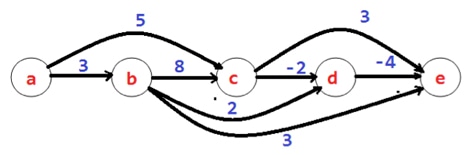
In the field of mathematics and computer science, a directed acyclic graph (DAG) is a finite directed graph with no directed cycles. In another word, it is composed of finitely a lot of nodes and edges, with each edge directed from one node to another, such that it is impossible to start at a given node v and follow a consistently-directed sequence of edges that finally loops back to node v again. Similarly, a DAG is a directed graph that has a topological ordering, a series of the nodes such that each edge is directed from earlier to later in the sequence. DAGs are a very important class of graphs, being part graph, part tree, and having a variety of applications, such as those relevant precedence among "events". Many problems in graphs become simpler using DAGs, such as game evaluation, expression tree evaluation, and path analysis.
GO is generally organized as a tree-like hierarchy of functional terms, in which each term can have children that are more-specific classes of the parent class. As a matter of fact, the GO hierarchy is a DAG rather than a tree, as a GO term can have several parents. Indeed, because GO covers three domains of biology knowledge and relationships are defined only within each kind of knowledge, GO is in fact three non-overlapping DAGs determined by their namespaces: biological process, molecular function and cellular component.
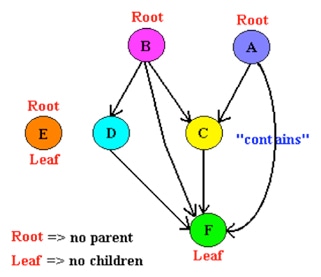
The DAGs provided by Creative Proteomics consists of the following sections:
- Nodes. Each node means some object or piece of data.
- Directed edges. A directed edge from one node to another represents some type of relationship between these two nodes. The arrow generally means “is based on”.
- A root node. At least one node will have no parent term. This is the root of the DAG.
- Leaf nodes. One or more of the nodes will have no child terms. They are called leaves or leaf nodes.
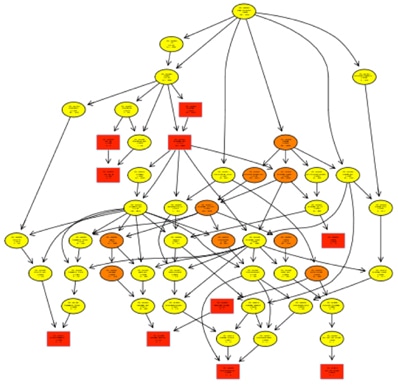
Since the GO is structured as a DAG where relationships of each term to one or more other terms has been defined in the same domain, and sometimes to other domains, an insightful way of looking at the results of the analysis is to investigate how the significant GO terms are distributed over the GO graph. Subgraphs induced by the most significant GO terms were plotted. In these plots, the significant nodes are described as rectangles. The plotted graph is the superior induced graph generated by those significant nodes. These graph plots are used to see how the significant GO terms are distributed in the hierarchy. It is a very useful tool to realize behaviour of various enrichment methods and to better understand which of the significant GO terms are really of interest.
How to place an order:

*If your organization requires signing of a confidentiality agreement, please contact us by email
As one of the leading omics industry company in the world! Creative Proteomics now is opening to provide DAG analysis service for our customers. With rich experience in the field of bioinformatics, we are willing to provide our customers the most outstanding GO DAG service! Contact us for all the detailed information!




