Background
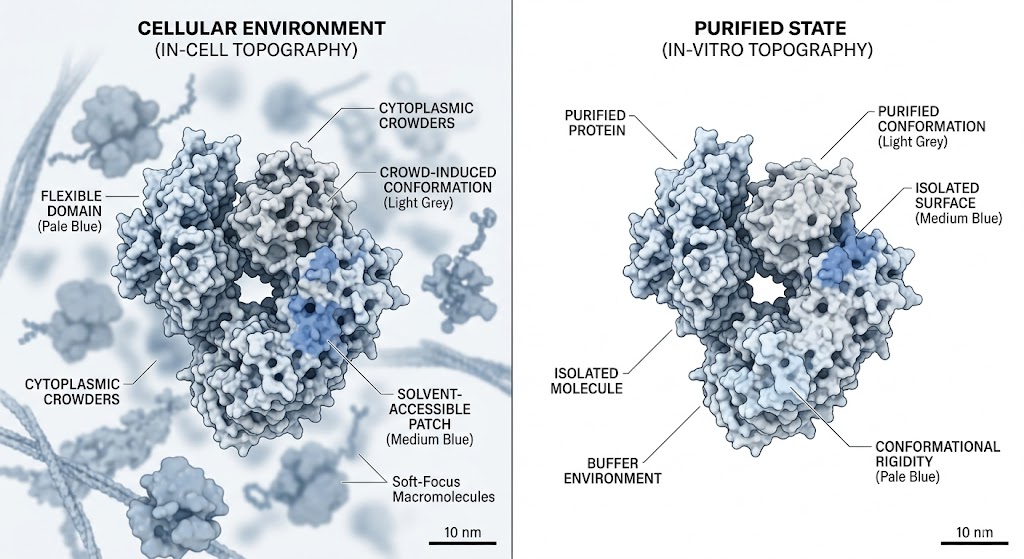
Traditional structural techniques often fail to capture highly dynamic or transient protein conformations. This is largely due to slow labeling speeds in footprinting assays or the requirement for rigid, unnatural crystallization in X-ray methodologies. Researchers required a powerful analytical method to map the true, solvent-accessible surfaces of proteins as they exist in a dynamic, liquid environment, without altering their natural folding pathways or capturing only a time-averaged structural blur.
Methods
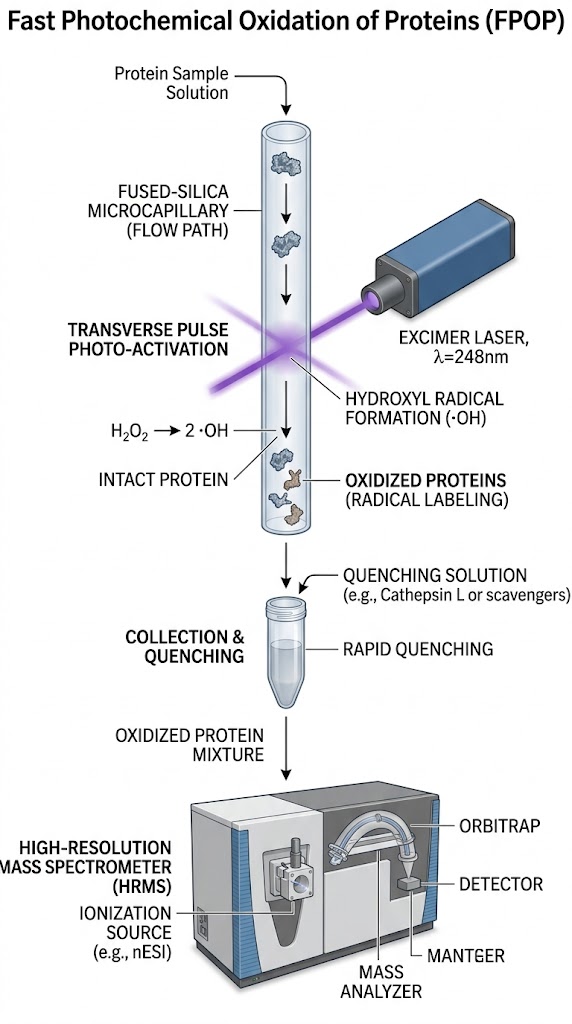
Researchers utilized FPOP-MS to overcome these fundamental biophysical limitations. The target proteins were exposed to a precisely controlled, pulsed UV laser system that photolyzed an optimized concentration of hydrogen peroxide, generating a brief, intense burst of hydroxyl radicals. These highly reactive radicals covalently modified the solvent-exposed amino acid side chains within a sub-microsecond timeframe. The radical lifetime was strictly limited by a scavenger molecule. The samples were then rapidly quenched, enzymatically digested, and analyzed using high-resolution liquid chromatography-tandem mass spectrometry (LC-MS/MS).
Results
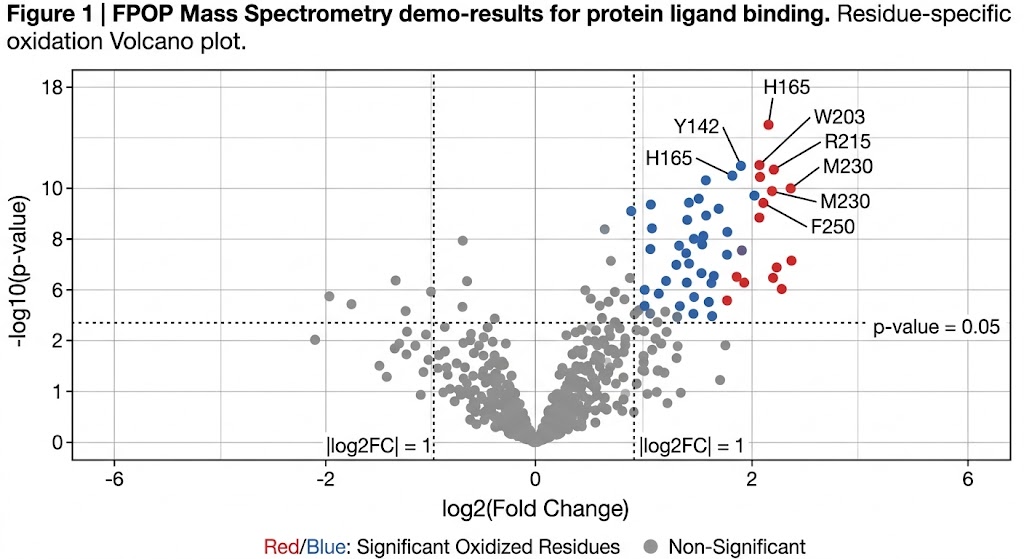
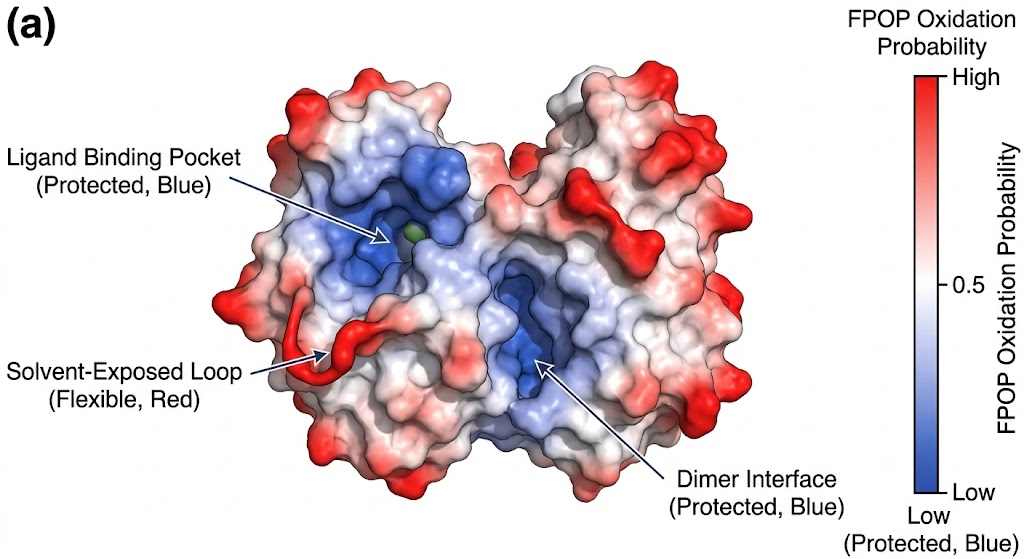
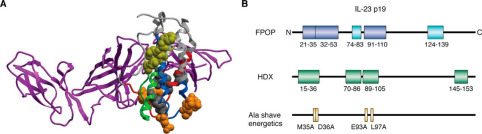
As detailed in the structural mapping data (specifically highlighted in the mass spectra and topographical mapping figures of the study), FPOP successfully labeled the target proteins faster than any unfolding events could physically occur. The high-resolution mass spectrometry analysis accurately identified specific oxidized residues (such as prominent modifications on exposed methionines, tyrosines, and tryptophans). By calculating the oxidation fractions, the researchers created a high-fidelity, residue-level topographical map of the protein's native conformation, successfully pinpointing previously hidden interaction interfaces and solvent channels.
Conclusion
FPOP-MS provides incredibly robust, residue-level structural proteomics data. It is an exceptionally powerful tool for investigating higher-order protein structures and fast-kinetic dynamics that easily evade classical analytical methodologies, supporting critical mechanism-of-action decisions in early-stage drug design and discovery.