Overcoming the Challenges of Target Deconvolution
In modern drug discovery, researchers frequently utilize phenotypic screening to determine which compounds effectively alter a specific cellular behavior. This approach is incredibly powerful because it guarantees the compound is cell-permeable and functional within a real biological system. However, it leaves researchers with a major developmental bottleneck: elucidating the unknown target, referred to as target deconvolution.
Traditionally, scientists have relied on "pull-down" affinity chromatography. While historically useful, this method frequently fails because many small molecules exhibit transient or relatively weak interactions with their targets. When the solid support is subjected to stringent washing steps, the authentic target simply dissociates and is lost.
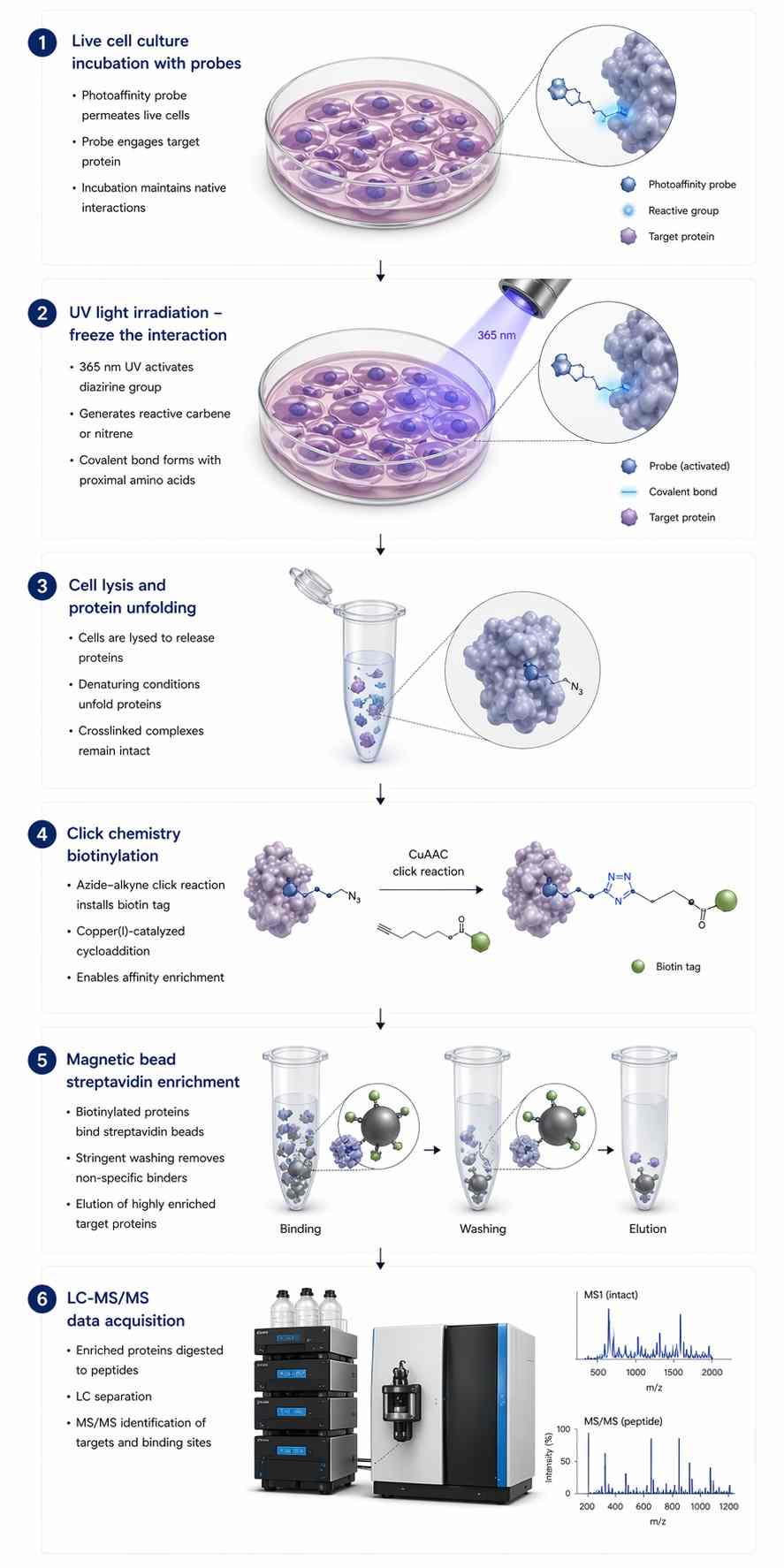
Photoaffinity Labeling (PAL) paired with Mass Spectrometry (MS) elegantly solves this dissociation problem. We synthesize a customized version of your active drug equipped with a photoreactive group. When we irradiate the live cells with ultraviolet (UV) light, this tag generates a highly reactive intermediate that forms a permanent, irreversible covalent bond with whatever amino acid residue it is physically touching at that exact microsecond. Because the interaction is now permanently locked, we can deploy extremely harsh denaturing washing conditions to quantitatively strip away all noisy background proteins, leaving only the true targets ready for identification.