Photo-crosslinking Structural MS Service for Binding Site Mapping and Mechanism Studies
Capture ligand engagement, interpret binding regions, and support structure-aware project decisions.
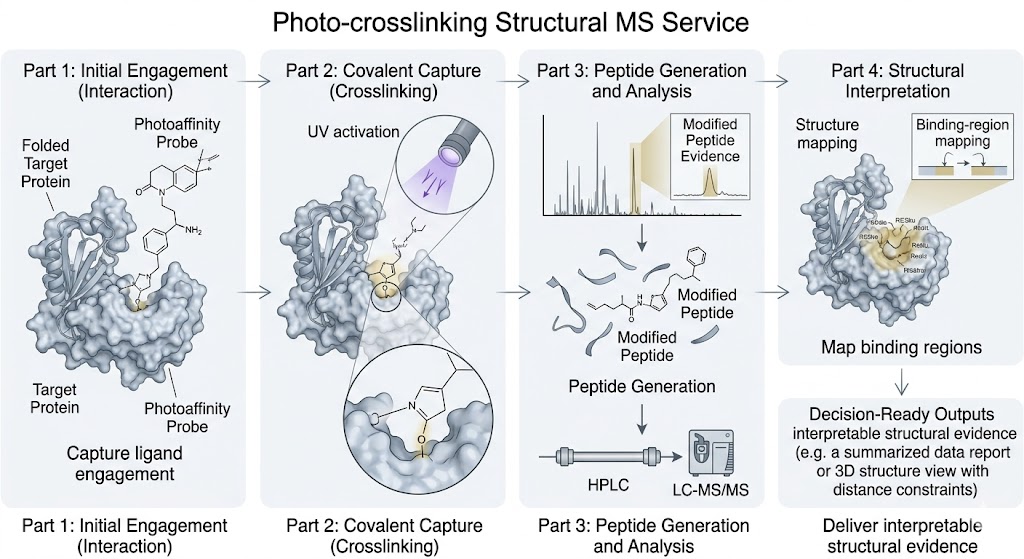
Photo-crosslinking Structural MS adds a direct evidence layer when a compound shows activity but the binding region is still unclear. By combining photoactivated capture with LC-MS/MS-based interpretation, we help you study ligand engagement, weak or transient interactions, and structure-aware mechanism questions in a format your team can review and act on.
This workflow is especially useful when standard function-only assays cannot provide enough spatial context, when interactions are difficult to stabilize, or when you need a method that bridges biochemical observations and deeper structural follow-up.
At Creative Proteomics, we do not treat photo-crosslinking as a standalone checkbox assay. We place it inside a broader MS evidence chain that can extend from hit support to target deconvolution, mechanism evidence, and orthogonal follow-up when your study needs more than one readout.
Key Advantages:
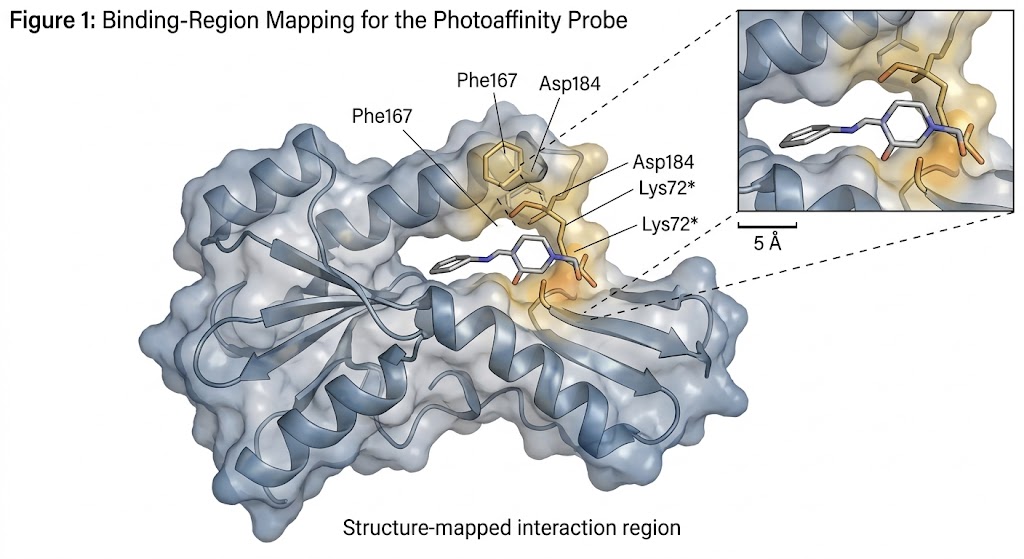
- Binding-region-focused structural evidence for small-molecule engagement.
- Strong utility for weak or transient interactions that are difficult to interpret with standard assays.
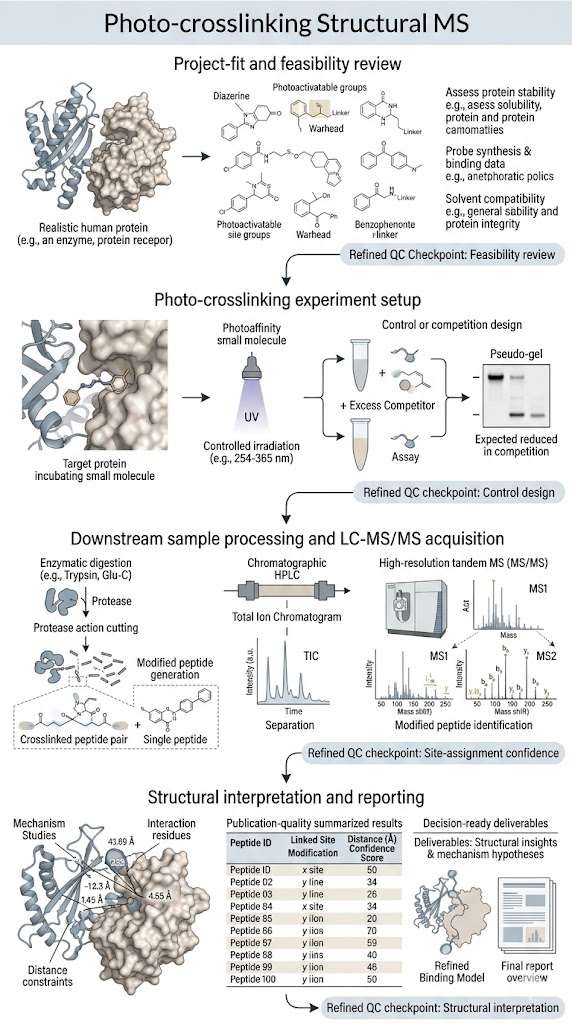
- Explicit workflow and QC checkpoints from feasibility review to final reporting.
- Decision-ready deliverables designed for technical review, not just raw spectra.