Affinity Selection Mass Spectrometry (ASMS) for High-Throughput Hit Discovery
Accelerating hit identification and ligand discovery through high-throughput, label-free screening.
Affinity Selection Mass Spectrometry (ASMS) has emerged as a critical platform for modern drug screening, enabling researchers to rapidly evaluate compound–target interactions without the need for reporter assays or target immobilisation.
At Creative Proteomics, our ASMS service is designed to address discovery-stage challenges: limited in-house mass spectrometry resources, long assay development times, and the need for robust, data-rich hit lists.
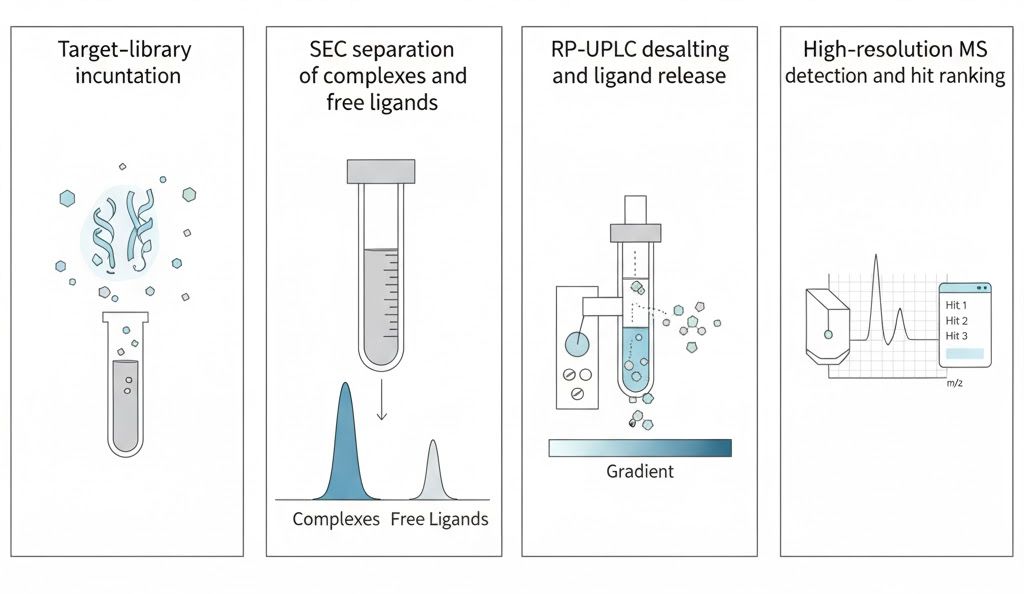
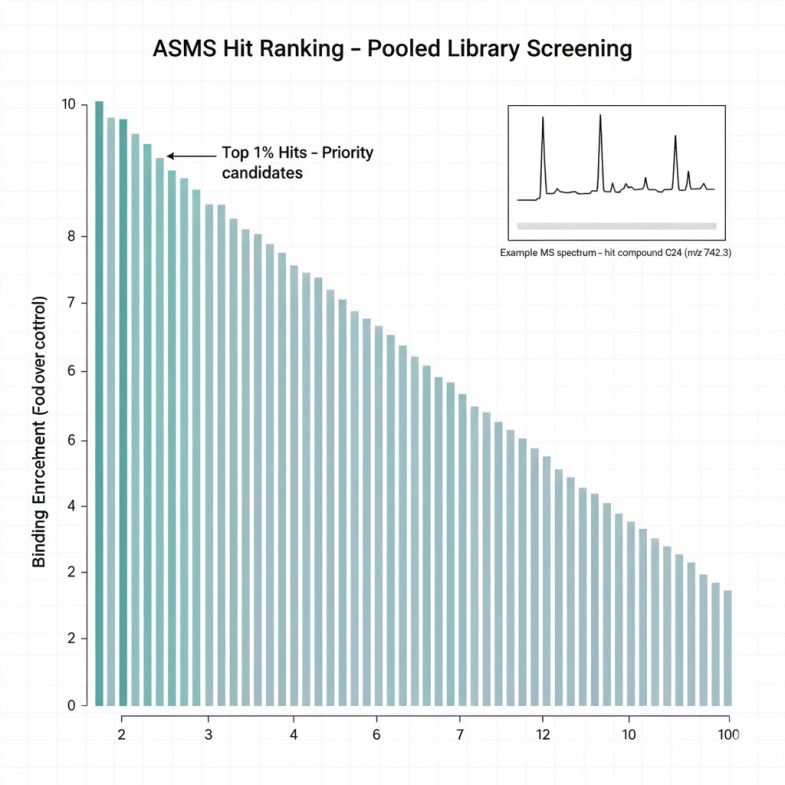
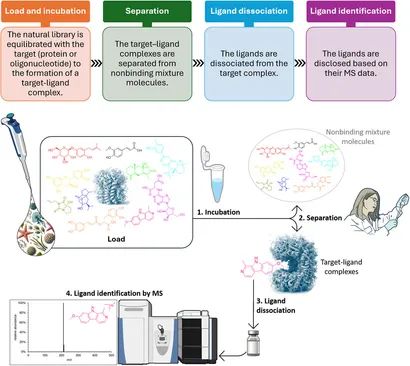
Leveraging our integrated SEC–RP-UPLC → high-resolution MS workflow, we deliver high-throughput, data-rich hit lists for small molecules, fragments, and macrocyclic/linear peptide libraries.
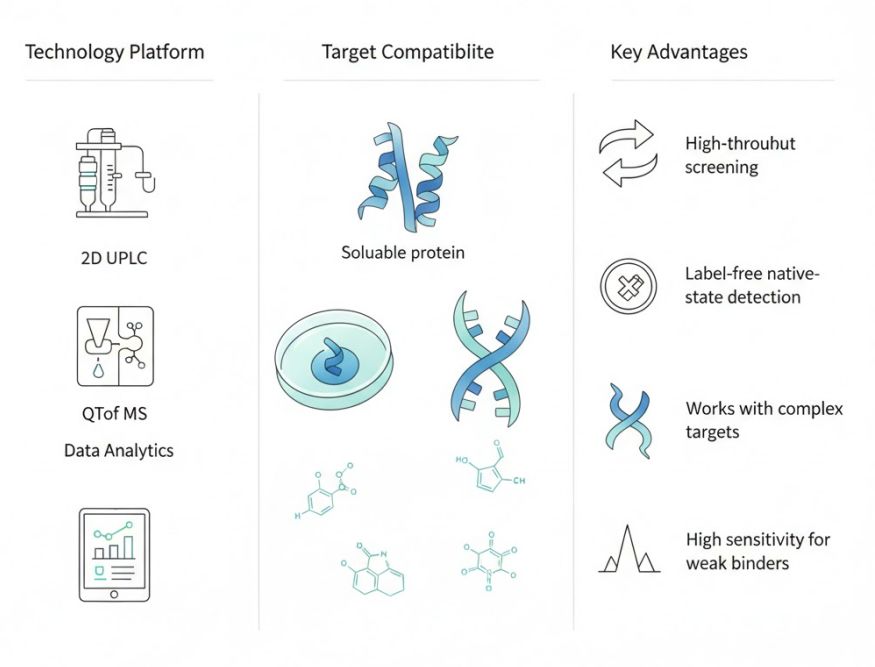
Key Advantages:
- High-throughput screening of mixed compound pools, compatible with affinity selection mass spectrometry screening workflows.
- Support for challenging targets, including membrane proteins, protein complexes, and induced-proximity modalities (e.g., molecular glues).
- Rapid turnaround times and scalable throughputs across biotech, pharma, and academic discovery programmes.
- Robust data output and documentation ready for both internal review and external partnerships.