Proteins control all biological systems in the cell. Although many proteins function independently, most proteins need to interact with other proteins to maintain complete biological activity. Therefore, in order to better understand the function and biological characteristics of intracellular proteins, researchers often need to identify and analyze protein-protein interactions. Co-immunoprecipitation and pull-down are two of the most popular techniques for identifying protein-protein interactions because of their robust stability.
What is Co-immunoprecipitation?
- The principle and workflow of co-IP
Co-IP is a classic method for studying protein interactions based on the specific interaction between antibodies and antigens. It effectively determines the physiological interaction of two proteins in intact cells. During the co-IP assay, when cells are lysed under non-denaturing conditions, many protein-protein interactions are preserved in intact cells. When protein A is immunoprecipitated with an antibody of protein A pre-cured on argarose beads, protein B bound to protein A in vivo can be precipitated together. Protein B can then be detected by western blotting or mass spectrometry to prove protein-protein interaction.
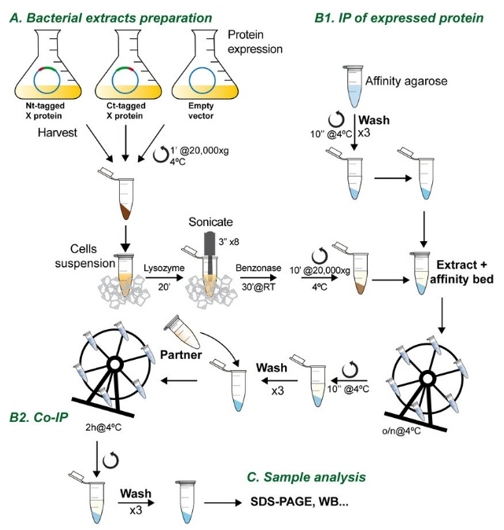
Flow chart of the co-immunoprecipitation procedure (Lechuga Ana et al, 2018).
- The advantages and disadvantages of co-IP
Co-IP can be used to detect both known protein-protein interactions in the body and to find unknown protein-protein interactions. The co-IP method can isolate and interact with protein complexes in a natural state, and the obtained protein has high reliability. This method also has certain limitations. One is that the binding of two proteins may not be direct, but a third party may play a bridge role. The other is that low affinity and transient protein-protein interactions may not be detected. Third, it is necessary to predict the target protein before the experiment in order to select the antibody to be detected. Therefore, if the prediction is incorrect, the experiment will not yield results.
What is Pull-down?
- The principle and workflow of pull-down
Pull-down assay is a form of affinity purification that are very similar to immunoprecipitation, except that bait proteins are used instead of antibodies. In a pull-down assay, a tagged bait protein is captured by a solid-phase affinity ligand that specifically binds to that tag. Commonly used tag proteins are GST-, His-, or Biotin-, producing a "secondary affinity support". Secondary affinity supports containing solid-phase decoy proteins can be incubated with various protein samples containing putative prey proteins. If the buffer and sample conditions are compatible with the target binding interaction and the bait protein can still function when tagged and immobilized, then the prey protein present in the sample will bind to the affinity support. If the affinity of the specific binding interaction is sufficiently strong, unbound sample components can be washed away, and the prey or bait-prey complex in purified form can be eluted from the support.
- The advantages and disadvantages of pull-down
The pull-down technology can determine the interaction between known proteins and fished proteins or purified related proteins, and can detect protein interactions from in vitro transcription or translation systems, such as the detection of proteins between yeast two-hybrid systems. It does not truly reflect the interaction between proteins, because they do not necessarily meet in space in the body. So it does not mean that they must be bound under physiological conditions.
- The types of pull-down
In order to use the pull-down technology more effectively, the protein to be purified can be expressed as a fusion protein, that is, a "bait" protein is fused with an easily purified ligand protein. According to different types of fusion proteins (common fusion tags), there are three types of pull-down methods: GST pull-down, polyhistidine pull-down, and biotin pull-down.
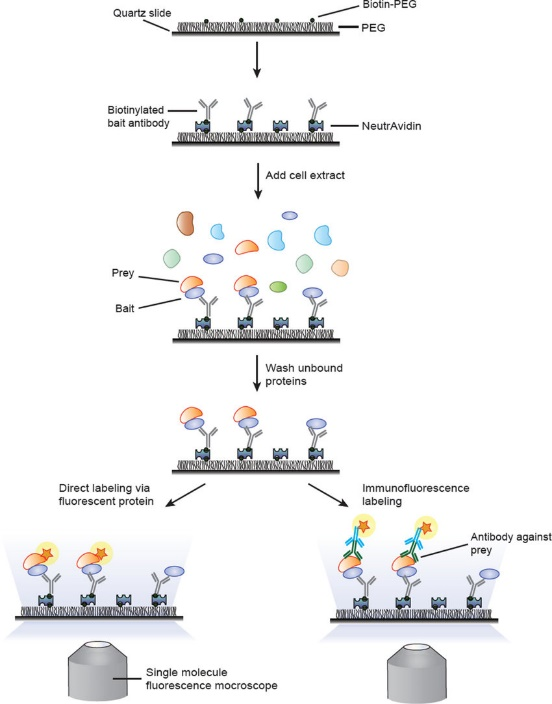
This protocol describes a single-molecule pull-down (SiMPull) assay for analyzing physiological protein complexes (Jain et al, 2012).
- GST pull-down assay
GST fusion protein can be adsorbed by the interaction of GST and GSH when passing through a glutathione (GSH) -immobilized column. When a cell extract is passed through the column, a protein of interest can be obtained to interact with the "bait" protein. The GST fusion protein pull-down method is used in two ways. One is to identify unknown proteins that can interact with known fusion proteins; the other is to identify whether there is an interaction between two known proteins. This method is relatively simple, avoids the use of isotopes and other dangerous substances, and has a wide range of applications in protein interaction studies.
- Polyhistidine pull-down assay
The basic technical principle of polyhistidine pull-down technology is the same as that of GST pull-down assay, except that the bait protein in this method is multihistamine acid.
- Biotin pull-down assay
Biotin pull-down assay is also an important branch of pull-down technology. The basic principles and steps of the assay are the same as GST pull-down assay, except that the bait protein in biotin pull-down assay is composed of biotin.
References
- Zhenyuan Tang, Yoshinori Takahashi. Analysis of Protein–Protein Interaction by Co-IP in Human Cells. Two-Hybrid Systems, 2018.
- S. Fletcher, S.E.H. Bowden, N.V. Marrion. False interaction of syntaxin 1A with a Ca2+-activated K+ channel revealed by co-immunoprecipitation and pull-down assays: implications for identification of protein–protein interactions. Neuropharmacology, 2003, 44(6):0-827.
- Lechuga Ana, Berjon-Otero M, Salas Margarita, et al. Analysis of Direct Interaction between Viral DNA-binding Proteins by Protein Pull-down Co-immunoprecipitation Assay, Bio-Protocol, 2018.
- Jain Ankur, Liu Ruijie, Xiang Yang, et al. Single-molecule pull-down for studying protein interactions. Nature protocols. 2012(7): 445-452.
- Lee, Hong-Won, Ryu, Ji Young, et al. Real-time single-molecule coimmunoprecipitation of weak protein-protein interactions. Nature Protocol, 2014, 8(10):2045-2060.






