A gene co-expression network is a graph, where each gene corresponds to a node and a pair of nodes is connected by an undirected edge if their pair-wise expression similarity score is above a specific threshold. Construction of co-expression networks from gene expression datasets has become a widely used alternative to the conventional analytic methods. Large-scale gene co-expression networks have been applied, e.g. to demonstrate that functionally related genes are often co-expressed across variety of datasets and across distinct organisms. By constructing co-expression networks for different conditions, such as normal and cancerous states, it is possible to investigate disease-mediated changes in the network connectivity patterns.
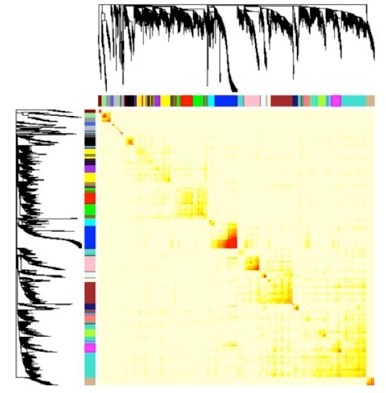
Correlation networks have been increasingly used in bioinformatics applications. For example, weighted gene co-expression network analysis is a systems biology approach for describing the correlation patterns among genes across different microarray samples. Weighted correlation network analysis, also called weighted gene co-expression network analysis (WGCNA), is a widely used correlation networks analysis method especially for the study of biological networks based on pair wise correlations between variables. While it can be used to most high-dimensional data sets, it has been most widely applied in genomic data mining. It allows one to define clusters (or modules), network nodes, and intramodular hubs according to module membership, to compare the relations between different co-expression modules, and to study the network topology of distinct networks.
Applications for Gene Co-expression Network
- Finding clusters or modules of highly relevant genes
- Summarizing such modules using an intramodular hub gene or the module eigengene
- Relating modules to one another and to external sample traits with eigengene network methodology
- Calculating the measures of module membership
How to place an order:

*If your organization requires signing of a confidentiality agreement, please contact us by email
As one of the leading omics industry company in the world! Creative Proteomics now is opening to provide gene co-expression network analysis service for our customers. With rich experience in the field of bioinformatics, we are willing to provide our customers the most outstanding service! Contact us for all the detailed information!




