Background
Houttuynia cordata, a well-known medicinal plant, has demonstrated significant potential in managing blood sugar levels, but identifying the exact pharmacodynamic substances responsible for this anti-diabetic effect within its highly complex botanical matrix has historically been a massive analytical challenge. Researchers sought a robust, high-resolution method to directly screen the plant's crude extract for specific molecules that inhibit α-glucosidase, a key enzyme in glucose metabolism directly linked to type 2 diabetes.
Methods
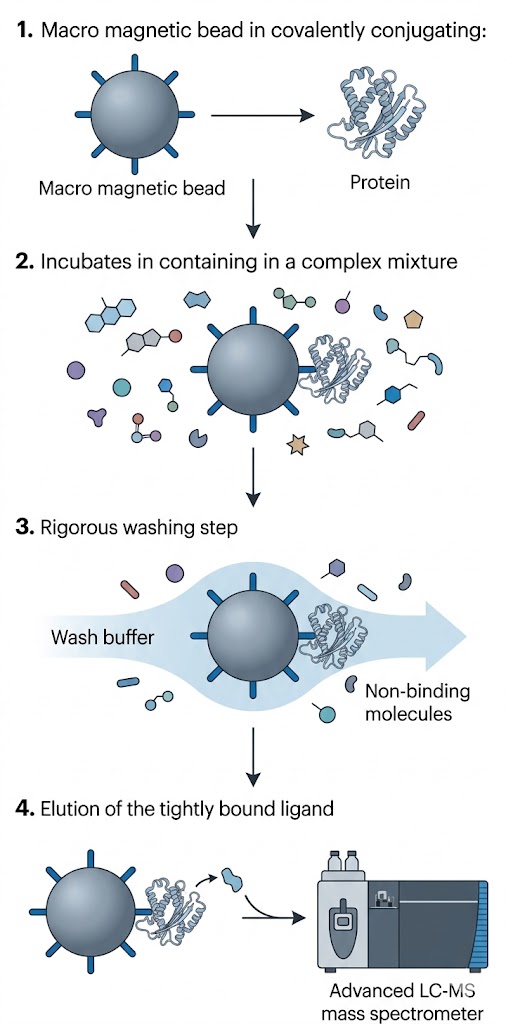
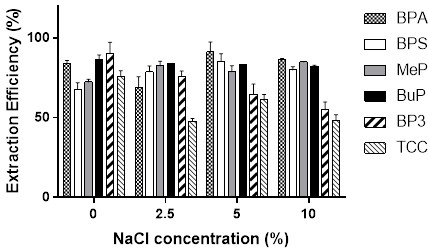
To bypass the slow, expensive, and tedious process of traditional bioassay-guided fractionation, the research team utilized a magnetic bead-based target fishing approach. They successfully immobilized the α-glucosidase enzyme onto carboxyl-terminated magnetic beads using an optimized covalent coupling strategy that protected the active site. The crude extract of Houttuynia cordata was then incubated with both the target-immobilized beads and a set of blank control beads. After rigorous washing with modified buffers to remove non-binding matrix interference and polar contaminants, the tightly bound ligands were released using a targeted solvent elution. The eluate was then analyzed using Ultra-High-Performance Liquid Chromatography coupled with Quadrupole Time-of-Flight Mass Spectrometry (UHPLC-QTOF MS) operating in both positive and negative electrospray ionization modes to ensure maximum coverage of potential phytochemicals.
Results
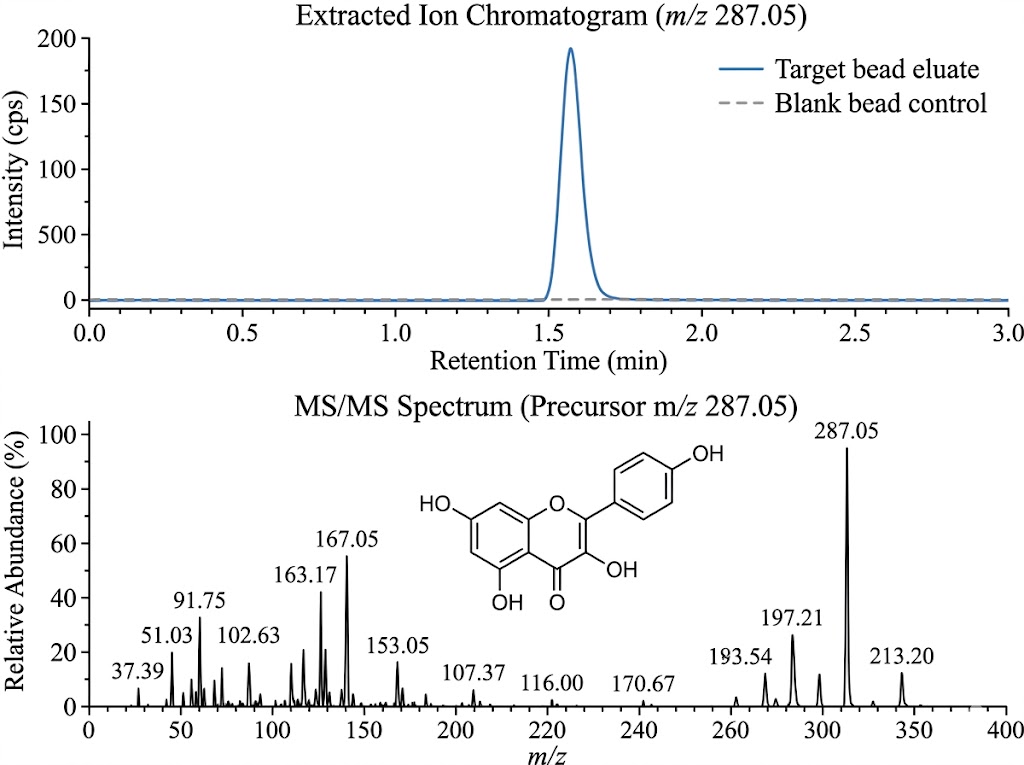
The target fishing workflow effectively isolated the active compounds from the chaotic background of the crude extract. By comparing the UHPLC-QTOF MS Extracted Ion Chromatograms (XIC) of the target beads against the blank beads, the bioinformatic team confidently eliminated false positives and isolated 9 distinct, high-affinity peaks that exclusively appeared in the target bead eluate. Leveraging high-resolution MS/MS fragmentation patterns and matching them against comprehensive natural product spectral libraries, the researchers successfully identified these 9 active α-glucosidase inhibitors. The structural elucidation revealed them to be 3 chlorogenic acid isomers, 2 flavonoid C-glycosides, and 4 flavonoid O-glycosides, effectively mapping the plant's multi-component mechanism of action.
Conclusion
This study clearly demonstrates that target immobilization coupled with high-resolution LC-MS/MS is an incredibly powerful and efficient tool for unraveling the mysteries of complex natural products. By deploying this target fishing workflow, discovery teams can rapidly bridge the gap between crude botanical extracts and identified, actionable lead compounds for metabolic disease therapeutics, saving months of analytical effort and preserving the integrity of the natural compounds.
Source verification: A magnetic beads-based ligand fishing method Coupled with UHPLC-QTOF MS for screening and identification of α-glucosidase inhibitors from Houttuynia cordata Thunb