Phospholipids are often introduced as simple amphiphiles with a polar head and two hydrophobic tails. That definition is accurate, but it is too blunt for anyone trying to understand membrane behavior, signaling logic, or species-level lipid analysis. In real systems, phospholipids are not generic membrane bricks. They are structurally encoded regulators of surface charge, lateral packing, curvature stress, leaflet asymmetry, and downstream signal generation. The same diversity that shapes membrane physics also shapes how lipids ionize, fragment, and overlap in LC-MS/MS. A serious phospholipid article therefore has to move past the standard bilayer cartoon and treat the phospholipidome as a coupled chemical and biophysical system.
This first half builds that foundation. It starts with molecular architecture, then moves into interfacial chemistry, membrane thermodynamics, packing constraints, and sterol-dependent lateral organization. That order matters. Without structural literacy, membrane fluidity becomes a vague slogan. Without membrane physics, lipid signaling seems disconnected from molecular form. And without both, later phospholipidomics becomes little more than a peak list.
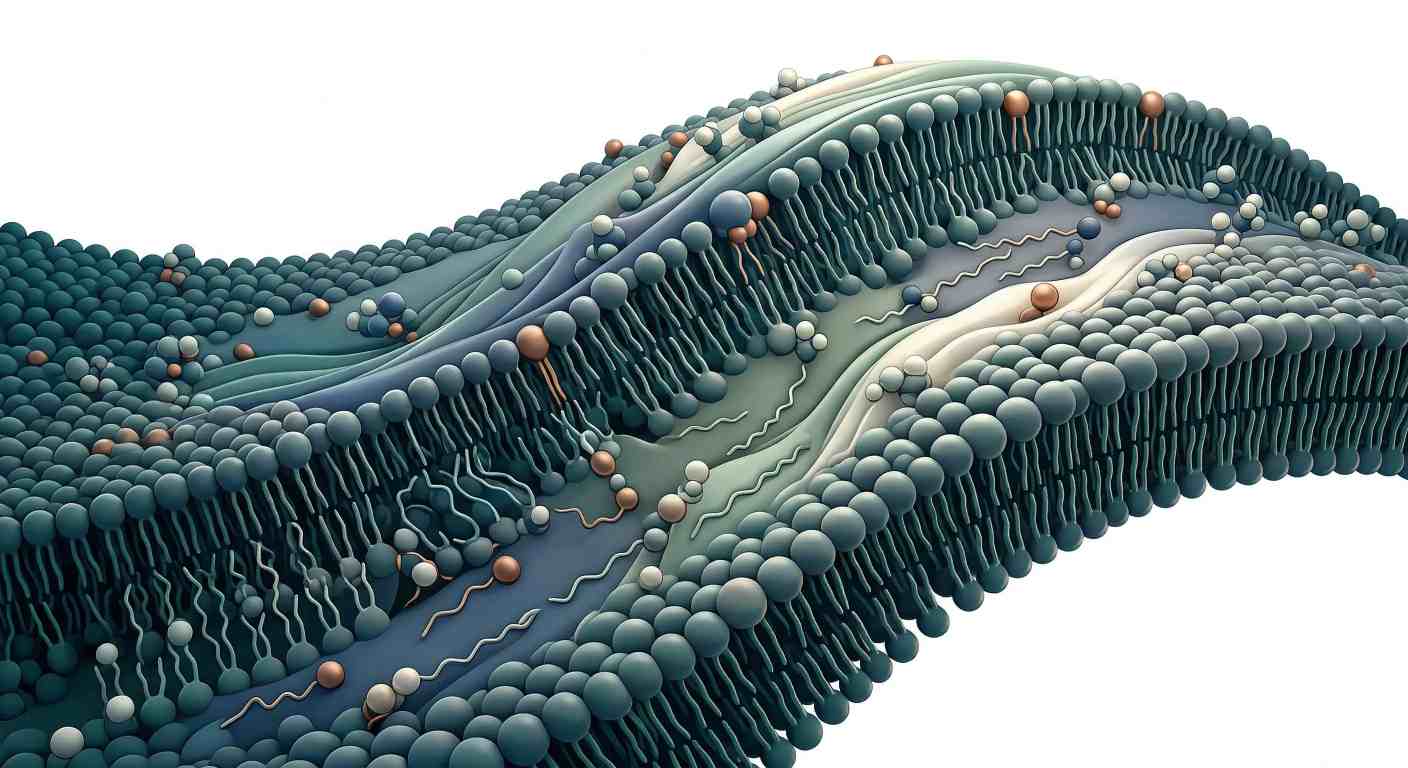 Figure 1: The Phospholipidome at a Glance: Chemical Diversity Across the Membrane Landscape
Figure 1: The Phospholipidome at a Glance: Chemical Diversity Across the Membrane Landscape
The Molecular Anatomy of Phospholipids
Glycerophospholipids and sphingophospholipids do not follow the same blueprint
The first distinction that matters is the one between glycerophospholipids and sphingophospholipids. Both belong to the phospholipid world, but they are not built on the same scaffold.
Glycerophospholipids use a glycerol backbone. Its three carbons are designated sn-1, sn-2, and sn-3 under the stereospecific numbering system. In the common arrangement, fatty acyl chains occupy sn-1 and sn-2, while the phosphate-bearing head-group region attaches at sn-3. This is not a naming convenience. It is a functional map. It tells you where hydrophobic mass sits, where the interface begins, and where enzyme action can generate different biological products.
Sphingophospholipids follow another structural logic. Instead of glycerol, they use a sphingoid base, typically sphingosine. One hydrophobic segment is embedded in that backbone, while the second hydrocarbon chain is attached through an amide linkage. In membrane biology, the dominant example is sphingomyelin. At the surface, sphingomyelin can resemble phosphatidylcholine because both can present phosphocholine. But the shared head-group appearance hides deeper differences. Backbone chemistry changes packing, hydrogen-bonding behavior, sterol compatibility, and fragmentation behavior in tandem MS.
That is why "phospholipid" should never be treated as one chemically uniform category. A glycerophospholipid carrying ester-linked chains and a sphingophospholipid carrying an amide-linked chain can coexist in the same membrane yet behave very differently. If the goal is only broad discovery across membrane lipid classes, Untargeted Lipidomics can reveal that class-level diversity. But once the question becomes structural or mechanistic, backbone type stops being background information and becomes part of the interpretation itself.
The sn system is a structural coordinate system with biological consequences
The sn-1 / sn-2 / sn-3 system matters because phospholipid behavior is position-sensitive. In many mammalian glycerophospholipids, the sn-1 position is often occupied by a saturated or monounsaturated chain, whereas the sn-2 position more often carries a more unsaturated chain. That pattern is not absolute, but it is common enough to have real consequences for membrane physics and downstream signaling.
Why? Because the two positions are not interchangeable. A phospholipid with a more saturated sn-1 chain and a polyunsaturated sn-2 chain already contains built-in asymmetry. One side supports stronger packing order. The other promotes disorder and often stores future signaling potential. If arachidonic acid occupies sn-2, the molecule is not just a membrane constituent. It is also a latent precursor pool waiting for phospholipase action.
This is the point where structural chemistry stops being abstract. Once the sn positions are taken seriously, it becomes easier to understand why some phospholipids depress local order, why some act as better reservoirs for lipid mediators, and why positional information later becomes valuable in MS/MS interpretation. The molecule is not just a class label with two tails. It is a positionally organized molecular unit.
Head-group diversity controls interfacial behavior, not just phospholipid names
The familiar class list—PC, PE, PS, PI, PG, PA—is often taught as though it were just a taxonomy. It is more useful to read these classes as different solutions to an interfacial design problem.
Phosphatidylcholine (PC) usually supports a more cylindrical geometry because its choline head group is relatively bulky. That shape tends to favor lamellar bilayer organization. Phosphatidylethanolamine (PE) has a smaller head group, which reduces effective interfacial area and increases its tendency to support tighter local packing and negative curvature stress. The difference sounds subtle until the membrane is forced to bend, fuse, or pass through a high-curvature intermediate. Then the contrast becomes decisive.
Phosphatidylserine (PS), phosphatidylinositol (PI), phosphatidylglycerol (PG), and phosphatidic acid (PA) contribute stronger anionic character. But the important point is not only that they are "negative lipids." They change surface potential, hydration, head-group spacing, hydrogen-bonding behavior, and cation affinity. A membrane enriched in anionic phospholipids presents a different interfacial field from one dominated by PC and sphingomyelin, even if total phospholipid abundance is similar.
This is why bulk summaries such as "total phospholipids increased" are often not enough. Two membranes can contain similar total phospholipid mass and still behave very differently if one is enriched in PC and sphingomyelin while the other contains more PE, PS, and PI. The first may support tighter sterol-compatible order. The second may present a more negative interface, stronger curvature stress, and a different protein-binding landscape. When that level of detail is required, Glycerophospholipids Analysis Service and Phospholipids Analysis Service become much more informative than a single bulk phospholipid readout.
Leaflet asymmetry makes phospholipid composition a spatial problem
Biological membranes are not symmetric lipid sandwiches. In many cells, PS and PE are enriched on the cytosolic leaflet, while PC and sphingomyelin are often more abundant on the exoplasmic leaflet of the plasma membrane. This asymmetry is actively maintained. It does not emerge automatically from equilibrium.
The functional consequence is that membrane identity is three-dimensional. A phospholipid is not defined only by class and acyl chains. It is also defined by which leaflet presents it. The same PS species on the inner leaflet and outer leaflet does not mean the same thing. Inner-leaflet PS contributes to membrane charge and cytosolic protein recruitment. Externalized PS can act as a context-dependent membrane-recognition cue in regulated biological processes.
That is why phospholipid structure should be treated as predictive. Once backbone type, sn-position logic, head-group chemistry, and leaflet context are understood together, many later questions become easier to answer. Packing behavior, surface electrostatics, curvature tendency, and analytical behavior all start to look less mysterious.
Surface Chemistry, Charge Distribution, and Interfacial Behavior
The membrane interface is a crowded chemical layer
The membrane interface is where phospholipid chemistry becomes physical. It is also where simplified explanations tend to fail.
A head group does not float in isolation. It sits in a crowded layer containing neighboring head groups, structured water, dissolved ions, and membrane-associated proteins. The effective behavior of that head group therefore depends not only on its formal structure, but also on spacing, hydration, ionic composition, and local crowding.
This is why net charge alone is not enough. Zwitterionic lipids such as PC and PE are net neutral, but they are not electrostatically blank. Their internal charge separation still contributes to dipolar behavior at the surface. Anionic lipids such as PS, PI, PG, and PA create stronger negative interfacial character, but even that depends on protonation state, ionic strength, and surrounding chemical context. Surface potential is therefore not a fixed label. It is an emergent interfacial property.
Head-group spacing changes what proteins and enzymes encounter
Membrane-binding proteins do not see "a membrane" in the abstract. They encounter a surface with a particular charge density, hydration structure, head-group spacing, and mechanical compliance. A PI-rich surface differs from a PC-rich one. A PS-enriched interface creates a different electrostatic landscape for polybasic proteins. A PE-rich region can alter local packing even before any protein binds.
This is one reason phospholipids should not be described only as building materials. They are also surface-defining variables. They determine what kind of interfacial environment a protein encounters, how close two membranes can approach one another, and how readily local membrane deformation can occur.
When the biological question centers on low-abundance signaling pools rather than total phospholipid mass, broad profiling often needs to be narrowed. That is where Targeted Phosphoinositides Analysis Service becomes more useful than a general phospholipid survey, because the aim shifts from class presence to mechanistically meaningful local lipid pools.
Hydration is part of membrane structure, not background solvent
Water at the membrane interface is not equivalent to bulk solvent. It forms a structured hydration layer around phospholipid head groups, and that interfacial water affects spacing, ion association, and the energetic cost of bringing membranes together.
This is easy to miss because hydration is invisible in most simple membrane cartoons. Yet many membrane processes depend on it. Fusion, adhesion, and membrane docking all require partial reorganization of interfacial water. A strongly hydrated head-group layer behaves differently from one whose spacing has already been compressed or whose ionic environment has changed.
Hydration also changes how membranes respond mechanically. A membrane interface that is loosely spaced and strongly hydrated is not equivalent to one that has been compressed by ions or enriched in tightly interacting head groups. The same phospholipid composition can therefore behave differently when its interfacial solvent environment changes.
Divalent cations can reorganize a membrane without changing lipid class abundance
Calcium and magnesium add another control variable. These cations associate preferentially with anionic phospholipids and can reduce repulsion between neighboring head groups. When repulsion falls, interfacial spacing can contract, hydration can shift, and local rigidity may increase.
The practical lesson is important: the same membrane composition can behave differently under different ionic conditions. That matters for vesicle preparation, protein-binding assays, extraction performance, and the interpretation of membrane-state changes. It also explains why phospholipid biology should never be separated too sharply from buffer chemistry.
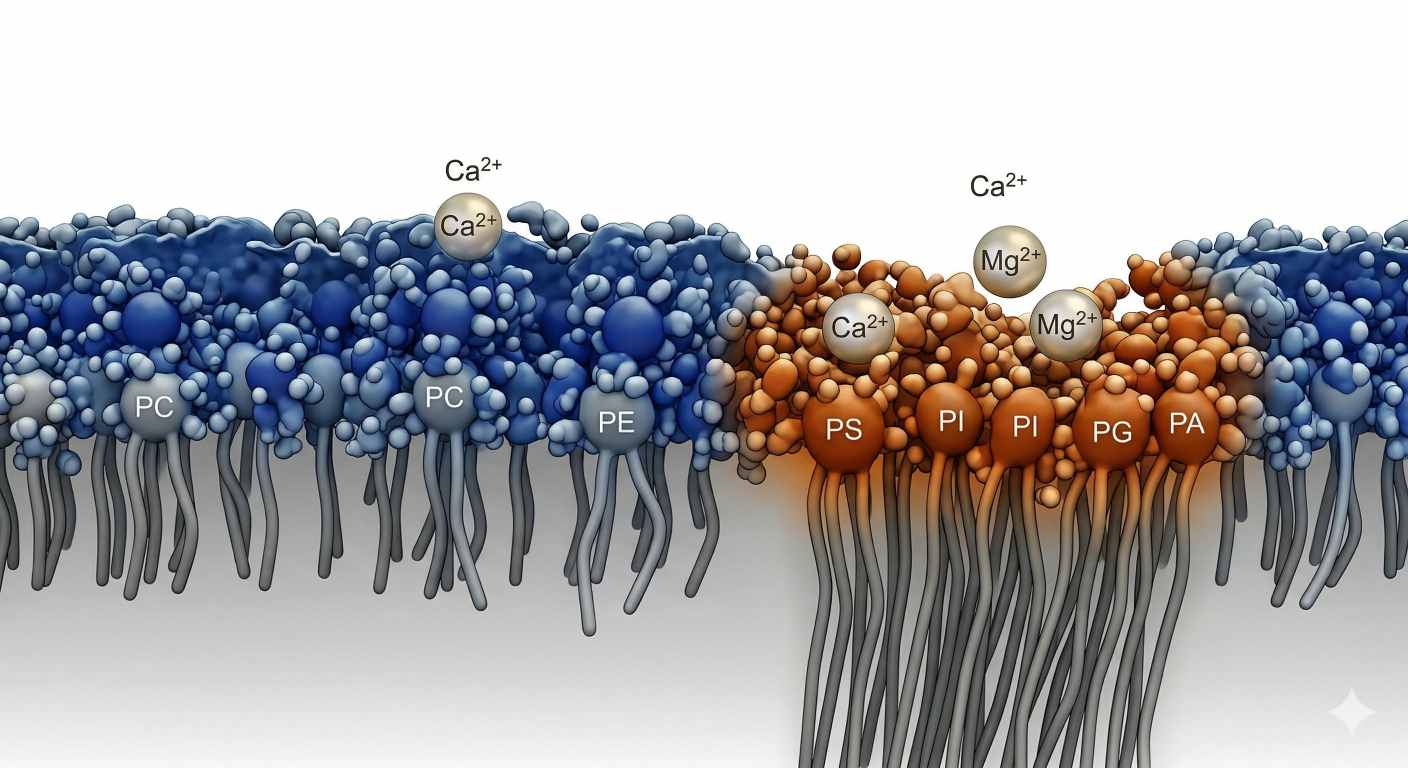 Figure 2: Charged Boundaries: Head-Group Electrostatics, Hydration, and Divalent Ion Binding
Figure 2: Charged Boundaries: Head-Group Electrostatics, Hydration, and Divalent Ion Binding
In real workflows, phospholipid profiling is often not the endpoint. It is the structural readout that has to be interpreted together with buffer composition, pH, and membrane context. This is one reason many studies pair composition data with Lipidomics Service workflows that can be matched to the actual biological question instead of treating every lipid dataset as analytically self-explanatory.
Membrane Thermodynamics: Why Phase State Matters
"Fluidity" is too vague unless phase behavior is specified
Membrane fluidity is often described as though it were one simple adjustable property. That language is convenient, but it hides the real issue. Membranes occupy phase states whose behavior depends on temperature, composition, and intermolecular interactions.
For phospholipid bilayers, the classic transition is the shift from the gel phase (Lβ) to the liquid-crystalline phase (Lα). In the gel state, acyl chains are more extended and closely packed. Lateral motion is restricted, free volume is lower, and order is higher. In the liquid-crystalline state, chain disorder rises, free volume increases, and the bilayer becomes much more dynamic.
This is not just descriptive language. It is a thermodynamic transition with a measurable melting or chain-order transition temperature, (Tm). Once that concept is in place, membrane behavior becomes easier to interpret because the discussion moves from vague fluidity language to a more exact relationship among temperature, composition, and order.
Acyl-chain length and unsaturation are the dominant thermodynamic levers
Two structural variables dominate phospholipid melting behavior: chain length and unsaturation.
Longer hydrocarbon chains raise (Tm) because they increase van der Waals interactions and support tighter ordered packing. Cis double bonds lower (Tm) because they introduce kinks that disrupt close chain alignment. A phospholipid with long saturated chains therefore tends to favor a more ordered state than one containing shorter or more unsaturated chains.
This is where many overview articles stop too early. They correctly say that saturation increases order and unsaturation increases fluidity, but they do not carry that idea through to phase behavior, thickness, compressibility, and membrane-state transitions. In real membranes, acyl chains are not passive tails. They are the primary structural drivers of chain-melting thermodynamics.
Composition acts like a thermodynamic control panel
Once chain composition changes, membrane state changes. A membrane enriched in saturated phospholipids tends to shift toward higher transition temperatures and stronger packing order. A membrane enriched in unsaturated species remains more disordered under the same thermal conditions. The head group still matters, but the acyl chains carry much of the thermodynamic weight.
A polyunsaturated membrane is therefore not simply "more fluid." It is also less tightly packed, more permissive to local disorder, and often less compatible with sterol-driven condensation than a membrane enriched in saturated species. That difference matters for permeability, protein coupling, membrane remodeling, and any attempt to interpret structural lipidomics in physical terms.
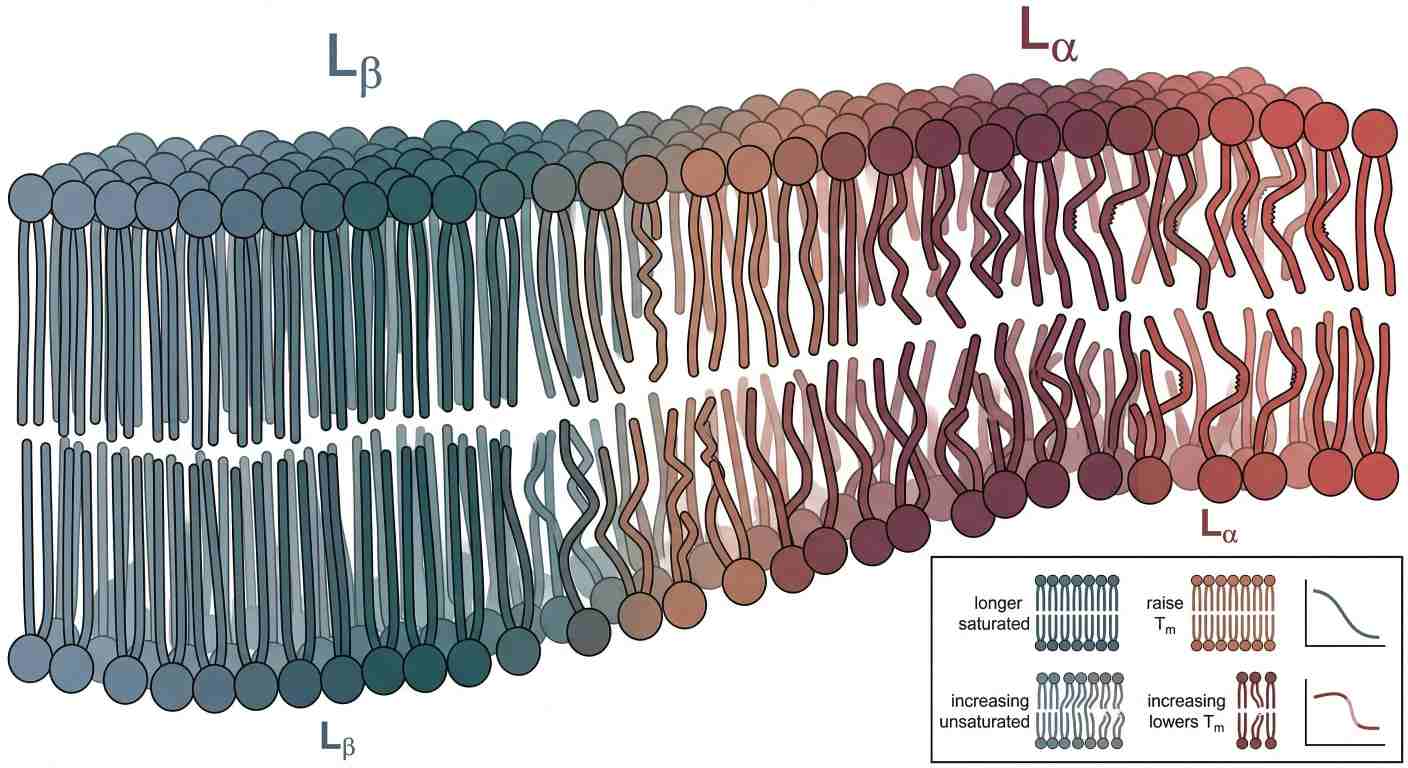 Figure 3: Chain Order Under Heat: The Gel-to-Fluid Transition in Phospholipid Bilayers
Figure 3: Chain Order Under Heat: The Gel-to-Fluid Transition in Phospholipid Bilayers
This is exactly where class abundance alone stops being enough. If the biological question concerns chain-dependent membrane-state changes, then species-level resolution becomes much more valuable than total phospholipid totals. In those cases, Targeted Lipidomics and Phospholipids Analysis Service are often better aligned with the question than a broad untargeted screen alone.
Packing Constraints and the Logic of Self-Assembly
The hydrophobic effect explains assembly, but not preferred geometry
Phospholipids do not remain dispersed in water because the hydrophobic effect drives them into aggregates. But that does not explain what aggregate they will form. For that, packing constraints matter.
A useful conceptual tool here is the critical packing parameter. Without turning it into a formula-first exercise, the logic is straightforward. Preferred assembly depends on the balance among hydrophobic volume, chain length, and effective head-group area. Cone-like amphiphiles tend toward micelles. More cylindrical lipids favor lamellar bilayers. Molecules with relatively smaller effective head-group area can support inverted curvature and non-lamellar organization such as the inverted hexagonal phase, (HII).
PE illustrates how head-group geometry changes membrane mechanics
This is why PE matters so much in membrane remodeling. Its smaller head group makes it more compatible with negative curvature than PC. In a flat bilayer, that preference is not fully expressed, so the membrane can store curvature stress. When the system enters a fusion intermediate, a budding neck, or another highly curved transition state, PE-rich composition can lower the energetic penalty relative to a PC-dominated system.
This is one of the most important upgrades from basic membrane biology to membrane biophysics. Lipids are not just present in membranes. They bias the energy landscape of membrane shape change.
Non-bilayer propensity matters even when the membrane remains lamellar
The biological point is not that cells constantly convert full membranes into hexagonal phases. The point is that lipids with non-bilayer propensity influence transition states, curvature stress, and the energetic cost of remodeling. They can stabilize intermediates, lower bending penalties, and alter how membrane-shaping proteins operate.
That makes curvature stress relevant to vesicle trafficking, organelle dynamics, membrane fusion, and protein insertion. Once again, phospholipid structure becomes predictive. A shift in head-group size, chain composition, or class abundance can change the energetic cost of a biological event.
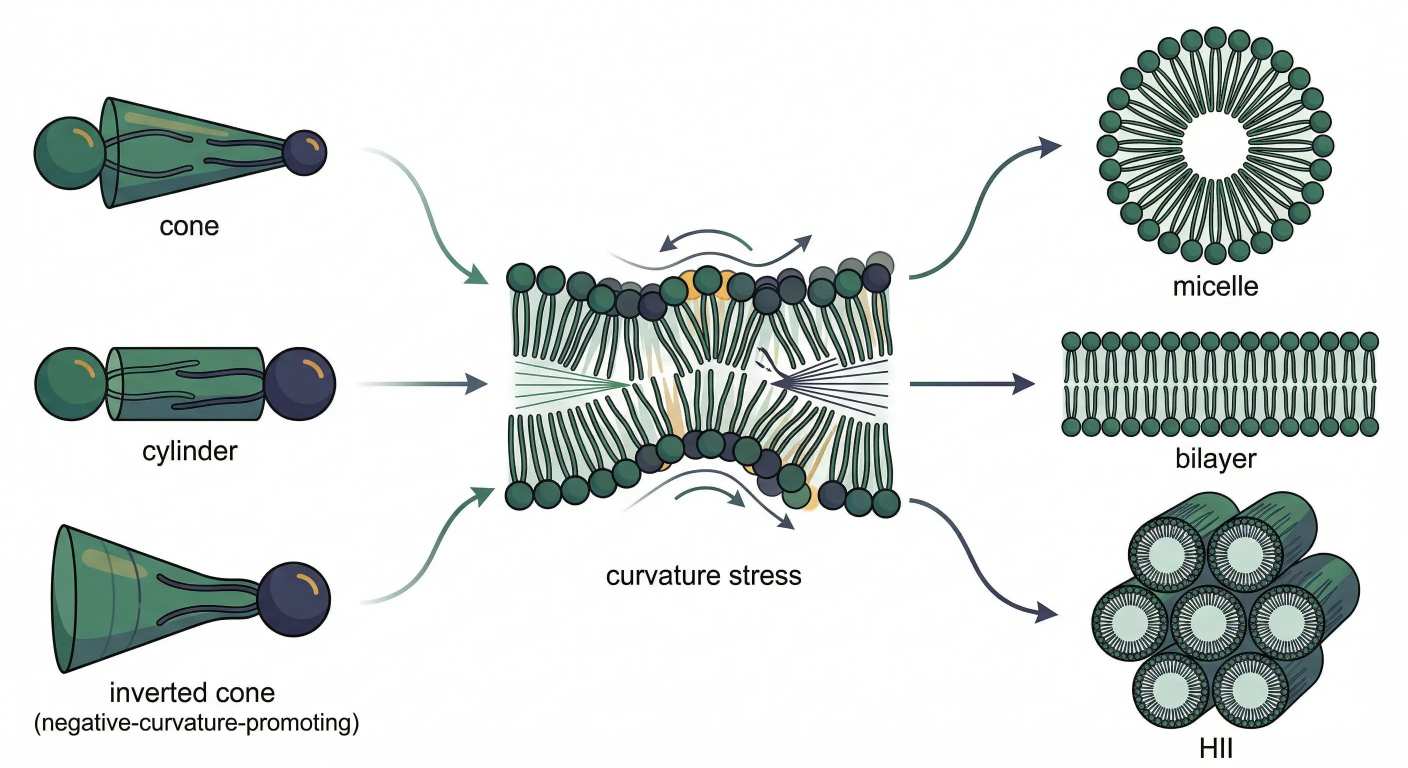 Figure 4: From Shape to Curvature: How Phospholipids Choose Micelles, Bilayers, or HII Phases
Figure 4: From Shape to Curvature: How Phospholipids Choose Micelles, Bilayers, or HII Phases
Cholesterol, Sphingomyelin, and Lateral Organization
Cholesterol is not a one-direction fluidity switch
Cholesterol is often described as a regulator of membrane fluidity. That statement is true, but too coarse. Cholesterol does not push membrane behavior in one fixed direction under all conditions. Its effect depends on the phospholipid background.
In highly ordered saturated systems, cholesterol can disrupt very tight chain packing and broaden otherwise sharp transitions. In more disordered unsaturated systems, it can condense chain motion and support a more ordered state. It is therefore better understood as a context-dependent packing modifier than as a universal rigidifier.
Sphingomyelin and cholesterol cooperate especially well
The relationship between cholesterol and sphingomyelin is especially important because sphingomyelin often carries relatively saturated chains and packs well with sterols. When sphingomyelin and cholesterol coexist within a broader background of more unsaturated glycerophospholipids, lateral heterogeneity can emerge.
That heterogeneity is not only compositional. It becomes physical. One membrane region may be more ordered, more tightly packed, and slightly thicker, while the surrounding membrane remains more disordered and more flexible. This is the basis of liquid-ordered membrane behavior.
Domain contrast matters because proteins do not experience membranes as uniform surfaces
The old raft model treated ordered domains too rigidly. Modern thinking is more careful. Lateral organization is dynamic, composition-sensitive, and strongly dependent on local thermodynamic conditions. But the central point remains: cholesterol–sphingomyelin cooperation can create a membrane state that differs from the surrounding phospholipid matrix.
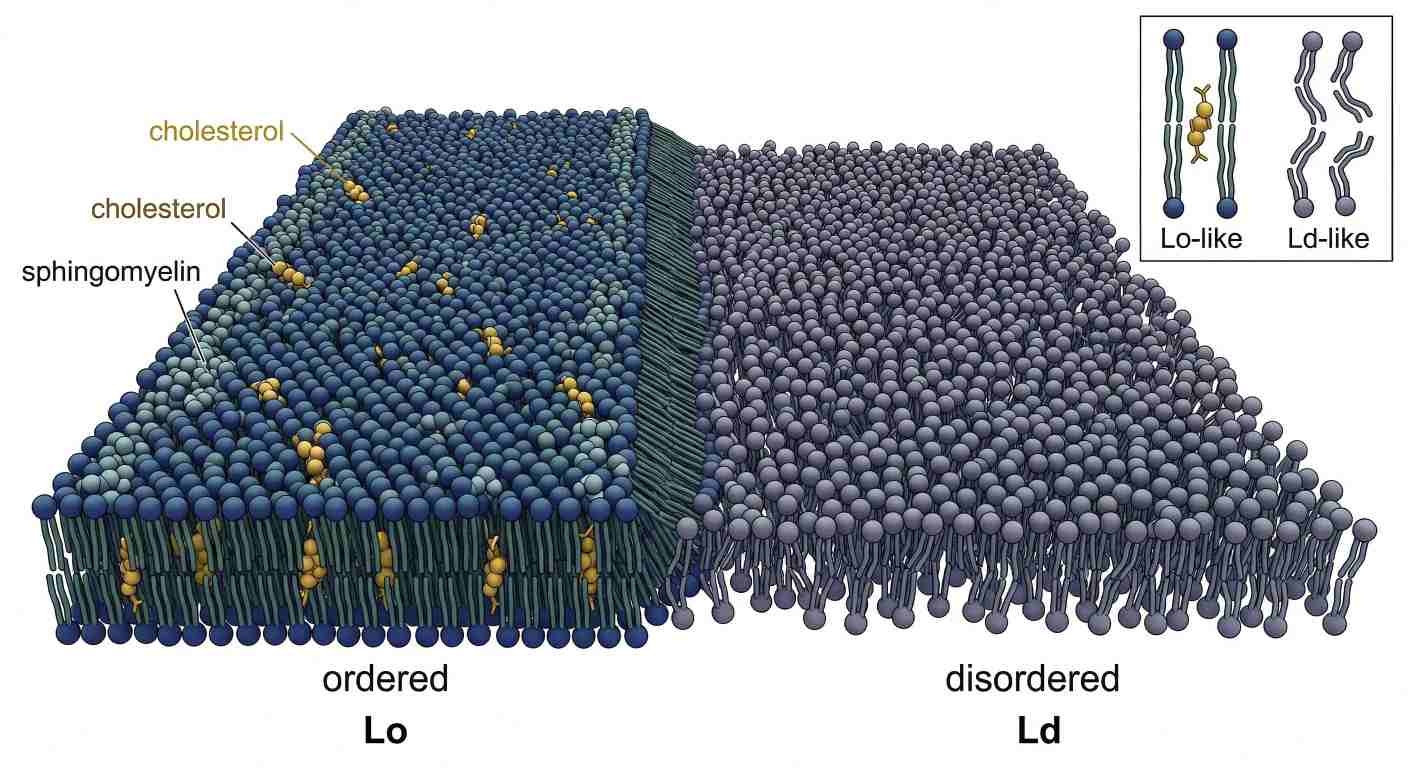 Figure 5: Order from Sterols: Cholesterol–Sphingomyelin Domain Formation in a Fluid Membrane
Figure 5: Order from Sterols: Cholesterol–Sphingomyelin Domain Formation in a Fluid Membrane
This matters because proteins interact with membrane states, not only with membrane surfaces. A more ordered, thicker, sterol-enriched region presents a different environment from a thinner, more unsaturated region. Membrane proteins, peripheral binders, and signaling complexes may partition differently as a result.
That is the key endpoint of the first half. Phospholipids are no longer just amphiphiles in a generic bilayer. They are structural variables that define interfacial chemistry, thermodynamic state, self-assembly logic, and lateral membrane organization. In the second half, that framework will carry directly into phospholipid signaling, phospholipase cleavage logic, and LC-MS/MS-based structural interpretation.
Phospholipids as Bioactive Signaling Molecules
Phospholipids do not merely support signaling. Many of them are the signaling substrate
Once membrane structure and membrane physics are in place, the next step is to understand why phospholipids matter so much in signaling. The key point is simple: phospholipids are not only structural components of membranes. They are also dynamic sources of information and bioactive intermediates.
Some phospholipids act by shaping the membrane environment in which receptors, channels, and enzymes operate. Others act more directly by serving as precursors for second messengers or lipid mediators. This dual role is what makes phospholipid biology so powerful. A single molecule can influence cell behavior in two ways at once: first by altering the physical properties of the membrane, and second by being enzymatically converted into an active signaling species.
That is why phospholipid signaling cannot be reduced to a short list of lipid classes. The important question is not only which phospholipid is present, but also where it is located, how fast it turns over, which enzyme can access it, and what product is generated after cleavage or phosphorylation.
The phosphoinositide cycle is a membrane-coded signaling system
Among all phospholipid families, phosphoinositides provide the clearest example of how low-abundance lipids can exert high regulatory control. Phosphatidylinositol itself is a membrane phospholipid, but its phosphorylated derivatives create a dynamic signaling code at membrane surfaces.
Phosphorylation can occur at different positions on the inositol ring, generating species such as PI4P, PI(4,5)P2, and PI(3,4,5)P3. These lipids are not abundant enough to dominate bulk membrane structure, but they do not need to be. Their power lies in local concentration, rapid turnover, and spatial precision. A small membrane patch enriched in PI(4,5)P2 can recruit a specific set of proteins, alter actin-linked behavior, regulate ion channels, or prime the site for enzymatic cleavage.
This is why phosphoinositides are best understood as membrane identity markers and signaling organizers. They help define where signaling should happen, which proteins should bind, and how quickly a membrane region can switch from one functional state to another. A membrane enriched in PI4P behaves differently from one enriched in PI(4,5)P2, even if the rest of the phospholipid background is similar.
The spatial aspect is critical. Phosphoinositide signaling is not mainly about large global concentration changes. It is about small, sharply controlled pools. Kinases and phosphatases act at membrane interfaces, and the kinetics of local synthesis and dephosphorylation determine whether a signaling platform is assembled, sustained, or erased. This gives membranes a form of chemical addressability. They are not just surfaces. They are spatially encoded reaction fields.
That is also why broad phospholipid profiling is not always enough. If the mechanistic question centers on low-abundance PI-derived signaling pools, class-level abundance alone will blur the answer. In those cases, Targeted Phosphoinositides Analysis Service is usually more informative than a general membrane lipid survey.
Phospholipases read phospholipid structure with site-specific logic
Phospholipases are not generic degradative enzymes. They are site-selective readers of phospholipid architecture. Each subtype cleaves a different bond and therefore produces a different signaling topology.
- Phospholipase A1 cleaves the sn-1 ester bond.
- Phospholipase A2 cleaves the sn-2 ester bond.
- Phospholipase C cleaves near the glycerol-phosphate region to produce diacylglycerol (DAG) and a phosphorylated soluble head-group product.
- Phospholipase D cleaves distal to the phosphate, generating phosphatidic acid (PA).
This cleavage logic is not a biochemical footnote. It explains why phospholipid structure has to be read positionally. If a bioactive polyunsaturated fatty acid sits at sn-2, PLA2 action releases it directly. If PLC acts on PI(4,5)P2, the outcome is not membrane destruction. It is the creation of two separate messenger streams: membrane-resident DAG and soluble IP3. If PLD acts, the product remains in the membrane as PA, where it can alter both signaling and curvature stress.
This site-specific logic turns phospholipid structure into an active signaling map. The same phospholipid class can support very different biological effects depending on which acyl chains are present, which position they occupy, and which enzyme family acts on them.
The sn-2 position is a built-in reservoir for lipid mediator biology
The sn-2 position deserves special emphasis because it often stores polyunsaturated fatty acids that later become signaling mediators. When arachidonic acid is esterified at sn-2, the phospholipid is effectively storing a future branch point. Once released, arachidonic acid can enter eicosanoid biosynthesis, feeding the production of prostaglandins, leukotrienes, and related lipid mediators.
This means that a phospholipid species is not only a membrane component with a certain packing behavior. It can also function as a pre-positioned precursor pool. The membrane is therefore not separate from inflammatory signaling. It is the substrate reservoir from which major mediator pathways can be launched.
From an analytical perspective, this has an important consequence. When a study observes remodeling in arachidonoyl-containing phospholipids, the interpretation should not stop at membrane composition. It should extend to downstream mediator potential. This is where Eicosanoids Analysis Service becomes a natural follow-up to phospholipid profiling.
Phospholipid signaling is always both chemical and physical
One of the most common interpretive mistakes is to separate membrane mechanics from signaling chemistry too sharply. In reality, they are tightly coupled.
A phospholipid can change signaling by being converted into a messenger. But it can also change signaling by altering surface charge, bilayer thickness, curvature stress, or local membrane order, which in turn changes how enzymes and proteins access the membrane. A PA-rich membrane is chemically different, but it is also mechanically different. A PI(4,5)P2-rich region recruits effectors chemically, but it also exists within a specific interfacial environment shaped by neighboring lipids, ions, and sterols.
That is why the deepest phospholipid studies do not ask only, "Which lipids changed?" They ask, "Which membrane state changed, which signaling axis changed, and how are those changes coupled?"
Advanced Lipidomic Analysis by LC-MS/MS
Why phospholipid analysis is harder than it first appears
Phospholipidomics looks straightforward from a distance. Extract the lipids, inject them into LC-MS/MS, identify the classes, and compare abundances. In practice, the problem is more difficult. Phospholipids exist as families of closely related molecular species whose masses can differ only slightly, whose ionization efficiencies are class-dependent, and whose fragmentation patterns contain both useful information and real ambiguity.
That means a phospholipid LC-MS/MS workflow has to solve several problems at once:
- detect diverse classes in a complex matrix
- preserve enough structural information to distinguish head-group identity from acyl composition
- reduce ion suppression
- manage isobaric and isomeric overlap
- annotate species with appropriate confidence
A phospholipid readout is therefore never just a peak list. It is the output of an interpretive pipeline.
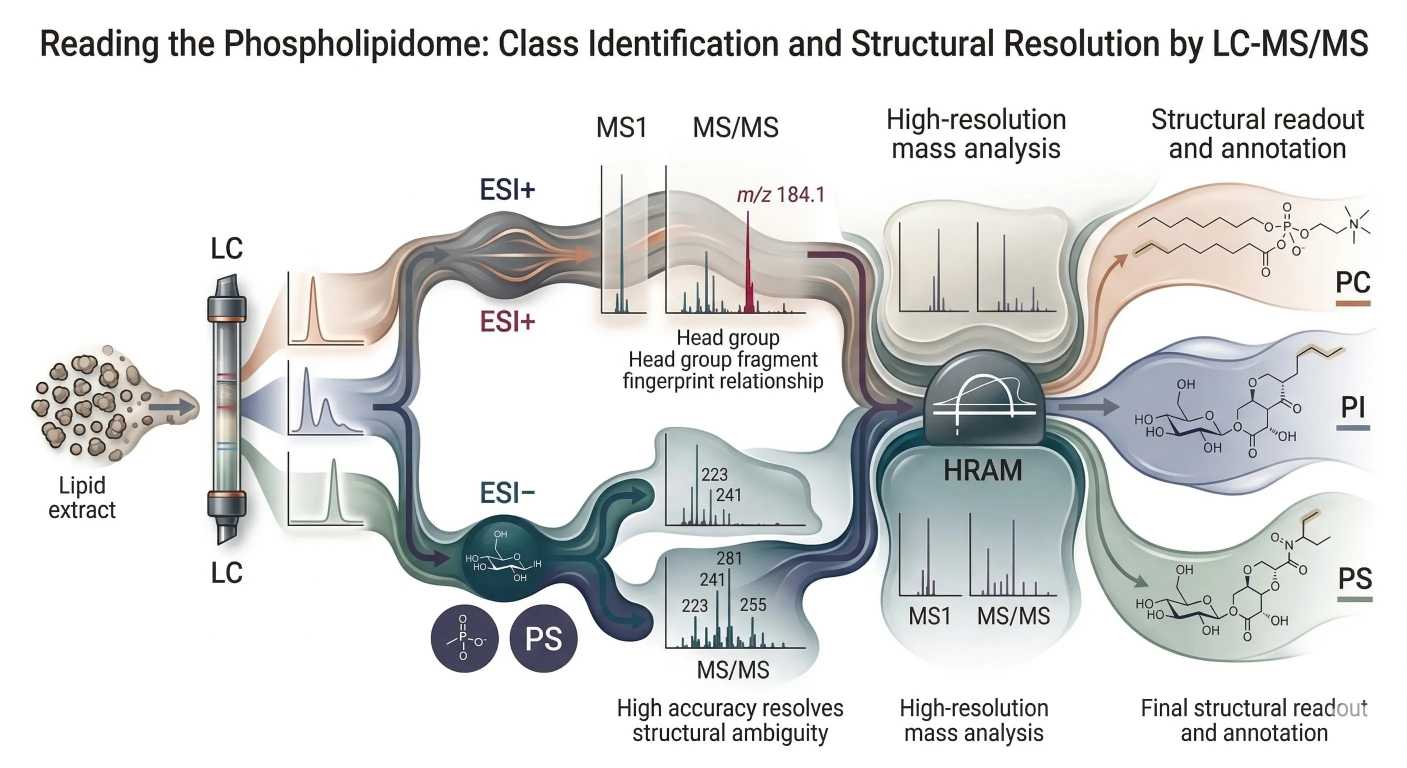 Figure 6: Reading the Phospholipidome: Class Identification and Structural Resolution by LC-MS/MS
Figure 6: Reading the Phospholipidome: Class Identification and Structural Resolution by LC-MS/MS
Ionization is class-dependent, so one mode never tells the whole story
The first analytical filter is ionization. In electrospray ionization, phospholipid classes do not behave equally in positive and negative mode. Their response depends on head-group chemistry and the stability of the ions they form.
Phosphatidylcholine (PC) is the classic positive-mode phospholipid. Its choline-containing head group favors strong signal in positive ESI, and PC often yields the characteristic phosphocholine-related fragment at m/z 184.1 in MS/MS. For phosphocholine-containing classes, positive mode is usually the fastest route to class recognition.
Sphingomyelin (SM) also benefits from positive-ion detection for the same broad reason: the phosphocholine-bearing interface strongly supports class-identifying behavior in positive mode. When the analytical priority is fast class assignment for phosphocholine-containing lipids, positive mode is usually the natural starting point.
By contrast, PI and PS often perform better in negative mode, where deprotonated ions are easier to form and class-informative fragmentation is often more interpretable. Negative mode is especially helpful when the goal is not just to detect the class, but to recover cleaner information about acidic head groups and associated acyl-chain features.
The practical lesson is straightforward: the best ion mode depends on what you are trying to learn. If the goal is rapid class identification for phosphocholine-containing species, positive mode is often favored. If the goal is stronger readout for acidic phospholipids or better chain-informative behavior in those classes, negative mode is often more useful. One mode rarely tells the whole story, and deep phospholipid work usually benefits from a class-aware acquisition strategy rather than a one-mode compromise.
MS/MS fragmentation works because phospholipids break in recognizable ways
Phospholipid MS/MS becomes readable when fragmentation is treated as structured logic, not random breakage.
At one level, fragmentation reveals the head group. This is where product ions and neutral losses become powerful. PC is the classic example because the phosphocholine fragment at m/z 184.1 provides a strong signature. Other classes are recognized through different fragment patterns, including characteristic neutral losses or class-specific head-group-derived ions.
At another level, fragmentation reveals acyl-chain information. Depending on ion mode and phospholipid class, MS/MS can generate signals corresponding to fatty-acid-related fragments or neutral losses that indicate which chains are attached. That is the bridge between "this is a phospholipid class" and "this is a specific molecular species."
The challenge is that these two layers of information are not always equally strong. Some fragmentation conditions produce very clear class identification but weaker chain detail. Others produce useful chain-level information but less straightforward class confirmation. That is why phospholipid MS/MS is best approached as a balance between class certainty and structural depth.
When the experiment requires stronger species-level resolution rather than broad discovery, Targeted Lipidomics and Phospholipids Analysis Service are often a better match than a single untargeted scan.
Determining sn-position is possible, but it is not a trivial readout
One of the most tempting goals in phospholipidomics is to assign sn-1 versus sn-2 chain placement directly from MS/MS. In some cases, this can be approached through fragmentation intensity patterns, because acyl loss from one position may be favored under specific class-dependent conditions. But this is suggestive logic, not universally definitive logic.
That boundary matters. Intensity asymmetry can support a plausible sn assignment, but it does not automatically prove it. The meaning of that asymmetry depends on the lipid class, adduct type, ion mode, and collision regime. A pattern that looks informative in one class may be far less reliable in another. Likewise, a spectrum collected under one fragmentation setting may not support the same confidence as a spectrum collected under another.
This is why high-confidence sn assignment often requires restraint. The biologically relevant question may be positional, especially when the sn-2 position carries the chain most relevant to membrane disorder or mediator release. But methodologically, analysts need to avoid overcalling. In many datasets, the correct statement is not "sn-1 and sn-2 are proven," but rather "the fragmentation pattern is consistent with a likely positional arrangement under these acquisition conditions."
The strongest practice is to treat sn-position interpretation as class-dependent and method-dependent. Where positional certainty is central to the study, orthogonal evidence or more specialized workflows may be needed. LC-MS/MS can strongly narrow the interpretation, but it does not always settle positional ambiguity on its own.
Shotgun lipidomics and LC-MS lipidomics answer different questions
A recurring method question is whether to use shotgun lipidomics or LC-MS-based lipidomics.
Shotgun lipidomics is fast and can profile many lipids directly from extracts with minimal chromatographic delay. That simplicity is appealing, especially for high-throughput discovery. But the tradeoff is that all analytes compete in the ion source together, which increases the risk of ion suppression and makes matrix complexity harder to manage.
LC-MS adds separation before ionization. That improves interpretability by spreading phospholipid classes and molecular species across retention time. In complex biological extracts, this often reduces overlap, supports cleaner fragmentation, and makes class-aware interpretation more robust. The cost is longer method time and greater method-development burden.
There is no universal winner. Shotgun methods are useful when throughput is critical and the question is broad. LC-MS becomes more attractive when class separation, species-level annotation, and control of matrix effects matter.
A practical workflow selection rule is often enough:
- Broad, rapid discovery across many lipid classes → shotgun can be efficient
- Complex extracts where separation and cleaner annotation matter → LC-MS is usually stronger
- Mechanistic follow-up on defined species or signaling axes → targeted follow-up is often the better endpoint
Ion suppression and matrix effects are not minor nuisances
Two terms appear in almost every phospholipidomics troubleshooting discussion: ion suppression and matrix effect. They are related, and in practice the key issue is the same. Complex extracts contain many co-eluting species, salts, and endogenous components that distort ionization efficiency. Measured signal therefore does not always scale cleanly with actual abundance.
This is one reason phospholipid data should never be interpreted as if every lipid class ionizes equally well. A strong signal may reflect favorable ionization as much as high abundance. A weak signal may reflect suppression instead of scarcity. Without internal standards and class-aware method design, these distortions can become severe.
That is why robust workflows often combine optimized extraction, careful chromatographic separation, and internal standards that match the lipid class under study. In larger studies, data quality also depends on downstream normalization and multivariate control, which is where Bioinformatic Data Preprocess and Normalization Service becomes part of the analytical pipeline rather than a post hoc cleanup step.
High-resolution accurate mass is essential when overlap becomes the real problem
As phospholipid mixtures become more complex, one of the hardest problems is not detection. It is overlap.
Different phospholipid species can sit so close in mass that low-resolution measurement struggles to distinguish them. In some cases, near-isobaric overlap can also arise between classes. A nominal mass alone may therefore be insufficient to assign a species correctly.
This is where high-resolution accurate mass (HRAM) becomes essential. Orbitrap-class instruments and related high-resolution platforms allow much finer mass discrimination, helping separate overlapping candidates and improve annotation confidence. That does not solve every ambiguity, especially where structural isomers are concerned, but it greatly reduces the risk of collapsing multiple species into one misleading call.
This matters because misassignment does not stay local. If the wrong class or species is called, the downstream interpretation of membrane state, signaling potential, and biological relevance can all be distorted.
Extraction method choice already shapes the lipidome you think you measured
Before LC-MS/MS begins, extraction has already started to shape the dataset. Different extraction methods do not recover all phospholipid classes equally, and they differ in solvent toxicity, phase handling, and matrix compatibility.
The three most commonly compared strategies are Folch, Bligh & Dyer, and MTBE-based extraction. Each has strengths and tradeoffs, and none is neutral.
Comparison Table: Folch vs Bligh & Dyer vs MTBE
| Method | Solvent System | Main Strength | Main Limitation | Phospholipid Recovery Tendency | Lipid Class Coverage | Practical Notes |
|---|---|---|---|---|---|---|
| Folch | Chloroform : methanol, typically 2:1 | Strong historical standard, robust total lipid extraction | Uses chlorinated solvent, more handling burden | Often strong for broad membrane lipid recovery | Broad | Reliable for many tissue-rich samples, but solvent safety and disposal matter |
| Bligh & Dyer | Chloroform : methanol : water in a different ratio scheme | Useful for lower-lipid samples and classic wet extraction workflows | Performance can vary more strongly with sample composition | Good, but matrix-dependent | Broad | Often considered for aqueous or lower-lipid matrices |
| MTBE | Methyl tert-butyl ether with methanol/water partitioning | Cleaner upper organic phase handling, reduced chloroform use | Recovery balance can differ by class and sample type | Often favorable in modern lipidomics workflows | Broad to very broad | Popular for workflow convenience and safer handling profile |
The most useful way to choose among them is not by tradition alone, but by sample type and analytical goal. For tissue-rich samples or broad membrane-lipid recovery, Folch remains a strong option. For lower-lipid aqueous matrices, Bligh & Dyer may still be considered. For many modern workflows where handling simplicity and reduced chlorinated solvent burden matter, MTBE is attractive.
The practical lesson is simple: there is no neutral extraction step. Extraction already biases what enters the phospholipidome you later interpret. That is why method selection is part of biological interpretation, not an invisible preprocessing step.
Conclusion
Phospholipids are often introduced as membrane constituents, but that framing is too small for the real subject. They are architectural molecules, thermodynamic regulators, signaling substrates, and analytical puzzles at the same time.
At the structural level, backbone class, head-group chemistry, and sn-position logic define how a phospholipid should be read. At the membrane level, those same variables determine phase behavior, interfacial charge, curvature stress, and lateral organization. At the signaling level, phosphoinositides and phospholipase products turn membrane composition into spatially controlled biochemical output. And at the analytical level, LC-MS/MS must recover all of that information from mixtures full of class-dependent ionization, fragmentation asymmetry, and overlap.
In practice, useful phospholipid interpretation starts when class identity, chain composition, membrane-state logic, and analytical constraints are evaluated together rather than in isolation.
FAQ
What is the difference between glycerophospholipids and sphingophospholipids?
Glycerophospholipids are built on a glycerol backbone with acyl chains typically at sn-1 and sn-2 and a phosphate-linked head group at sn-3. Sphingophospholipids use a sphingoid backbone, and one hydrophobic chain is part of that backbone while the second is attached through an amide bond. This difference influences packing, sterol interactions, and MS/MS behavior.
Why is the sn-2 position so important in phospholipids?
The sn-2 position often carries more unsaturated fatty acids, including biologically important polyunsaturated chains such as arachidonic acid. That makes it central to membrane disorder, signaling potential, and phospholipase A2-mediated release of lipid mediator precursors.
What determines the melting temperature (Tm) of a phospholipid bilayer?
The main structural drivers are acyl-chain length and unsaturation. Longer saturated chains increase (Tm), while cis double bonds lower it by disrupting tight packing.
Why does phosphatidylethanolamine promote curvature more strongly than phosphatidylcholine?
PE has a smaller head group and therefore a reduced effective interfacial area relative to its hydrophobic region. That geometry supports negative curvature more strongly than the more cylindrical PC structure.
Why is cholesterol not simply a membrane rigidifier?
Cholesterol acts in a composition-dependent way. In highly ordered saturated systems, it can disrupt very tight packing. In more disordered unsaturated systems, it can condense chain motion and support liquid-ordered behavior.
Why is phospholipid LC-MS/MS more difficult than basic lipid measurement?
Because phospholipid classes ionize differently, fragment differently, and frequently overlap in complex biological extracts. Accurate annotation requires class-aware ionization logic, informative fragmentation, and often high-resolution accurate mass.
Why is PC often measured in positive ion mode?
PC carries a phosphocholine head group that responds strongly in positive electrospray ionization and often yields the characteristic fragment at m/z 184.1, which helps confirm class identity.
Can LC-MS/MS alone always resolve phospholipid sn-position?
No. Positional assignment depends on lipid class, adduct type, ion mode, fragmentation regime, and the quality of supporting evidence. In some cases, MS/MS strongly suggests a likely arrangement, but it does not always prove it definitively.
References
- van Meer G, Voelker DR, Feigenson GW. Membrane lipids: where they are and how they behave. Nat Rev Mol Cell Biol. 2008;9(2):112-124. DOI: 10.1038/nrm2330
- Harayama T, Riezman H. Understanding the diversity of membrane lipid composition. Nat Rev Mol Cell Biol. 2018;19(5):281-296. DOI: 10.1038/nrm.2017.138
- Casares D, Escribá PV, Rosselló CA. Membrane Lipid Composition: Effect on Membrane and Organelle Structure, Function and Compartmentalization and Therapeutic Avenues. Int J Mol Sci. 2019;20(9):2167. DOI: 10.3390/ijms20092167
- Levental I, Lyman E. Regulation of membrane protein structure and function by their lipid nano-environment. Nat Rev Mol Cell Biol. 2023;24(2):107-122. DOI: 10.1038/s41580-022-00524-4
- Balla T, Hammond GRV, Irvine RF. Phosphoinositides as membrane organizers. Nat Rev Mol Cell Biol. 2022;23(12):797-816. DOI: 10.1038/s41580-022-00490-x
- Dennis EA, Norris PC. Eicosanoid storm in infection and inflammation. Nat Rev Immunol. 2015;15(8):511-523. DOI: 10.1038/nri3859
- Holčapek M, Liebisch G, Ekroos K. Lipidomic analysis. Anal Chem. 2018;90(7):4249-4257. DOI: 10.1021/acs.analchem.7b05395
- Hsu FF, Turk J. Electrospray ionization with low-energy collisionally activated dissociation tandem mass spectrometry of glycerophospholipids: mechanisms and applications. J Chromatogr B Analyt Technol Biomed Life Sci. 2009;877(26):2673-2695. DOI: 10.1016/j.jchromb.2009.02.033
- Pulfer M, Murphy RC. Electrospray mass spectrometry of phospholipids. Mass Spectrom Rev. 2003;22(5):332-364. DOI: 10.1002/mas.10061
- Cajka T, Fiehn O. Comprehensive analysis of lipids in biological systems by liquid chromatography-mass spectrometry. TrAC Trends Anal Chem. 2014;61:192-206. DOI: 10.1016/j.trac.2014.04.017
- Ulmer CZ, Jones CM, Yost RA, Garrett TJ, Bowden JA. Optimization of Folch, Bligh-Dyer, and Matyash sample-to-extraction solvent ratios for human plasma-based lipidomics studies. Anal Chim Acta. 2018;1037:351-357. DOI: 10.1016/j.aca.2018.07.004
- Klose C, Surma MA, Gerl MJ, et al. Flexibility of a eukaryotic lipidome—insights from yeast lipidomics. PLoS One. 2012;7(4):e35063. DOI: 10.1371/journal.pone.0035063
Disclaimer: For research use only. Not for use in diagnostic procedures.








