- Service Details
- Case Study
What is the Gene Ontology (GO)?
The Gene Ontology Consortium has established the GO, a globally recognized and comprehensive database. It aims to be applicable to various species, providing precise definitions and descriptions of the functions of genes and genes products. And its contents can be continuously updated as research progresses, both in terms of quantity and quality. The GO current (2023-07-27) developed formal ontologies: 42,887 GO terms | 7,608,910 annotations 1,527,452 gene products | 5,319 species, is reviewed and updated almost every day. Most importantly, any biological data sources resulting in gene/protein lists can be retrieved from the GO for function annotation analysis. And the GO annotations of genes can be searched on Uniprot as well.
The most fundamental concept of the GO is the GO term, and a standard GO annotation is created by associating a gene product with a GO term. These terms are used to describe the characteristics of genes and gene products so that researchers can quickly find the target gene through the "term". For instance, in terms of genes, after microarray, ChIP-seq, transcription analysis, we obtain a large amount of data. Subsequently, we identify numerous up-regulated and down-regulated differential genes; however, the specific functions of these genes remain unknown. Nevertheless, since each gene possesses its own GO annotation. By searching for the GO annotations of these differential expressed genes, we can elucidate their respective functions. The biological interpretation of high-throughput gene or protein lists derived from omics or multi-omics experiment poses a challenging task. Therefore, it is widely adopted in the life sciences to help experimentalists as well as computational biologists working on biological interpretation of omics data.
The GO annotations of each gene product have three different domains:
1. Molecular Function (MF)
It is employed to elucidate the molecular-level functionalities of gene products, such as enzyme activity, DNA binding, receptor activity, etc. (Single step tasks).
2. Cellular Component (CC)
It is used to depict the cellular localization or constituents of gene products, such as the nucleus, cell membrane, mitochondria, etc. (Where do gene products localize and become active?)
3. Biological Process (BP)
It is used to describe the activities of gene products in biological processes, such as cell division, signal transduction, metabolic processes, etc. A recognized series of events. (How are gene products activated?)
For instance, the gene product of "NADPH-dependent methylglyoxal reductase GRE2" can be characterized by the Molecular Function "3-beta-hydroxy-delta5-steroid dehydrogenase activity", "methylglyoxal reductase (NADPH-dependent) activity", "3-methylbutanol: NAD oxidoreductase activity", and "3-methylbutanol: NADP oxidoreductase activity". The Cellular Component shows that it is localized in both the "cytoplasm" and "nucleus", and the Biological Process tell us it involves in biological processes such as "filamentous growth", "ergosterol metabolic process", and "steroid biosynthetic process".
Our Go annotation analysis service
As one of the leading omics CRO companies in the world, Creative Proteomics has over ten years of experience in the field of bioinformatics in proteomics, enabling us to provide our customers with outstanding, credible, and reliable Go annotation service!
Our specialized bioinformatics analysis specialists are proficient in R, Python, Shell, and Perl. We can effortlessly handle both whole genome and differentially expressed proteins for GO functional annotation, enabling us to classify gene product function and location with ease. The GO annotations to gene products have become an indispensable component of functional analysis, with the inclusion of statistical tests using GO data becoming a standard practice for researchers when disseminating functional information.
1) Annotate genes and gene products, and present data through tables and figures for visualization purposes.
2) Maintain and develop its controlled vocabulary of gene and gene product attributes.
3) Refine and distribute GO annotation data.
4) Provision of tools for facilitating convenient access to all facets of the data enables functional interpretation of experimental data.
5) For new species or unknown genes and gene products, we can conduct GO annotation by incorporating experimental results, sequence similarity, and curator judgment.
GO Analysis Service at Creative Proteomics can be used for the following:
1) Integration of proteomic data across different species.
2) Categorize the differential proteins.
3) Predict the functions of specific protein domains accurately.
4) Identify the specific genes implicated in certain diseases.
5) The integration of GO annotation and enrichment analysis can enhance the comprehension of datasets and streamline downstream verification experiments.
In all, the utilization of GO functional annotation analysis in our Creative Proteomics enables all of researchers to gain comprehensive insights into the collective biological significance of gene sets.
How to place an order
Creative Proteomics possesses the professional expertise and proficiency to provide a customized experimental scheme that aligns precisely with your specific requirements and projects. Experienced bioinformatics team with strict and skillful techniques, our service is high-quality for reliable results. Please feel free to contact us via email (info@creative-proteomics.com) for a comprehensive discussion regarding your specific requirements. Our customer service representatives are available round the clock, seven days a week.

Proteomics Analysis of Cellular Proteins Co-Immunoprecipitated with Nucleoprotein of Influenza A Virus (H7N9)
Journal: International Journal of Molecular Sciences
Published: 2015
Main Technology: Co-immunoprecipitation (Co-IP), Proteomics, Bioinformatics Analysis (GO annotation analysis)
Abstract
Avian influenza A viruses are serious veterinary pathogens that normally circulate among avian populations, causing substantial economic impacts. Some strains of avian influenza A viruses, such as H5N1, H9N2, and recently reported H7N9, have been occasionally found to adapt to humans from other species. In order to replicate efficiently in the new host, influenza viruses have to interact with a variety of host factors. In the present study, H7N9 nucleoprotein was transfected into human HEK293T cells, followed by immunoprecipitated and analyzed by proteomics approaches. A series of host proteins co-immunoprecipitated were identified with high confidence, some of which were found to be acetylated at their lysine residues. Bioinformatics analysis revealed that spliceosome might be the most relevant pathway involved in host response to nucleoprotein expression, increasing our emerging knowledge of host proteins that might be involved in influenza virus replication activities.
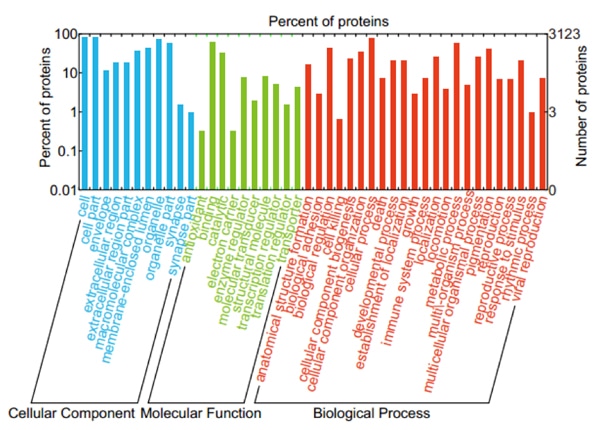
 Figure 3. GO annotation of identified NP-related proteins in three categories: biological process (BP), cellular component (CC) and molecular function (MF).
Figure 3. GO annotation of identified NP-related proteins in three categories: biological process (BP), cellular component (CC) and molecular function (MF).




