Epilepsy is the second most common disorder of the nervous system, plaguing the physical and mental health of people of all ages, and is listed by the World Health Organization as one of the top five neuropsychiatric disorders of global priority. The pathogenesis of epilepsy is complex and it is generally believed that genetic factors play a major role.
Proteomics is an emerging field in epilepsy research that focuses on the study of proteins and their functions in the context of the disease. Proteins play a crucial role in various cellular processes and are involved in the development and progression of epilepsy. By using various proteomic techniques, such as mass spectrometry, researchers aim to identify changes in the levels, expression, and modification of proteins in the brain that are associated with epilepsy. This information can then be used to better understand the underlying mechanisms of the disease, develop new diagnostic markers, and identify new therapeutic targets. Additionally, proteomics has the potential to provide a more comprehensive view of the complexity of epilepsy compared to traditional genetic studies, which only examine a limited number of genes. Thus, proteomics holds great promise for advancing our knowledge of epilepsy and improving the lives of those affected by this debilitating disease.
Case 1 Alterations in neuronal networks in patients with early onset genetic epilepsy (1)
The paper utilizes proteomics and phosphoproteomics based on mass spectrometry to detect abnormal protein expression changes and phosphorylation changes in neurons that have been induced to differentiate in the absence of CDKL5 protein. Several drugs were selected through a high-throughput drug screening platform that could rescue the abnormal neural network of CDD. The overall study reveals the cellular, molecular, and neural network mechanisms of hereditary epilepsy, which may ultimately bring new treatment opportunities to patients.
The samples for proteomics and phosphoproteomics included neural progenitor cells (NPCs), cortex-like organs, neuronal cells, and their respective control group samples. Proteomics quantified 6505 proteins, with downregulated proteins enriched in microtubule-based processes (23 proteins). Differential protein IPA analysis indicated impaired cellular activity and neuronal development in CDD. By evaluating the changes in cell count over time in vitro, the authors observed that CDD neural stem cells proliferated slowly compared to the control group, and changes in the cell cycle were also observed. In addition, an increase in DNA breaks, mitochondrial depolarization, and caspase activity was also observed, indicating an increase in cell death.
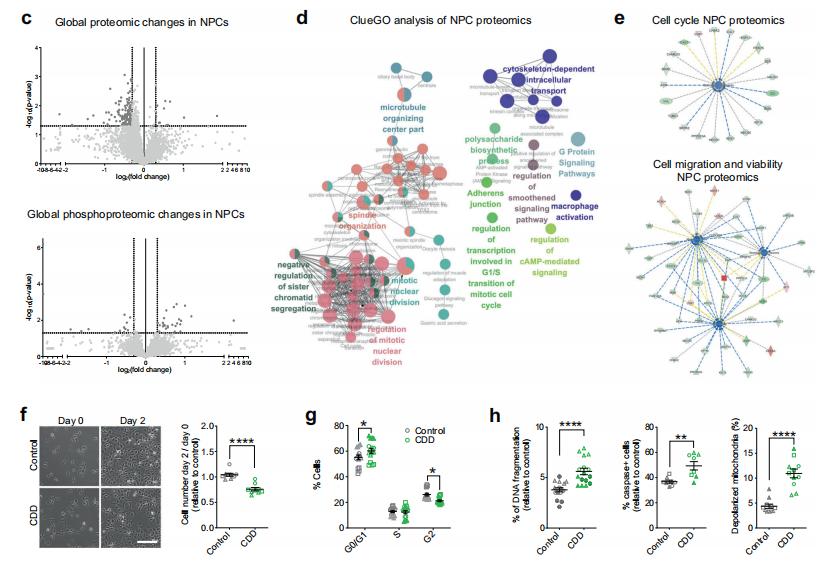
Altered proliferation and viability in CDD progenitor cells.
c Volcano plot. d ClueGO analysis. e Ingenuity pathway analysis (IPA). f Representative phase-contrast images. g Percentage of cells at different phases of the cell cycle compared to control. h Cell death analysis of CDD NPCs compared to control.
Phosphorylated proteomics reveals changes in CDD neuronal development and mTOR activation. In neurons, a total of 4115 phosphorylated peptides from 1485 proteins were identified and quantified. In cortical-like organs, a total of 6769 phosphorylated peptides from 2212 proteins were identified and quantified. Motif analysis of up-regulated phosphorylated peptides observed overexpression of pSXXE, a site phosphorylated by eosinophilic kinases such as CK2. IPA analysis and kinase enrichment analysis revealed that proteins known to be phosphorylated by CK2 were increased in CDD neurons, indicating that CK2 is over-activated. Phosphorylated proteomic analysis also revealed several dysregulated phosphorylated proteins in CDD neurons and organoids, such as microtubule-associated proteins, brain failure response regulatory protein family proteins, and cytoskeletal organizing proteins. Overall, phosphorylated proteins in neurons and organoids were enriched in the entries of neurogenesis, morphology, migration, synaptogenesis and neurotransmission. In addition, enrichment of the mTORC1 pathway was identified in the phosphorylated proteomic analysis of neurons. mTORC1 pathway activation was also confirmed by amino acid starvation experiments.
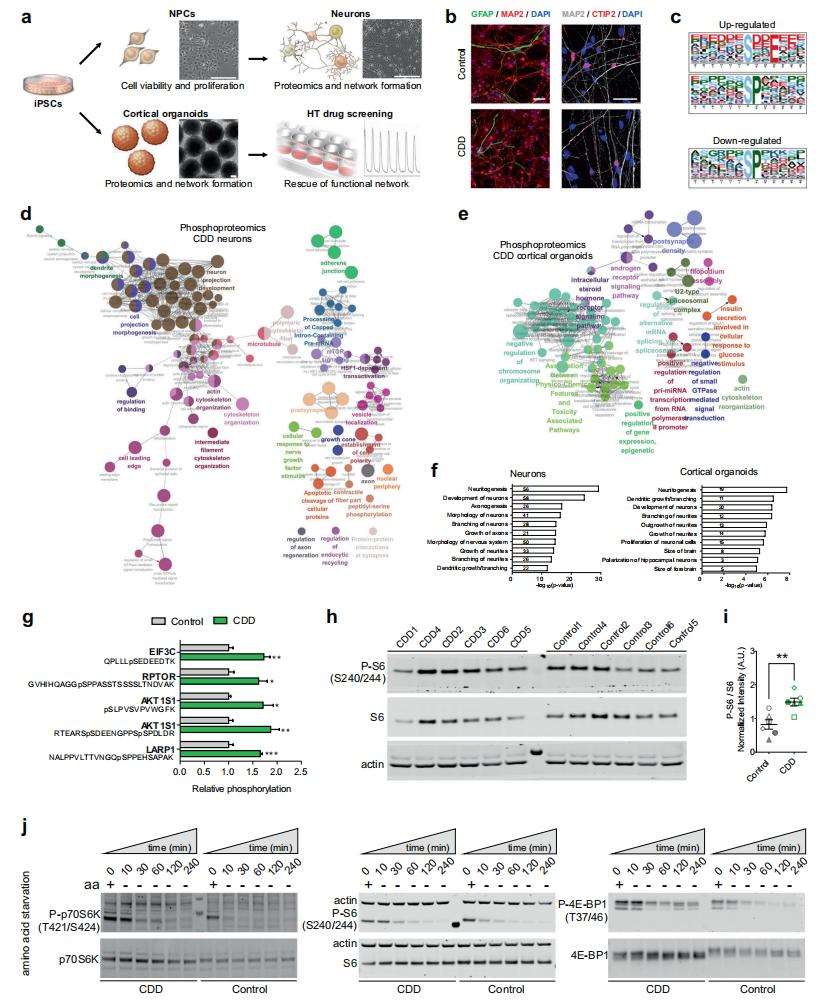
Proteomics and phosphoproteomics analysis reveal mTOR alterations in CDD neural cells.
The authors used proteomics and phosphorylated proteomics to reveal potential downstream molecules and pathways affected by CDKL5 mutations in CDKL5-deficient (CDD) brain cells and to describe the cellular and functional phenotypes associated with the molecular alterations.
Case 2 Proteomics of neurodevelopmental migration pathways (2)
Granular cell dispersion (GCD) is a common pathological feature observed in the hippocampus of patients with medial temporal lobe epilepsy (MTLE). The aim of this study was to compare the proteomes of hippocampal basal and dispersed granule cells in eight MTLE patients with GCD to identify proteins that may mediate GCD in MTLE.
Frozen hippocampal samples were used for proteomic studies. Samples were denatured using in-solution trypsin digestion and prepared for MSe label-free quantitative proteomics. Protein identification was obtained by searching the UniProt reference human proteome and adding porcine trypsin sequences. Bioinformatics analysis was then performed.
Quantitative proteomics identified 1,882 proteins, of which 29% were present in basal granule cells only, 17% in dispersed cells only and 54% in both samples (Fig. B).
Based on hierarchical clustering, some similar changes in protein expression were observed between the younger cohort of two cases (E1 and E2) and the three older cases, E5, E6 and E8 (Fig. C). Common neuronal markers such as MAP2, calcium binding protein and calreticulin were detected in the MTLE samples (Fig. D). Volcano plots (Figure E,H) show log-2 fold changes with -log P values for these proteins that were expressed in both dispersed and basal samples or in both young and old cohorts. Significantly differentially expressed proteins with more than 1.5-fold change are highlighted in the figure (P < 0.05). The abundance of each protein in Figures F and G is compared between the basal and dispersed samples, and between the younger and older cohorts. Proteins uniquely expressed in only one cohort, and proteins significantly expressed 1.5-fold more than the comparison cohort (fold change > 1.5, P < 0.05) generated clusters of dispersed, basal, young, and old. Each cluster was submitted to a bioinformatics platform for gene ontology and pathway analysis. Network diagrams show links between proteins in the small GTPase-mediated signal transduction (Fig. F) and ribosome annotation clusters ( Fig. G). Proteins in both the discrete and basal clusters are clearly involved in a number of pathways (Fig. I,J).
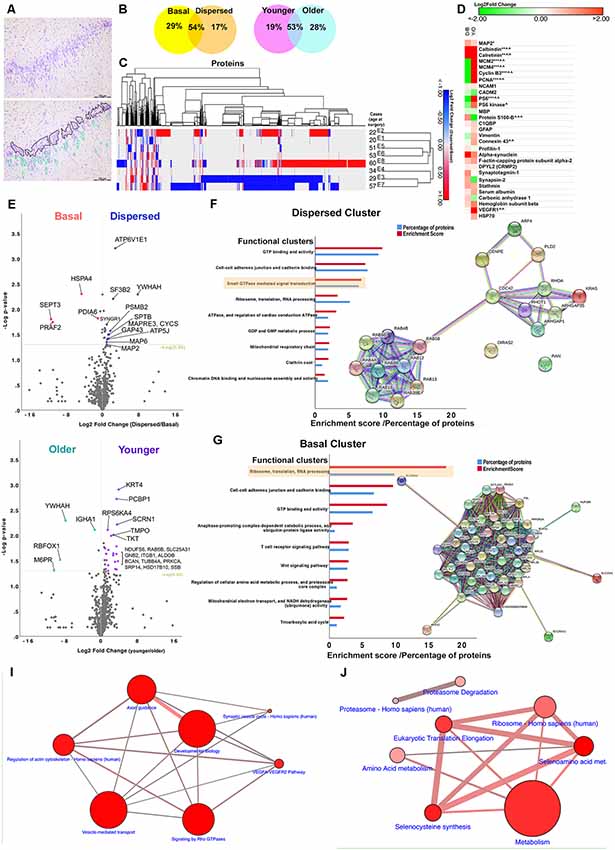
A key finding of this study was the identification of proteins specifically involved in axon guidance (including β-myostatin, growth-related protein 43, postsynaptic density protein 95, profilins), regulation of the actin cytoskeleton (cytoplasmic FMR1-interacting proteins. WAVE complex proteins), cytoskeletal and synaptic remodelling in dispersed DGCs from GCD cases (DPYL2, alpha-synaptic proteins, synaptic marker 1, synaptophysin II and stathmin1), and Rho GTPases, signaling (RhoA, Cdc42, Rac1, brain-specific angiogenesis inhibitors, Rho GTPases activating proteins). These processes are active in neurogenic niches in the neurodevelopmental and adult mammalian brain. Compared to controls, MTLE patients have increased expression of DPYL2 (also known as collapse response mediator protein 2, CRMP2), a cytoplasmic phosphoprotein that binds microtubules and promotes neurite growth during neurogenesis and neuronal migration, in the hippocampus.
Reference
- Negraes, Priscilla D., et al. "Altered network and rescue of human neurons derived from individuals with early-onset genetic epilepsy." Molecular psychiatry 26.11 (2021): 7047-7068.
- Liu, Joan YW, et al. "Granule cell dispersion in human temporal lobe epilepsy: proteomics investigation of neurodevelopmental migratory pathways." Frontiers in cellular neuroscience 14 (2020): 53.






