Proteome Integral Solubility Alteration (PISA) Service
Accelerate your hit deconvolution and ligand discovery with our high-throughput, label-free Proteome Integral Solubility Alteration (PISA) service.
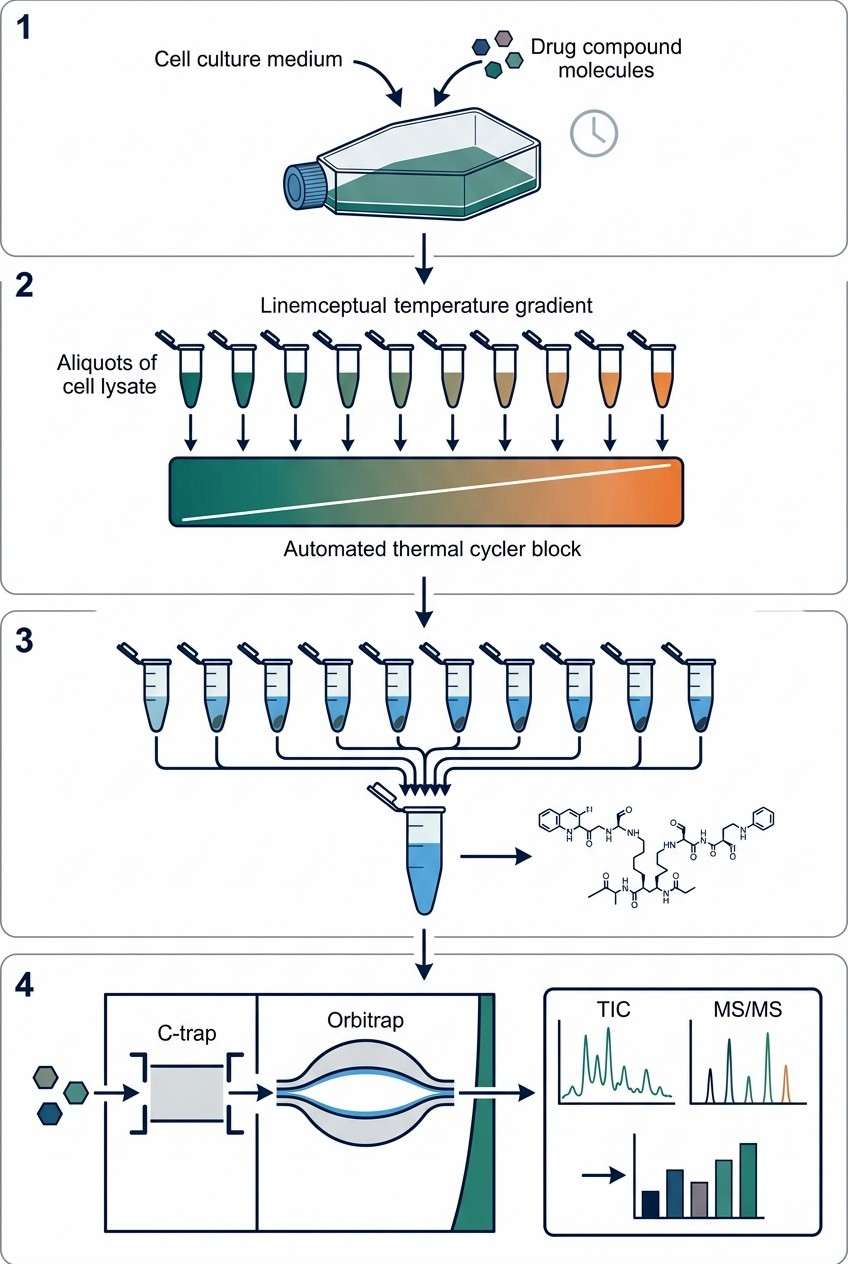
When you have multiple phenotypic hits or are working with incredibly scarce primary cell samples, traditional thermal profiling methods often hit a wall. Generating detailed thermal melt curves for every single compound requires massive amounts of biological material, lengthy mass spectrometry run times, and significant budgets. At Creative Proteomics, we help you overcome these early-stage discovery bottlenecks.
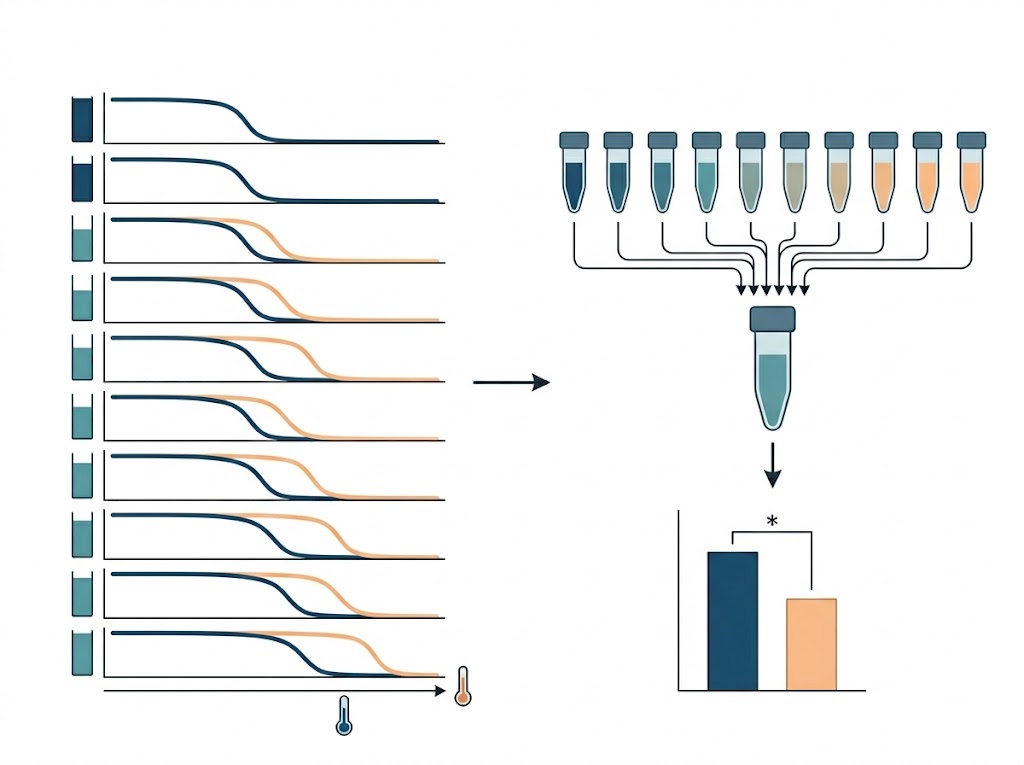
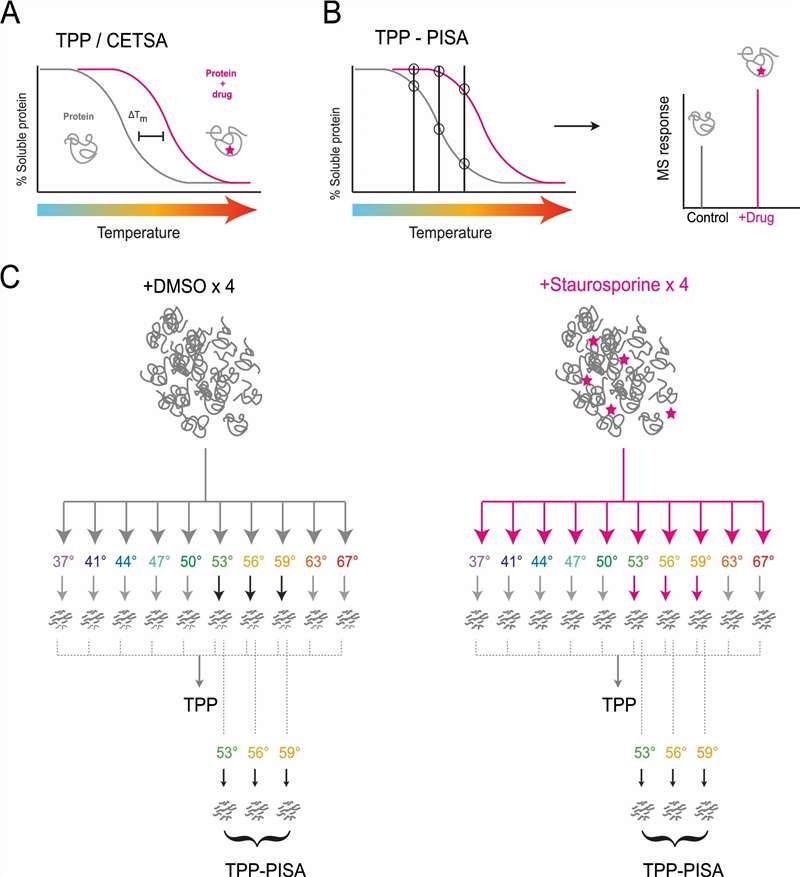
Our PISA service is specifically engineered to compress the thermal shift workflow. By leveraging intelligent sample pooling and high-resolution multiplexed mass spectrometry, we deliver up to a 10-fold increase in screening throughput. This allows you to rapidly evaluate compound-target interactions and conduct proteome-wide toxicity triaging without compromising the depth of your data or forcing you to use artificial chemical tags.
Key Benefits for Your Discovery Pipeline:
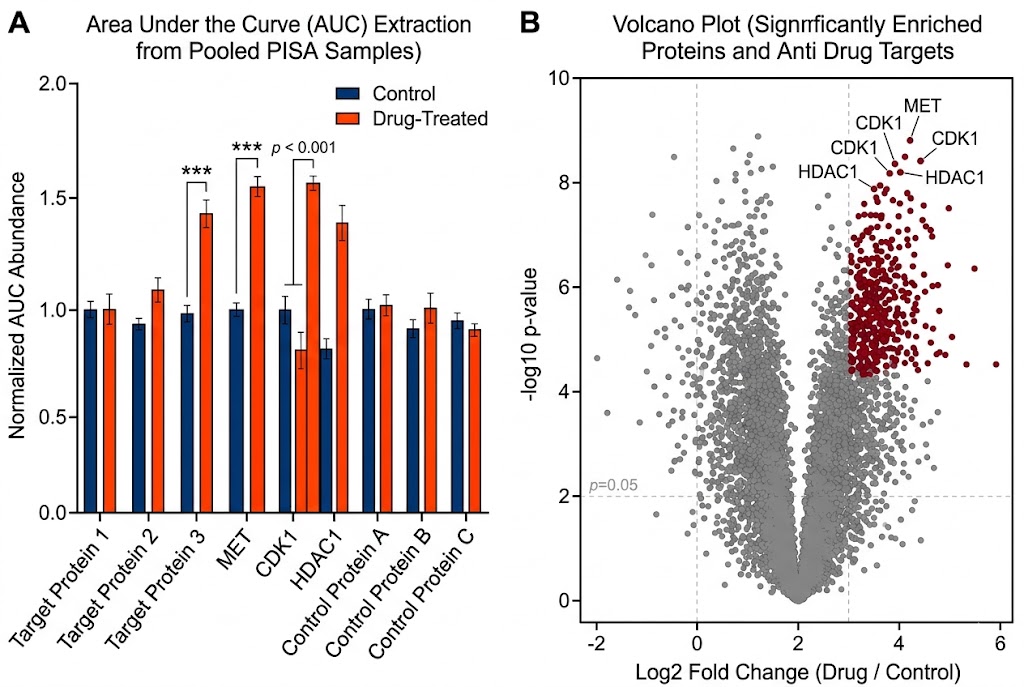
- Massive Throughput Gains: By analyzing the pooled AUC, we can screen multiple compounds, concentrations, or physiological states in a fraction of the time required by conventional assays.
- Unprecedented Sample Savings: Because we compress the analysis into pooled fractions, PISA requires significantly less starting material, making it ideal for precious patient biopsies or hard-to-culture primary cell lines.
- True Native State Screening: Your compounds require absolutely no chemical modification, linkers, or immobilization. We test the drug exactly as it will enter the clinic, ensuring we capture true physiological binding affinities.