Significance analysis of microarrays (SAM) is a widely used statistical technique, established in 2001 by Virginia Tusher, Robert Tibshirani and Gilbert Chu, for determining statistically significant changes in gene expression. With the advent of DNA microarrays, it is now possible to measure the expression of up to thousands of genes in a single hybridization experiment. The data generated is enormous, and a method for sorting out what is significant and what isn't is critical.
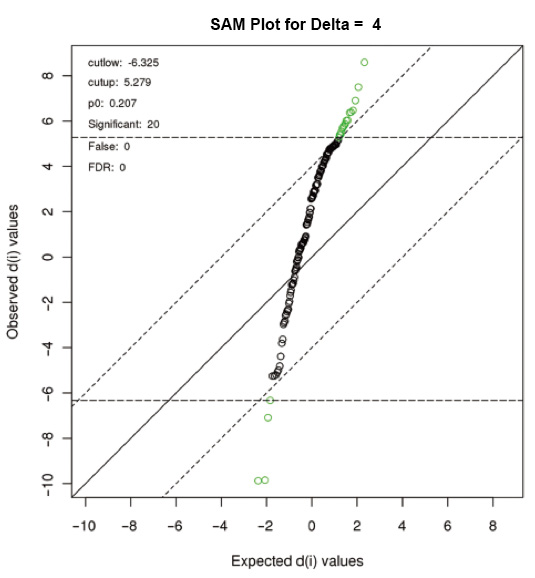
SAM identifies statistically significant genes by carrying out a set of gene specific t-tests and computes a statistic dj for each gene j, which measures the strength of the relationship between gene expression and a response variable. Each gene is assigned a score on the basis of its change in gene expression relative to the standard deviation of repeated measurements for that gene. A gene with scores higher than the threshold is considered to be potentially significant. The posibility of these genes being identified by chance is the false discovery rate (FDR). In order to estimate the FDR, analyzing those permutations from the measurement identifies of the nonsense genes. The threshold can be adjusted to identify smaller or larger collections of genes, and FDRs are computed for each collection.
Non-parametric statistics was used in this analysis, since the data may not follow a normal distribution. The response variable distinguishes and groups the data based on experimental conditions. In this method, repeated permutations of the data are applied to determine if the expression of any gene is statistically significant related to the response. The use of permutation-based analysis accounts for correlations in genes and avoids parametric assumptions about the distribution of each gene. This is an advantage over other techniques (e.g., ANOVA and Bonferroni), which assume equal variance or independence of genes.
How to place an order:

*If your organization requires signing of a confidentiality agreement, please contact us by email




