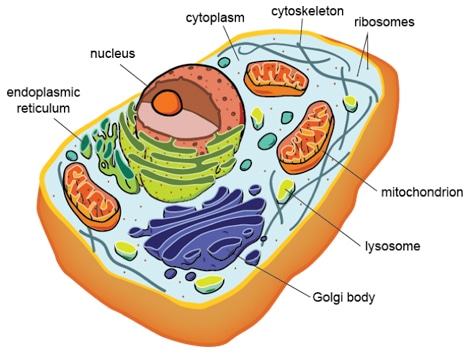
The cells of eukaryotic organisms are elaborately divided into functionally different membrane bound compartments. Some major components of eukaryotic cells are: extracellular space, mitochondria, nucleus, cytoplasm, Golgi apparatus, nucleoplasm, peroxisome, cytoskeleton, nucleolus, vacuoles, endoplasmic reticulum (ER), ribosomes and nuclear matrix.
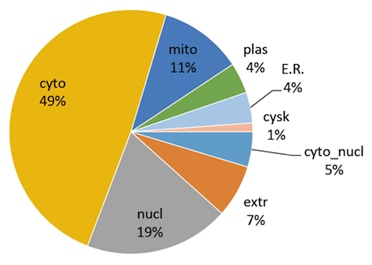
Bacteria also have protein subcellular localizations that can be disconnected when the cell is fractionated. The most generic localizations referred to include the cytoplasm, the cell wall (which is generally thicker in Gram-positive bacteria), the cytoplasmic membrane (also referred to as the inner membrane of Gram-negative bacteria) and the extracellular environment. Most Gram-negative bacteria also have an outer membrane and periplasmic space. Compared to eukaryotes, most bacteria do not have membrane-bound organelles, however there are some exceptions. Protein subcellular localization is a significant part of bioinformatics based prediction of protein function and genome annotation, and it can aid the identification of drug targets.
The reasons why we recommend Subcellular Localization include:
- Identifying subcellular localization is very important for understanding protein function and is a vital step in genome annotation.
- Abnormal subcellular localization of proteins has been discovered in the cells of a variety of diseases, such as Alzheimer's disease and cancer.
- Knowledge of the subcellular localization of a protein can greatly improve target identification in the course of drug discovery. For instance, plasma membrane proteins and secreted proteins are easily accessible by drug molecules due to they are located in the extracellular space or on the cell surface.
- Bacterial secreted proteins and cell surface are also of interest for their potential as diagnostic targets or as vaccine candidates.
- Some secreted proteins from archaea that can survive in severe environments have industrially important applications.
How to place an order:

*If your organization requires signing of a confidentiality agreement, please contact us by email
As one of the leading omics industry company in the world! Creative Proteomics now is opening to provide Subcellular Localization service for our customers. With rich experience in the field of bioinformatics, we are willing to provide our customer the most outstanding service! Contact us for all the detailed informations!




