Understanding the causes of variation in evolutionary rates of protein is central for many fields, including molecular evolution, structural biology and comparative genomics. Determining the rate of protein evolution is arguably the most powerful tool to quantify the relative importance of selection and genetic drift, and to identify selective forces from genomic data. Analysis of protein evolution also provide a useful tool for studying issues such as the evolution of speciation, senescence and social lifestyle; and they facilitate the identification of functionally important sites, peptides that are involved in human genetic diseases, drug targets or protein interaction partners. Observed rates of protein evolution can also be used to study how different mutations might contribute to disease. In addition to sequenced-based and structure-based methods, genome-based method is also one of the most useful techniques to study protein evolution. Now, bioinformaticians at Creative Proteomics are proud to tell you we are open to help you with Genome-based Evolution Analysis of Proteins Service!
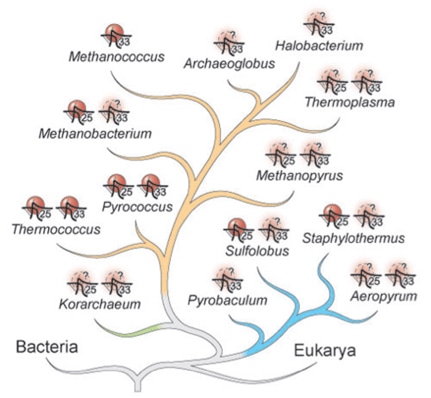
Protein evolution has a great relationship with changes and selection of DNA polymorphisms and mutations because protein sequences change responding to alterations in the DNA sequence. Amino acid sequences and nucleic acid sequences do not mutate at the same rate. Owning to the degenerate nature of DNA, bases may change without affecting the amino acid sequence. For instance, there are six codons that code for leucine. Thus, despite the difference in mutation rates, it is necessary to incorporate nucleic acid evolution into the discussion of protein evolution. At the end of the 1960s, two groups of scientists, Kimura (1968) and Jukes and King (1969), independently proposed that a majority of the evolutionary changes observed in proteins were neutral. Since then, the neutral theory has been expanded upon and debated.
How to place an order:

*If your organization requires signing of a confidentiality agreement, please contact us by email
As one of the leading omics industry company in the world! Creative Proteomics now is opening to provide genome-based evolution analysis of proteins service for our customers. With rich experience in the field of bioinformatics, we are willing to provide our customer the most outstanding service! Contact us for all the detailed information!




