What Are Kinase MS Activity Assays?
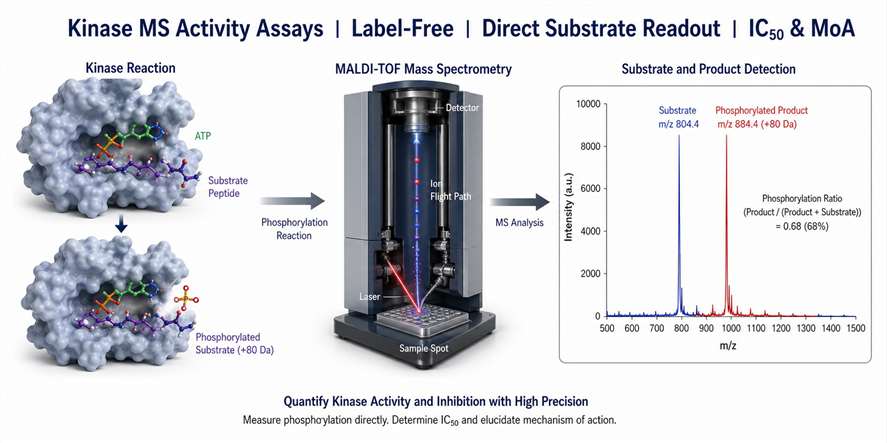
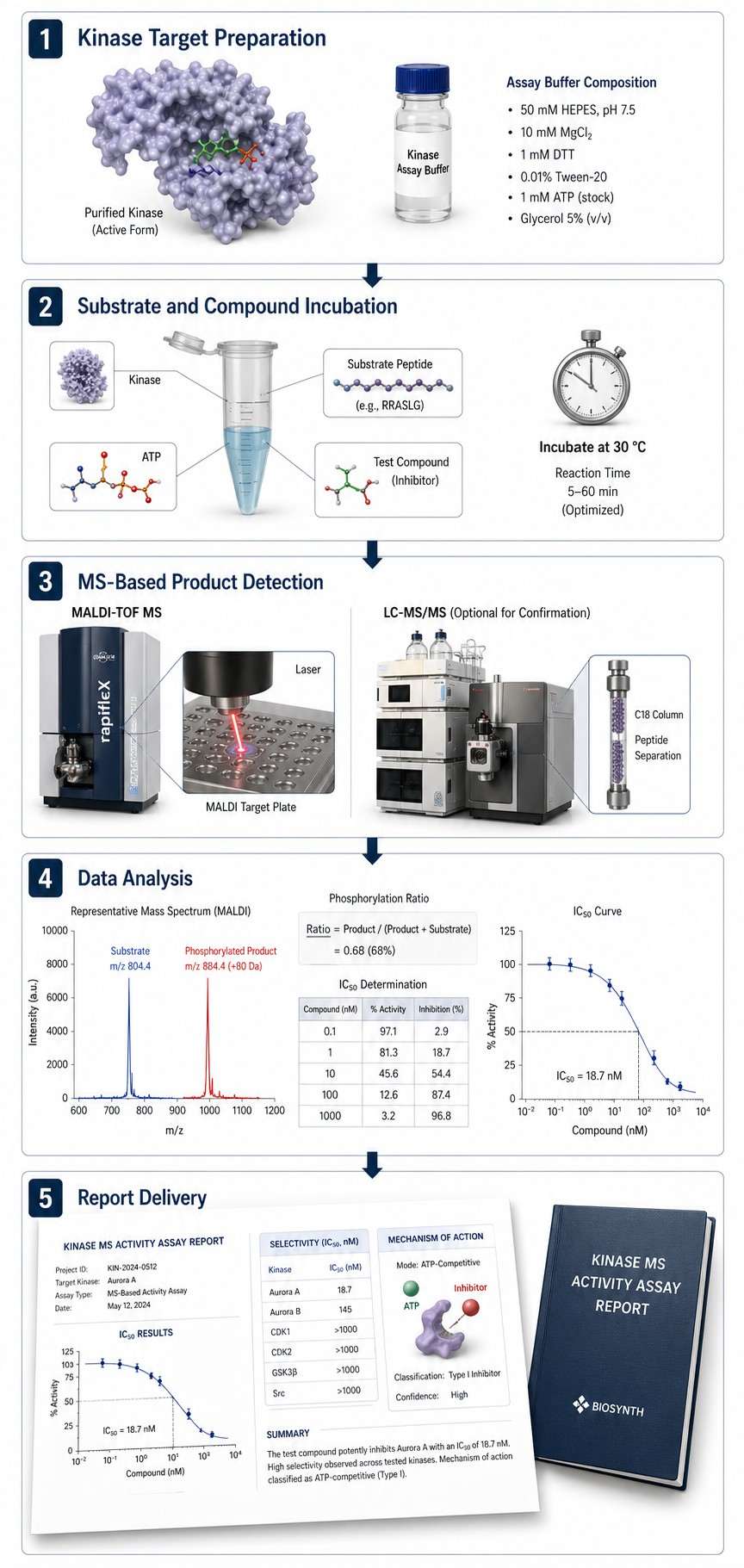
Kinase MS activity assays use mass spectrometry to measure protein kinase catalytic activity in vitro. A purified kinase is incubated with its substrate (peptide or full-length protein), ATP, and test compounds. Mass spectrometry then directly detects and quantifies the phosphorylated product and unphosphorylated substrate, yielding a precise phosphorylation ratio that reports on kinase activity.
Unlike traditional biochemical kinase assays that rely on indirect readouts — fluorescence polarization, FRET, luminescence (ADP-Glo), or radioactivity (³³P-ATP) — MS-based detection measures the actual mass shift of phosphorylation (+80 Da) on the substrate. This direct measurement eliminates several common sources of assay interference: compound autofluorescence in fluorescence-based formats, signal quenching in luminescent readouts, and the need for coupled enzyme systems that can introduce artifacts.
The approach is compatible with both MALDI-TOF and LC-MS/MS platforms. In MALDI-TOF mode, samples are spotted directly onto a target plate after the reaction is quenched, enabling rapid acquisition at rates suitable for moderate-throughput screening. LC-MS/MS mode adds chromatographic separation as an extra dimension of specificity — especially useful for complex substrate mixtures or low-conversion reactions where higher sensitivity is needed.
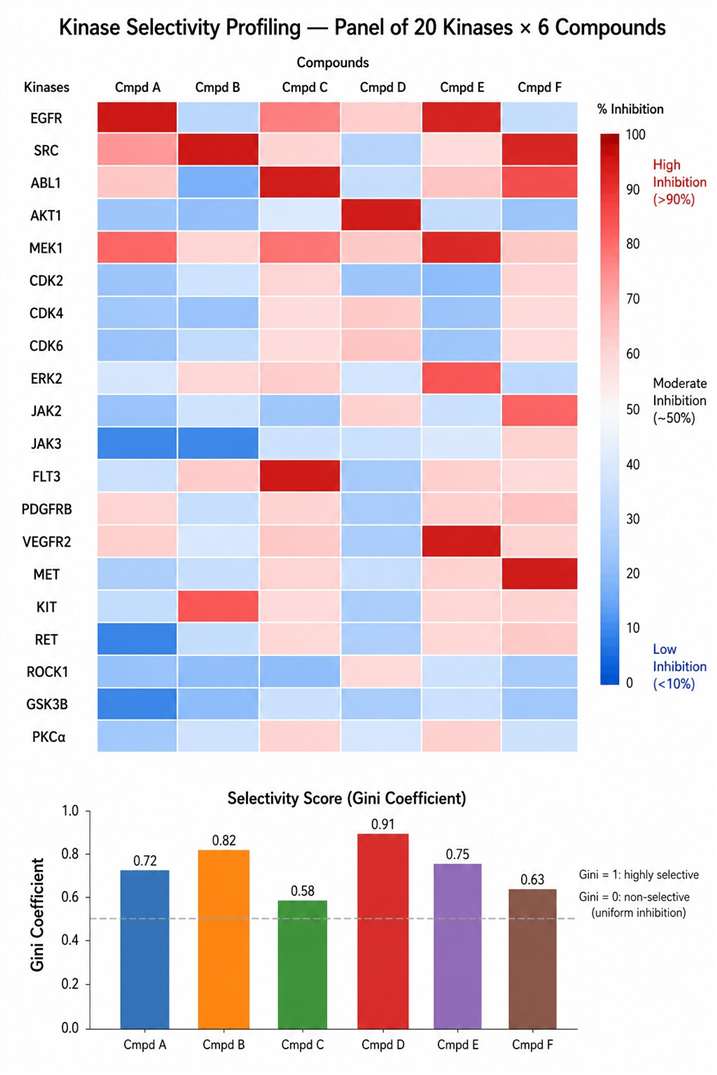
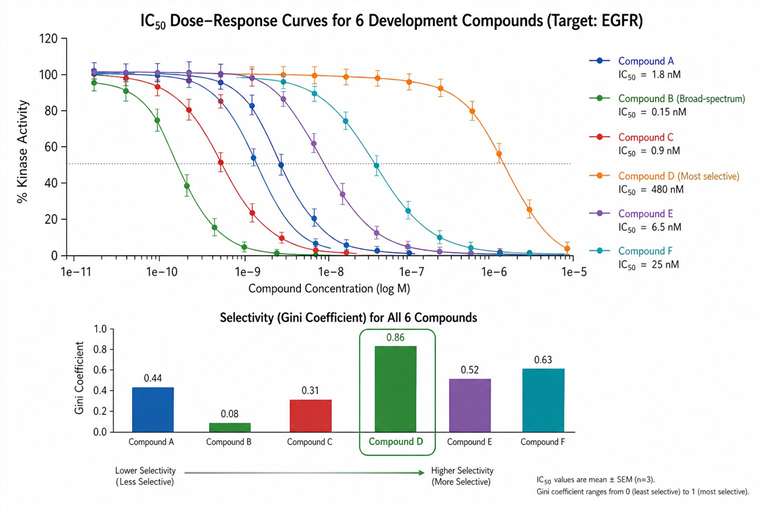
Kinase MS activity assays are particularly well suited for drug discovery programs where compound interference with optical readouts is a known problem, where direct substrate phosphorylation data is preferred over surrogate signals, and where detailed kinetic characterisation (Kₘ, Vₘₐₓ, mechanism of inhibition) is required alongside routine IC₅₀ determination.