High-Throughput IM-MS and CCS Profiling for Compound Screening
We deliver high-throughput ion mobility-mass spectrometry and collision cross-section profiling — adding an orthogonal dimension of structural information to your compound screening workflow for isomer resolution, metabolite identification, and CCS database building.
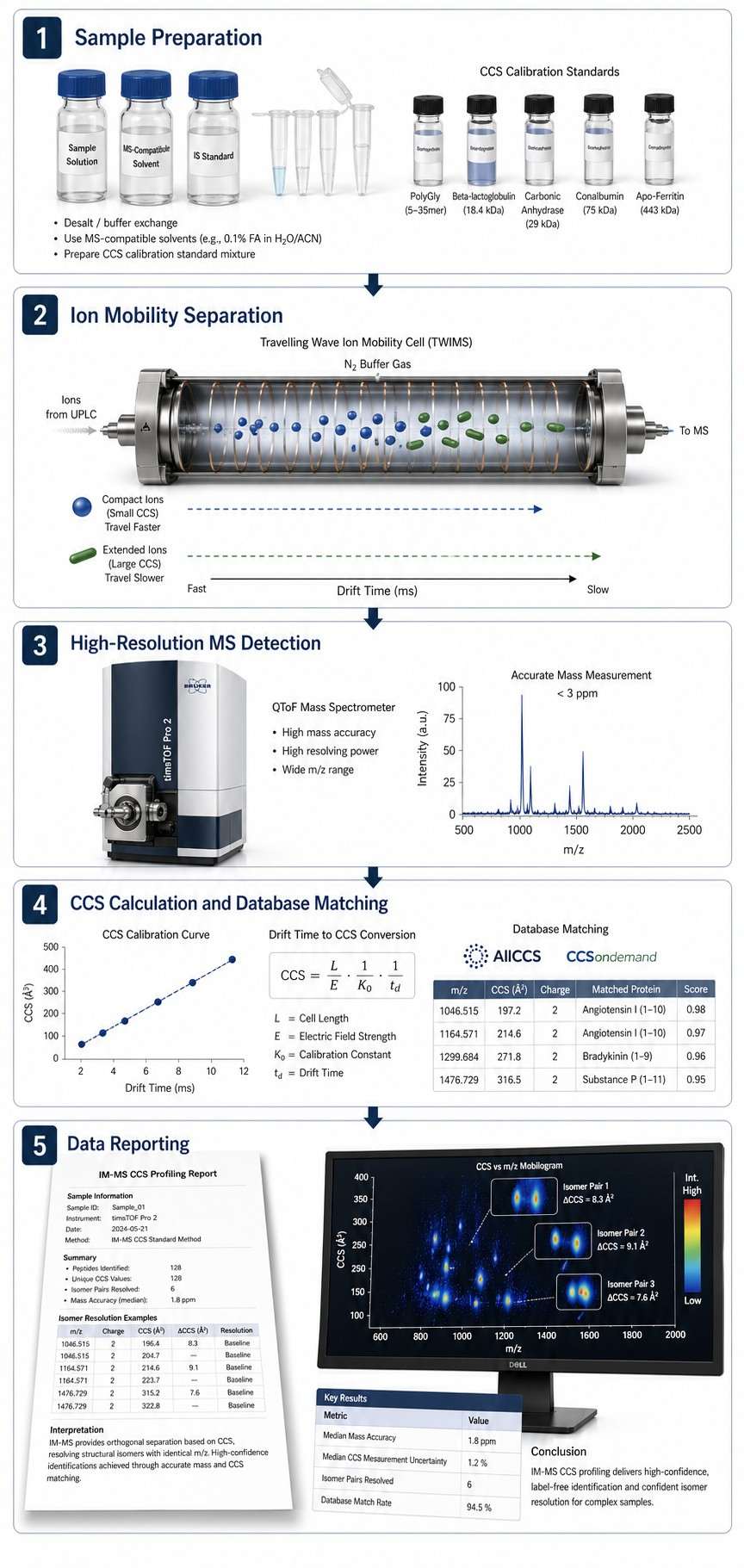
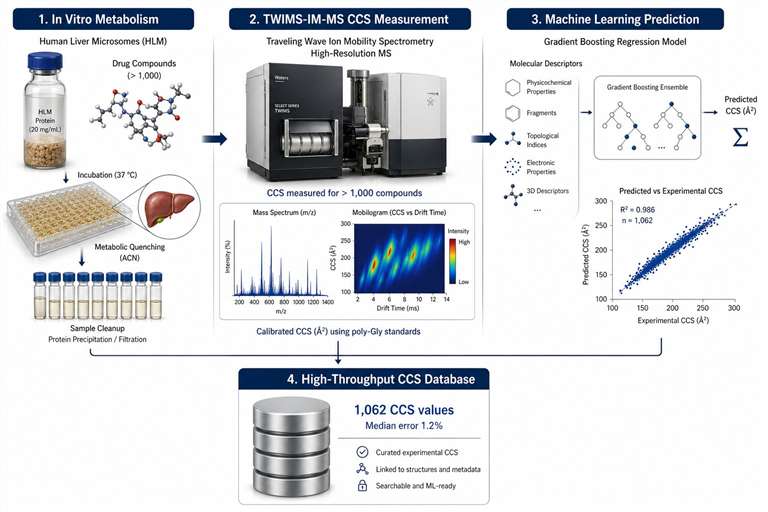
Ion mobility-mass spectrometry (IM-MS) adds a dimension that standard mass spectrometry cannot reach — ion shape and size. In an IM-MS experiment, ions are separated in the gas phase by how easily they move through a buffer gas. The resulting collision cross section (CCS) value tells you how large and what shape an ion is in the gas phase, giving you a physicochemical fingerprint that works alongside m/z and retention time for compound identification.
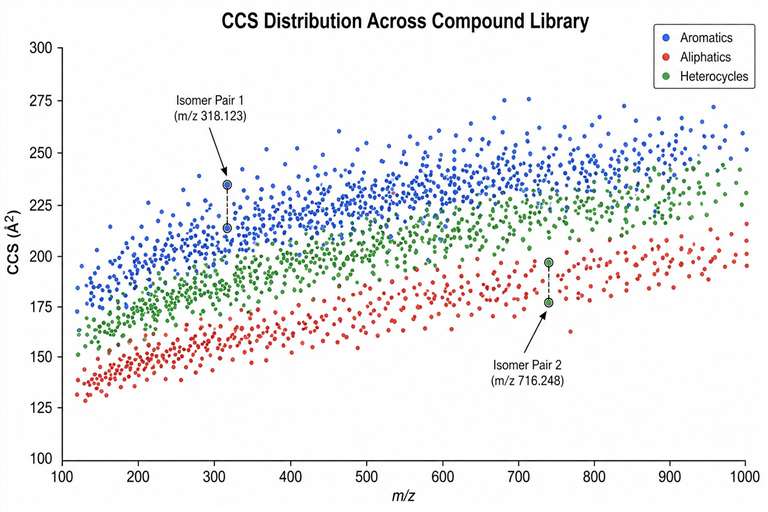
CCS profiling means measuring these values systematically and using them to screen compounds. Build a CCS library of known compounds, compare unknowns against it, and you can identify what you are looking at with far more confidence than mass alone provides — especially when dealing with isomers or compounds that look identical on a standard LC-MS run.
Key Advantages:
- Orthogonal identification dimension — CCS values are independent of chromatography and matrix effects, providing a third dimension alongside m/z and retention time.
- Isomer and isobar resolution — Structural isomers and isobaric compounds that co-elute or share identical mass spectra are routinely distinguished by CCS.
- High-throughput capability — CCS data acquired in milliseconds per compound on your existing LC-MS run. No time penalty, no extra sample preparation.
- Database-driven identification — Match against AllCCS, CCSondemand, or your own proprietary library. Machine learning prediction extends coverage to compounds without reference standards.