Proteins lipidation refers to the covalent binding of lipid molecules to proteins, thereby revising the structure and function of proteins. Among the known covalent chemical modifications, the unique physicochemical properties of lipid molecules endow proteins with special structures and functions as well as greatly affect their membrane anchoring ability, transport and localization pathways, signal transduction, and protein interactions. The diversity of lipid structures and binding modes determines the functional complexity of lipidated proteins. Meanwhile, these lipidated proteins act in concert with other known or unknown modified proteins for the fine regulation of physiological activities. The structural and functional analysis of protein lipidation modifications is an important way to reveal the mechanisms of microbial infection, immune regulation, and tumor development and is important for discovering and screening disease markers and mining novel drug targets.
Existing analytical techniques have detected a very small fraction of lipids bound to proteins. Fatty acids, isopentenyl lipids, cholesterol, and glycosylphosphatidylinositol (GPI) are four common types of lipid molecules that can be covalently bound to proteins. The first three are lipid groups, and the last one is a glycolipid complex. Nevertheless, these four lipid molecules are covalently bound to proteins mainly by the N-, S-, and O- linkages.
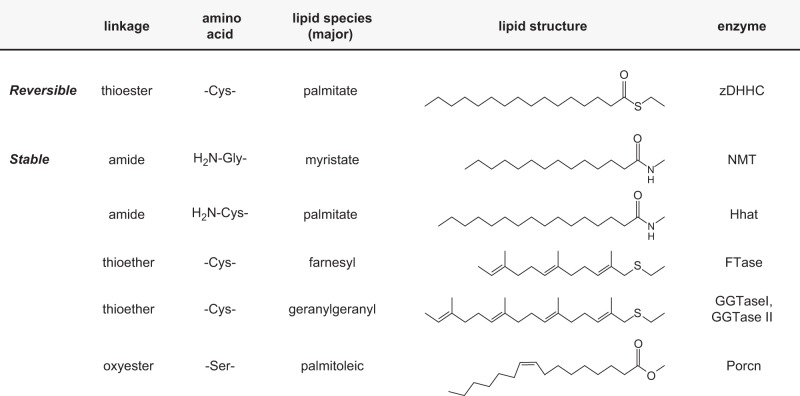
Major lipid modifications of proteins
Major Analytical Methods of Protein Lipidation
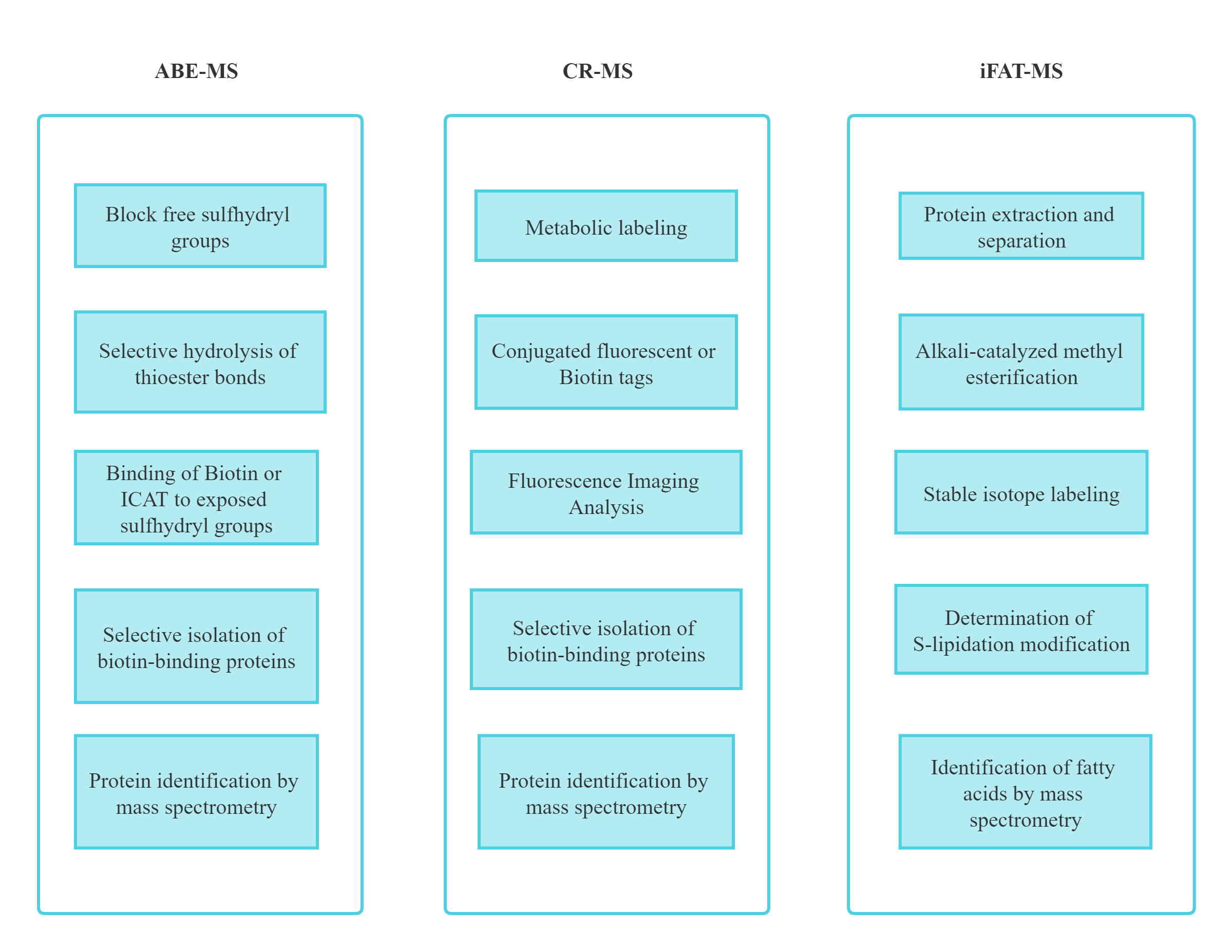
Our Protein Lipidation Analysis Service
Creative Proteomics uses Thermo Fisher's Q ExactiveHF Mass Spectrometry platform, Orbitrap Fusion Mass Spectrometry platform, and Orbitrap Fusion Lumos Mass Spectrometry platform in combination with Nano-LC to provide protein lipidation identification services.
Customized Bioinformatics Services
| Standard Analysis | Customized Analysis |
|---|---|
a) Collection of output statistics | Modified peptide motif analysis |
Sample requirements
- Fresh animal tissue: ≥600 mg
- Fresh plant tissue: ≥6 g
- Cell culture: ≥1×107 cells/tube x 3 tubes
- Fungi and bacteria: ≥600 mg
- Serum, plasma: 450 μL × 4 tubes
- Protein solution: total protein of 5-10 mg
- Body fluid samples: urine of 15 mL × 4 tubes (centrifuge at 1000 x g for 5 minutes and discard sediment); or other body fluids (saliva, amniotic fluid, cell culture supernatant, etc.) > 15 mL
Advantages
- High specificity and enrichment efficiency
- Large-scale identification of lipidated protein with mass spectrometry of high resolution and high throughput
- Combining commercially available quantitative techniques to analyze, compare, and correlate differences in lipidation levels between different samples quantitatively
- A research team with extensive experience in PTMs
References
- Luke H. Chamberlain, et al. The Physiology of Protein S-acylation. Physiol Rev. 2015 Apr; 95(2): 341–376.
- PinJoe Ko, et al. Protein palmitoylation and cancer. EMBO Rep. 2018 Oct; 19(10): e46666.
- Benjamin R. Sabari, et al. Metabolic regulation of gene expression through histone acylations. Nat Rev Mol Cell Biol. 2017 Feb; 18(2): 90–101.
- Baoen Chen, et al. Protein lipidation in cell signaling and diseases: function, regulation and therapeutic opportunities. Cell Chem Biol. 2018 Jul 19; 25(7): 817–831.





