The Important Biological Functions at the N-terminus of Proteins and Peptides
Protein synthesis starts at the N-terminus, and the sequence composition of the N-terminus of the protein has an influence on the overall biological function of the protein. For example, the N-terminus sequence affects the half-life of the protein and is associated with the location of protein subcellular organelles. N-terminal sequencing analysis of proteins helps to analyze the high-level structure of proteins and reveal the biological functions of proteins. With the development of the modern pharmaceutical industry, a large number of protein and peptide drug molecules have appeared. The analysis and confirmation of the N-terminal sequence of these protein drug molecules is also an important link in the quality control of the pharmaceutical industry. In particular, the ICH Q6B guidelines require the provision of protein drugs. The N-terminal region is also an important structural and functional site of proteins and peptides, and most proteins can be identified by the few amino acid residues at the N-terminal. For example, the identification of artificial modification sites on the N-terminus of protein and peptide drugs, such as cyclization modification and methylation modification, which can lay the foundation for improving its degradation stability and prolonging the efficacy.
N-terminal Sequencing Technology for Proteins and Peptides
Edman degradation
- The principle of Edman degradation sequencing
Edman degradation is a very mature and classic method for N-terminal sequencing of proteins and peptides, and is widely used in the field of biotechnology. The principle of Edman degradation sequencing mainly refers to the identification of amino acid types one by one from the N-terminus of the protein through a cycle reaction, so as to determine the N-terminus sequence of the protein. The phenyl isothiocyanate (PITC) is reacted with the N-terminal amino group of the peptide to be analyzed under alkaline conditions to form a derivative of aniline thioformamide, and then the coupled product is treated with acid. The N-terminus of the peptide chain is selectively cleaved, releasing the thiazolinone aniline derivative of this amino acid residue. The extracted amino acid derivatives are converted into stable hydantoin thiourea amino acids (PTH-amino acids) under strong acidic conditions, and the degraded PTH-amino acid species can be analyzed using HPLC or electrophoresis to obtain proteins or peptides N-terminal sequence information.
- The advantages of Edman degradation sequencing
The Edman degradation method has been widely used as the gold standard for N-terminal sequence testing of existing protein samples. It is a valuable research tool for N-terminal sequence analysis of the entire purified protein and the most reliable sequencing method.
- The disadvantages of Edman degradation sequencing
Edman degradation method is subject to many restrictions, such as the protein or peptide used for sequence analysis must be of high purity, and is not suitable for high-throughput analysis, and the sensitivity is not enough.
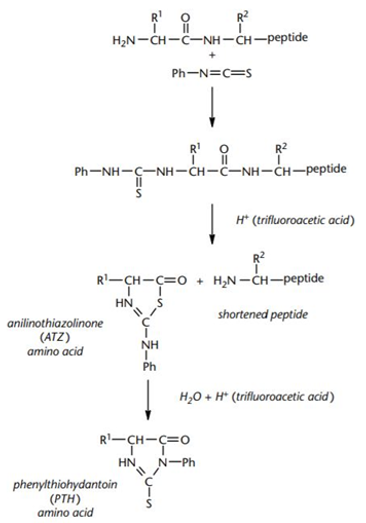
Edman chemistry for N-terminal sequencing of polypeptides (John Bryan Smith, 2011).
Mass spectrometry
Mass spectrometry-based protein N-terminal sequence sequencing technology can simultaneously determine the protein N-terminal sequence at a time, especially electrospray ionization (ESI) and matrix-assisted laserd desorption / ionization time-of-fight (MALDI-TOF), the application of mass spectrometry in protein structure analysis has undergone a revolutionary leap. High-sensitivity, high-accuracy, high-resolution, and high-throughput biological mass spectrometry technologies provide an important choice for protein N-terminal sequencing. The N-terminal sequencing technology based on mass spectrometry can realize the sequence determination of N-terminal blocking and PEGylated protein, which is complementary to Edman sequencing. This analysis can be used to confirm and identify the high-level structural integrity of recombinant protein drugs and the modification site of the N-terminal sequence of the recombinant protein. For this reason, N-terminal sequence sequencing can lay the foundation for the comparison of the sequence and modification of the original antibody.
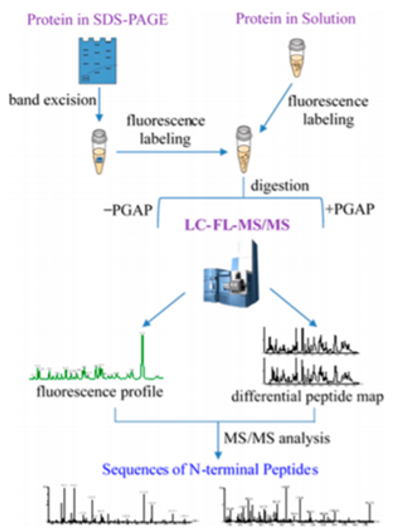
Workflow for identification and sequencing of N-terminus by mass spectrometry (Vecchi M M et al. 2019).
Chemical labeling combined with mass spectrometry
Many research methods for N-terminal peptides use a combination of mass spectrometry technology and a variety of chemical methods and biological enzymatic methods. For example, the protein is blocked by reduction, alkylation and guanidylation of side chain amino groups. The free N-terminal is labeled with different biotin reagents. After the labeled protein is digested with trypsin, the labeled N-terminal peptide is separated by the avidin affinity system, and then passed through MALDI-TOF / MALDI-TOF- PSD MS de novo sequencing to obtain the sequence of N-terminal peptide.
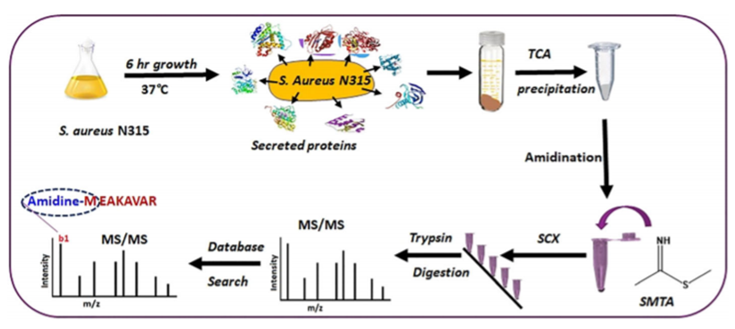
Chemical labeling / mass spectrometry work flow (Misal S A et al. 2019).
Future development of N-terminal sequencing
With the continuous development and improvement of classical methods, various chemical modifications based on mass spectrometry, and enzymatic assisted technologies, N-terminal sequence analysis of proteins and peptides has obtained rich terminal peptide sequence information, which provides a powerful basis and accelerate the identification of high-level structures of protein drugs and the study of modification sites. The N-terminal sequencing analysis technology of thousands of proteins in complex biological systems is still a huge challenge we face, especially for a more detailed determination of the N-terminal modification diversity on a large scale, a more targeted research strategy is also needed. The future development direction may be the exploration of chemical modification reagents and modification conditions with better selectivity and more controllable modification.
References
- John Bryan Smith. Peptide Sequencing by Edman Degradation. Encyclopedia of Life Sciences,2001.
- Vecchi M M, Xiao Y, et al. Identification and Sequencing of N-Terminal Peptides in Proteins by LC-Fluorescence-MS/MS: An Approach to Replacement of the Edman Degradation. Anal Chem, 2019, 91(21):13591-13600.
- Misal S A, Li S, et al. Identification of N-terminal protein processing sites by chemical labeling mass spectrometry. Rapid Commun Mass Spectrom, 2019;33(11):1015-1023.






