Ubiquitination, the process by which proteins are modified with ubiquitin, has important implications in agriculture and forestry. It plays a critical role in regulating plant growth, development, and stress responses. For example, ubiquitination has been shown to be involved in the degradation of proteins that are essential for the regulation of plant growth hormones. In addition, the manipulation of ubiquitination pathways has been used to enhance crop yield and stress tolerance. This has been achieved through the development of genetically modified crops that overexpress or downregulate specific ubiquitin-related genes. Furthermore, the use of ubiquitin-based proteomics has been employed to identify proteins involved in plant stress responses and to understand the molecular mechanisms underlying plant-microbe interactions. In forestry, the study of ubiquitination has implications for the management of forest ecosystems and the development of sustainable forestry practices. Overall, the study of ubiquitination modifications in agriculture and forestry has the potential to significantly impact crop yield, stress tolerance, and forest management practices.
Case 1 Ubiquitination modification profile of young rice spikelets (1)
To obtain information on ubiquitination modifications in young rice spikelets, researchers conducted a large-scale ubiquitination modification histology study and identified 1,638 ubiquitination modification sites on 916 proteins.
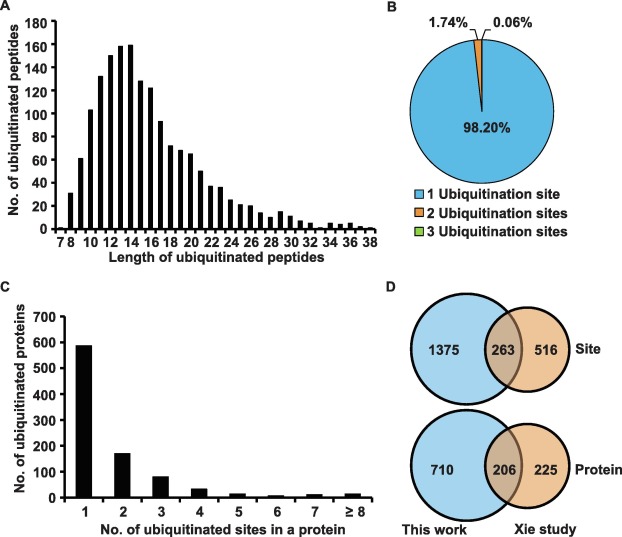
Characteristics of the identified lysine-ubiquitinated peptides in rice young panicles A
Functional analysis of the identified ubiquitinated modified proteins was carried out by GO and KEGG databases. The results showed that ubiquitination-modified proteins are involved in many essential cellular processes that are critical for anther development in young rice spikelets, such as protein degradation and DNA repair, revealing the role of protein ubiquitination in signal transduction of rice anther development. The PPI results suggest that ubiquitinated modified proteins are widely involved in different pathways, including general metabolism, protein degradation, RNA biology, DNA damage repair, plant-pathogen interaction, autophagy and innate immune signaling.
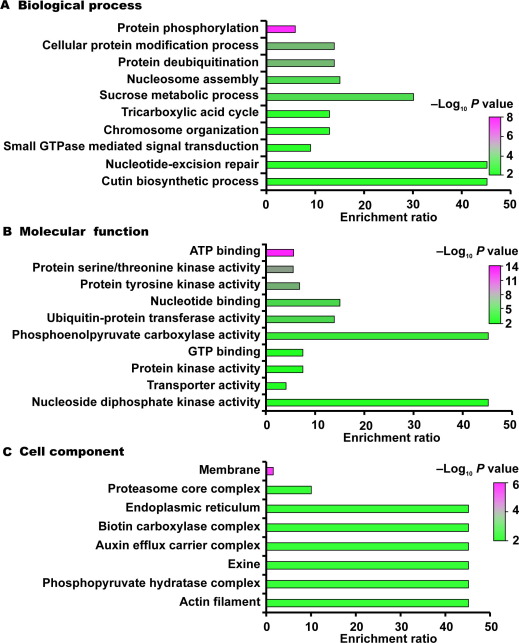
Enrichment analysis of GO annotations in identified ubiquitinated proteins
Excessive accumulation of ubiquitin causes male sterility in rice at the pollen mother cell and meiotic stages. To investigate the role of ubiquitination in rice anther development researchers identified seven proteins associated with anther development (MSP1, UAM3, WDA1, OsMADS7, NRL2, OsDRUS1, OsDRUS2) identified in this study. sRS5, HGW and qNPT1 were associated with rice seed size.
To further validate the protein ubiquitination modifications in young rice spikelets, Co-IP and WB experiments were performed on OsDRUS1 and OsOTUB1, and the results confirmed that OsDRUS1 and OsOTUB1 had ubiquitination modifications in young rice spikelets, which demonstrated the reliability of ubiquitination modification histological analysis.
Case 2 Ubiquitination modifications and abiotic stresses (2)
Among the many stress response pathways, ubiquitination can mediate the degradation of many transcription factors through the ubiquitin-proteasome pathway, which in turn is involved in stress response. In addition, plants accumulate misfolded proteins when subjected to stress, and ubiquitination can also mitigate plant injury by degrading these toxic proteins. Therefore, studying ubiquitination modifications in plant stress is biologically important and has great potential application in improving plant response to stressful environments.
First, the researchers used genetic analysis to identify Thermo-Tolerance1 (OgTT1), a target gene associated with heat tolerance, which encodes a protein that allows the proteasome in cells to degrade ubiquitinated substrates at a faster rate at high temperatures. To further investigate the mechanism by which OgTT1 controls high-temperature resistance in rice, researchers conducted quantitative ubiquitinylation histology studies in African and Asian rice. African rice was found to accumulate less ubiquitin-modified proteins after heat treatment, suggesting that African rice can more effectively eliminate the cytotoxicity of denatured proteins while maintaining the heat response process more efficiently. Using quantitative ubiquitinomic analysis, the results of all ubiquitin modification differences can be presented by bioinformatic analysis (Wayne diagram, heat map, etc.) as shown in the figure below, allowing for a more comprehensive and systematic observation of modification changes.
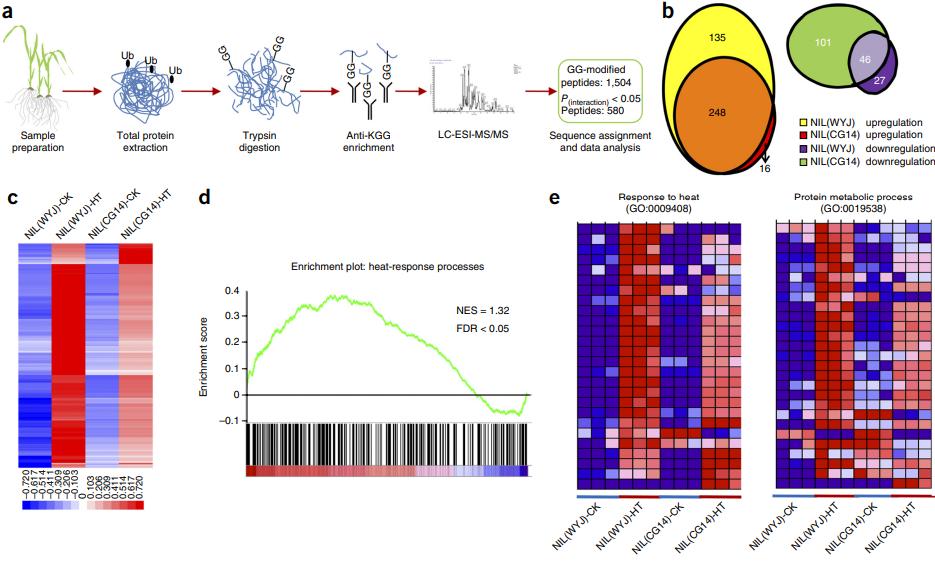
Role of TT1 in the heat response.
Reference
- Zhu, Liya, et al. "Ubiquitinome profiling reveals the landscape of ubiquitination regulation in rice young panicles." Genomics, Proteomics & Bioinformatics 18.3 (2020): 305-320.
- Li, Xin-Min, et al. "Natural alleles of a proteasome α2 subunit gene contribute to thermotolerance and adaptation of African rice." Nature genetics 47.7 (2015): 827-833.






