Carboxyl Group Footprinting Mass Spectrometry and Molecular Dynamics Identify Key Interactions in the HER2-HER3 Receptor Tyrosine Kinase Interface. https://www.jbc.org/article/S0021-9258(20)49130-6/fulltext
Background
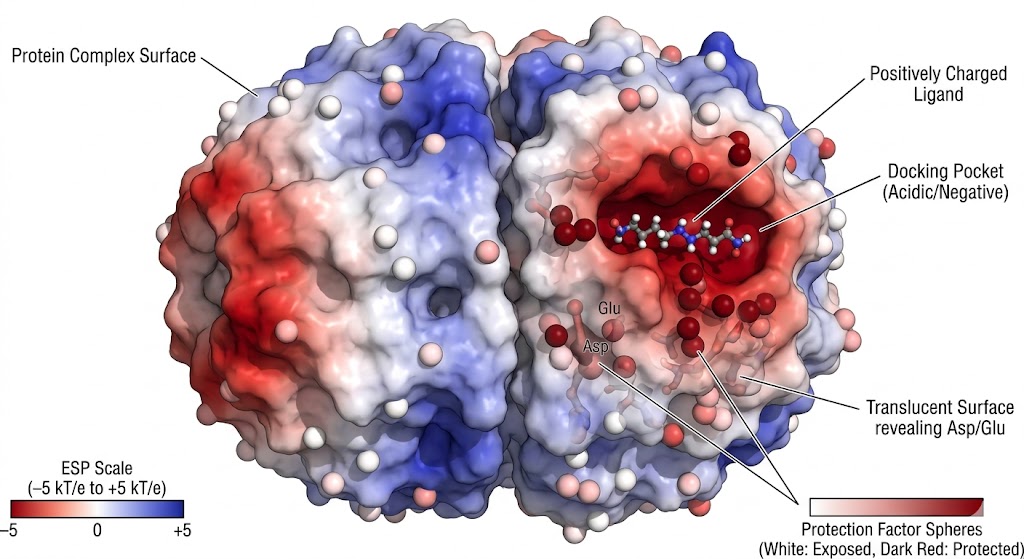
The HER2-HER3 receptor tyrosine kinase heterodimer is a critical, potent oncogenic driver in numerous breast and gastric cancers. Developing therapeutics that successfully block this specific dimerization is a major biopharmaceutical industry goal. However, capturing the highly dynamic, transient nature of the HER2-HER3 interface in solution using static X-ray crystallography is incredibly challenging. Researchers required an advanced, high-resolution footprinting method to map the precise electrostatic interactions driving the complex formation in a native liquid environment.
Methods
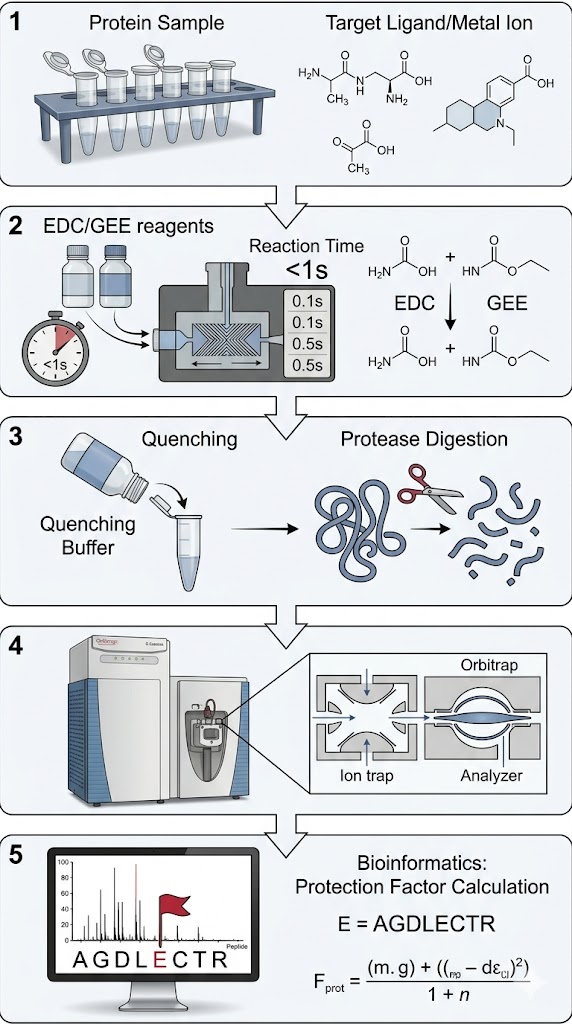
Researchers deployed Carboxyl Group Footprinting Mass Spectrometry, combined with computational Molecular Dynamics (MD) simulations, to deeply interrogate the acidic residues at the heterodimer interface. The isolated HER2 and HER3 kinase domains, as well as the fully assembled complex, were gently labeled in parallel using EDC and a stable nucleophile. The covalently labeled proteins were subsequently digested and analyzed via high-resolution LC-MS/MS to pinpoint and quantify the exact modification extent of the solvent-exposed Asp and Glu residues across the entire receptor.
Results
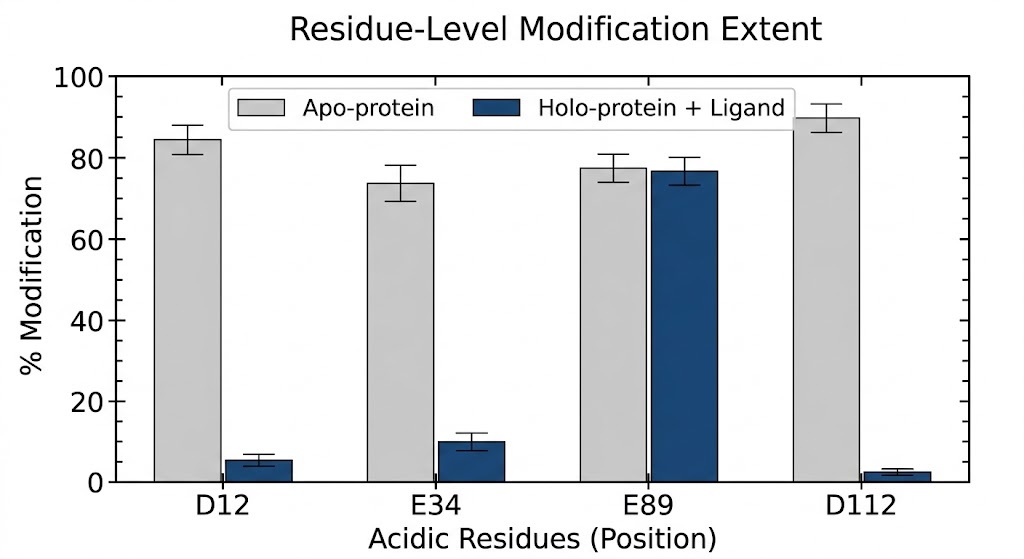
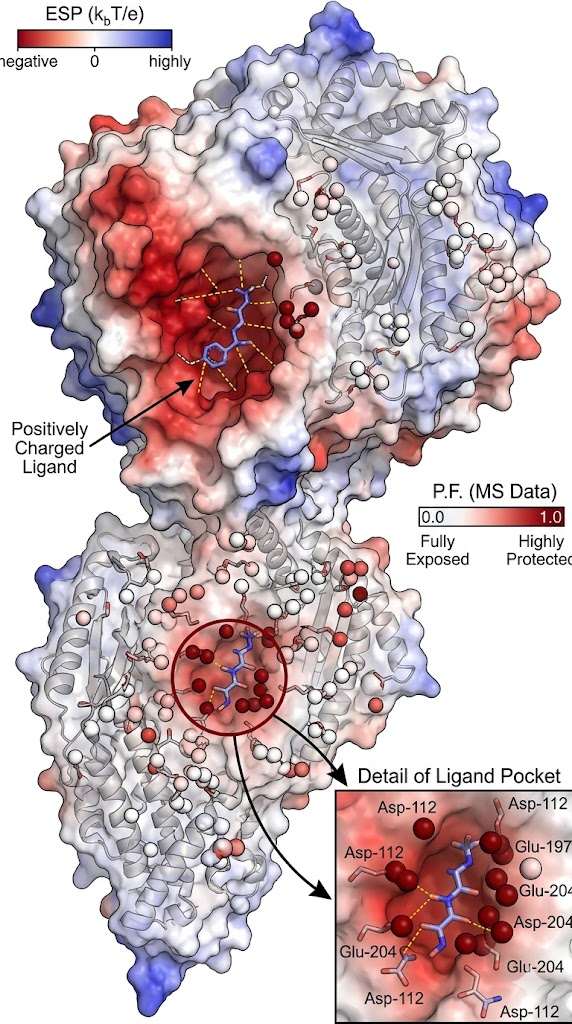
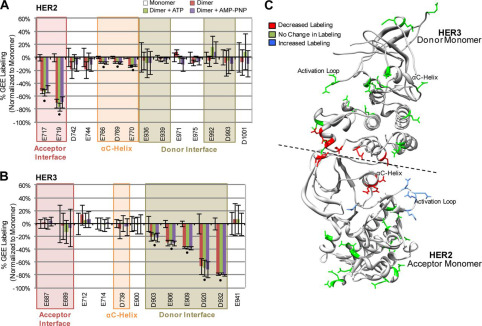
The advanced mass spectrometry data successfully pinpointed the key electrostatic "hotspots" driving the oncogenic dimerization. As specifically illustrated in Figure 2 of the referenced structural study, the footprinting analysis revealed significant, highly localized structural protection directly at the dimer interface. Crucially, the data achieved true single-residue resolution, demonstrating a massive, statistically significant decrease in labeling specifically at residues Glu-922 and Asp-948 when the distinct proteins formed the intact complex. These experimental mass spectrometry results perfectly correlated with the computational MD simulations, proving definitively that these two acidic residues act as essential, solvent-shielded structural anchors for the heterodimer.
Conclusion
Carboxyl Footprinting MS provided unprecedented, residue-level structural evidence of the HER2-HER3 interaction interface. By precisely locating the protected acidic residues in native solution, the biophysical study provided medicinal chemists with a highly accurate 3D roadmap for developing novel, allosteric small-molecule inhibitors designed to physically disrupt this critical oncogenic dimerization event.