What is Edman Degradation Sequencing?
Edman degradation sequencing is to sequentially cut out each amino acid at the N-terminal of the protein through chemical reaction, and then use HPLC liquid chromatography to identify the ID of the amino acid, and the obtained amino acid sequence information is the sequence of the N-terminal of the protein. Next, we use the determination of the first amino acid at the N-terminus of a protein as an example to explain the entire process of a single Edman sequencing reaction.
Determination of the First Amino Acid at the N-terminal of Protein by Edman Degradation Method
First, the α-amino acid at the N-terminal end of the protein is reacted with PITC, allowing PITC to be attached to the N-terminal end of the protein. The PITC-protein complex is then treated with trichloroacetic acid (TFA), under the action of which the PITC and the first amino acid attached to it can be specifically cleaved off to form a PITC-amino acid complex. The PITC-amino acid is then extracted and isolated. The PITC-amino acid is then converted to the more stable PTH-amino acid form, which is identified by HPLC chromatography and compared with the standard amino acid chromatogram to obtain information on the first amino acid. Similarly, the remaining amino acid sequences are sequentially determined in this way to obtain information on the amino acid sequence at the N-terminal end of the protein.
When to use Edman sequencing?
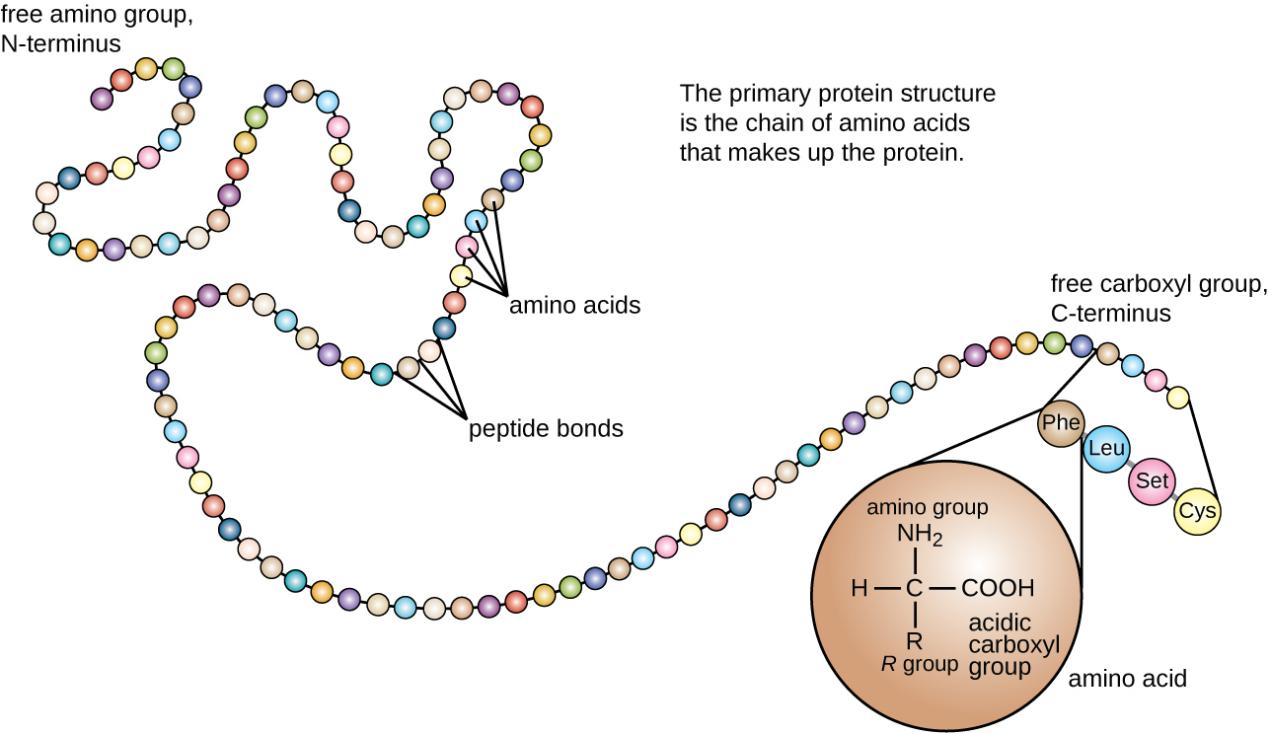
According to the principle of Edman sequencing, we can know that the starting point of Edman sequencing is the free α-amino group at the N-terminal of the protein, and then the amino acid sequence is sequentially determined from the N-terminal to the C. Therefore, Edman sequencing can be applied to the N-terminal sequence determination of various proteins, including recombinant proteins, protein isoforms, antibodies, peptides, etc.
Advantages of Edman sequencing over traditional mass spectrometry
Edman sequencing can analyze amino acid information directly without the help of any protein database, therefore, Edman sequencing can analyze the N-terminal sequence of brand new proteins without database information. In contrast, the traditional mass spectrometry sequencing method requires the use of a known protein database to compare the determined polypeptide sequence with the protein database, and then infer the protein ID and sequence. Therefore, traditional mass spectrometry cannot sequence proteins that are not contained in the database. Also, mass spectrometry is not able to accurately distinguish between amino acids with similar molecular weights (I/L, Q/K) and mutation sites on proteins. Edman sequencing directly determines the sequence of N-terminal mutated/modified proteins more accurately. This is why Edman sequencing is still known as the gold standard for N-terminal sequence analysis of proteins.
Limitations of Edman sequencing
Edman sequencing has several major limitations: the throughput is low and it is not possible to analyze multiple proteins simultaneously; the analysis process is also relatively time-consuming, taking 45 minutes per residue, and the cost is high. In addition, because Edman sequencing starts from the free α-amino group at the N-terminal end of the protein, it is not possible to directly sequence the N-terminal end of proteins with blocked N-terminal (α-amino groups with modifications such as acetylation, methylation and pyroglutamylation).
Since Edman generally determines the sequence of about 50 amino acids at the N-terminal end of a protein, it has some limitations for analyzing the full-length sequence of a protein. However, it is possible to analyze the full-length protein sequence by using Edman sequencing method. It is only need to digest the protein into several short peptides, and then analyze the sequence of each short peptide by Edman sequencing method to splice into the full-length protein sequence.
How to address the limitations of Edman sequencing?
In response to the problem of throughput, we can combine the advantages of mass spectrometry identification and Edman sequencing to achieve high-throughput protein identification and precise sequence determination. In addition, for the case of N-terminal blocking, the corresponding enzyme can be selected to remove the modification of the N-terminal of the protein, or the method of mass spectrometry identification can be used to identify the modification of the N-terminal. In the process of N-terminal sequencing of protein samples, Edman sequencing and mass spectrometry sequencing can complement each other and learn from each other.






