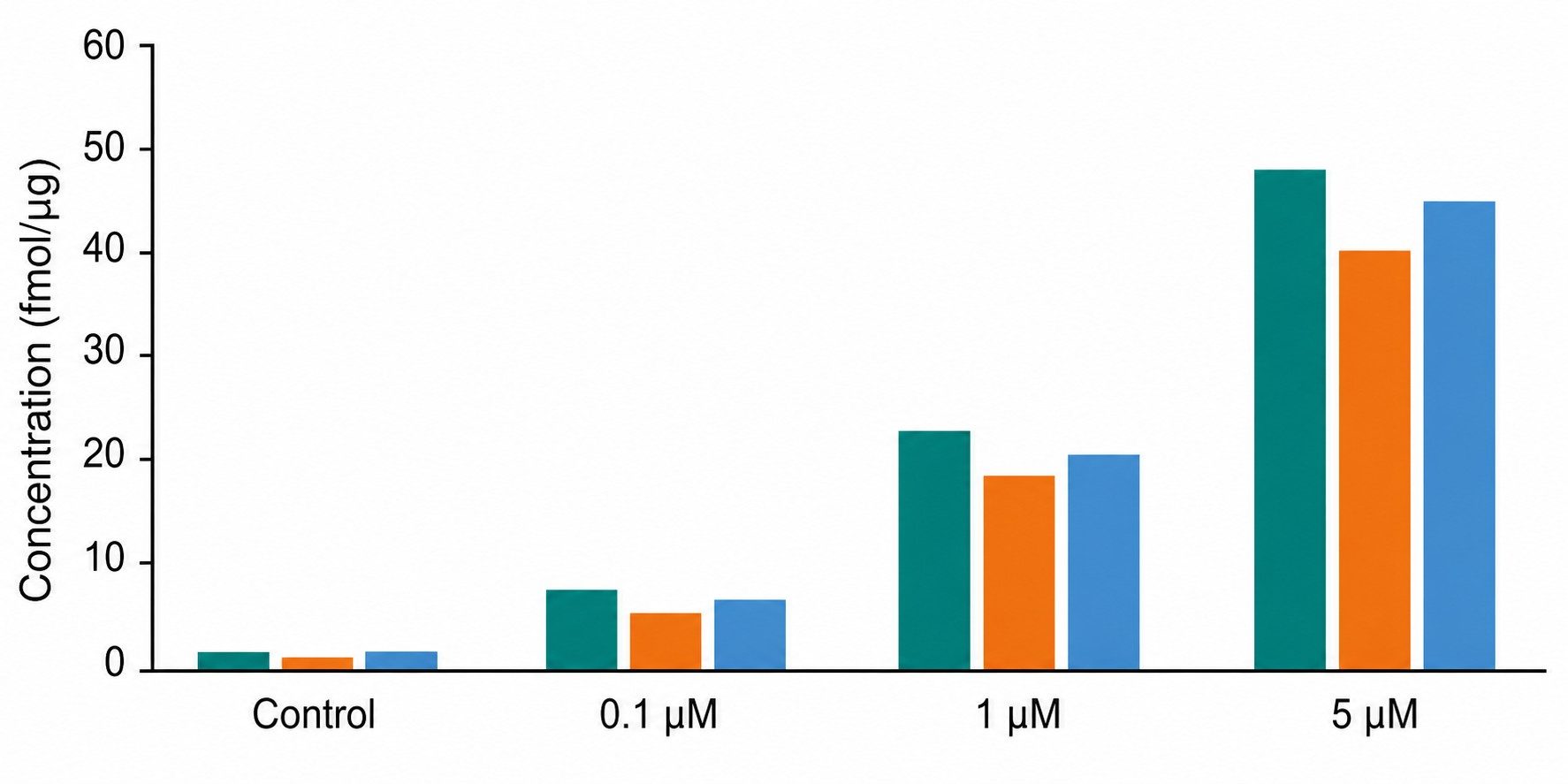
- Absolute quantification with stable isotope-labeled internal standards.
- Isoform-specific detection of cleaved vs. full-length caspases and PARP.
- 50-100× wider dynamic range than Western blot.
- Requires 5-10× less sample than equivalent ELISA panel.
- Custom panel design for your specific apoptosis markers of interest.