What is KEGG
KEGG, abbreviation of Kyoto Encyclopedia of Genes and Genomes, is a collection of databases, which is used for bioinformatics research, including data mining in genomics, proteomics, metabolomics and other omics studies, modeling and simulation in systems biology, and translational research in drug R & D. Through the KEGG databases can be categorized into systems, genomic, chemical, and health information, which is distributed in the KEGG PATHWAY, KEGG BRITE, KEGG MODULE, KEGG GENES and so on. KEGG, connects known information on molecular interaction networks, such as pathways and complexes (the "Pathway" database), information about genes and proteins generated by genome projects (including the gene database) and information about biochemical compounds and reactions (including compound and reaction databases). These databases are different networks, known as the "protein network", and the "chemical universe" respectively. There are efforts in progress to add to the knowledge of KEGG, including information regarding ortholog clusters in the KEGG Orthology database.
KEGG PATHWAY database (the wiring diagram database) is the core of the KEGG resource, which contains the higher order functional information, including graphical representations of cellular processes, such as metabolism, membrane transport, signal transduction and cell cycle. A collection of pathway maps integrating genes, proteins, RNAs, metabolites, glycans, and chemical reactions, as well as genes involved in specific diseases and drug targets, are all stored as individual entries in the other databases of KEGG.
KEGG Annotation Analysis Service at Creative Proteomics include:
- KEGG Pathway annotation
- KEGG Pathway classification
The annotated proteins can be classified into the following sections with the aid of KEGG Annotation Analysis Service:
- Metabolism
- Genetic information processing (transcription, translation, replication and repair, etc.)
- Environmental information processing (membrane transport, signal transduction, etc.)
- Cellular processes (cell growth, cell death, cell membrane functions, etc.)
- Organismal systems (immune system, endocrine system, nervous system, etc.)
- Human diseases
- Drug development
Why Use KEGG Metabolic Pathways
KEGG provides exceptional integrated pathway queries, encompassing metabolism of carbohydrates, nucleotides, amino acids, and organic compound biodegradation. It not only offers comprehensive metabolic pathways but also annotates enzymes catalyzing each step of reactions, including amino acid sequences, links to the PDB library, and more. KEGG is a powerful tool for conducting intracellular metabolic analysis and metabolic network research.
- In comparison to other databases, KEGG stands out with its robust graphical capabilities, using visuals rather than verbose text to illustrate numerous metabolic pathways and their relationships.
- Starting from functionality, researchers can swiftly pinpoint genes associated with specific functions within pathways.
- Beginning with genes, one can determine a gene's role (upstream or downstream) in signaling pathways and its biological functions.
- Identify variations and functional distributions related to pathways.
- Intuitive graphics provide an overall impression of a gene's involvement.
How to Interpret KEGG Metabolic Pathways
In KEGG analysis results, it is crucial to understand the biological significance of identifiers like K01000 and ko01000:
K01000: Comprising an uppercase K followed by five digits, it corresponds to the identification of a specific class of proteins in the KEGG database. It represents proteins associated with certain genes, with one such major K identifier per gene.
ko01000: Comprising lowercase ko followed by five digits, it represents pathway identifiers. This code corresponds to a specific metabolic pathway. Within a pathway, multiple genes may collaborate in regulation. Consequently, multiple major K identifiers may exist within a pathway. Similarly, a single gene can participate in multiple metabolic pathways, resulting in multiple ko annotations.
Other symbols and their meanings in metabolic pathways:
M+ num: Module name.
C+ num: Compound name.
E-,-,-,-: Enzyme name.
R + num: Reaction name.
RC+ num: Reaction type.
RP+num: Reactant pairs.
Legend for Functional Relationships:
Legend for Functional Relationships:
Linking to Another Pathway

Various Metabolic Pathway Diagrams
Reference Pathway:
A reference pathway, also known as a reference metabolic diagram, is a comprehensive, detailed, and generalized representation of a metabolic pathway. These diagrams are constructed based on existing knowledge and serve as overarching references for understanding fundamental metabolic principles. In these diagrams, rectangular boxes within the pathway remain uncolored. In the KEGG database, these pathways are identified by names commencing with "map," such as map00010, which designates the reference pathway for glycolysis. These reference pathways offer invaluable insights into the common and widely accepted aspects of metabolic pathways.

Species-Specific Pathway:
A species-specific pathway, also referred to as a pathway specific to a particular organism, is a metabolic diagram that pertains exclusively to a specific species. In these diagrams, green boxes represent genes or enzymes unique to that particular species. Detailed information is available only for these green boxes, providing insights into the distinctive aspects of the species' metabolism.
In KEGG, these pathways are identified by the English abbreviation of the specific species, such as "hsa" for humans, followed by the pathway code, as seen in the example of the glycolysis pathway for humans, designated as "hsa00010."

KEGG Pathway Enrichment:
Pathway enrichment analysis is frequently applied in the functional annotation of differentially expressed genes, providing insights into the associated functions and biological pathways of these genes. In pathway enrichment plots, upregulated differentially expressed genes are delineated by red borders, while downregulated differentially expressed genes are indicated by green borders.
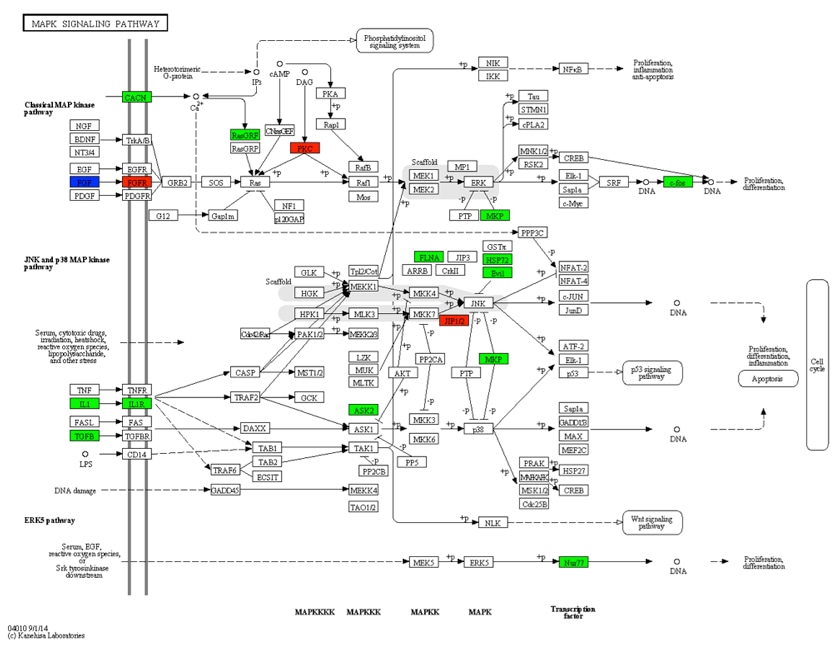
In living organisms, different metabolic products coordinate with each other to collectively regulate overall metabolism. Interpreting and selecting differentially enriched metabolites in pathway enrichment analysis plots often requires human intervention. Currently, after KEGG annotation enrichment, visualization techniques such as bubble charts and bar graphs are employed to analyze whether there is differential expression of genes along a specific pathway, providing enhanced visual insights.

In addition to providing professional bioinformatics services, Creative Proteomics also possesses a range of analytical instruments, including liquid chromatography triple quadrupole mass spectrometers, high-resolution liquid chromatography with quadrupole-Orbitrap mass spectrometers, gas chromatography-mass spectrometers, gas chromatographs, liquid chromatographs, amino acid analyzers, scanning electron microscopes, transmission electron microscopes, nuclear magnetic resonance spectrometers, and more. These instruments enable us to undertake a wide array of tests, including metabolomics, proteomics, transcriptomics, genomics, multi-omics integrative analysis, and polysaccharide structural elucidation.
How to place an order:

*If your organization requires signing of a confidentiality agreement, please contact us by email
As one of the leading omics industry company in the world! Creative Proteomics now is opening to provide KEGG annotation analysis service for our customers. With rich experience in the field of bioinformatics, we are willing to provide our customer the most outstanding service! Contact us for all the detailed informations!




