Protein Full Spectrum Analysis Service
- Home
- Applications
- Proteomics Analysis Services
- Protein Identification Services
- Protein Full Spectrum Analysis Service
Service Details
Full-spectrum protein analysis is to use mass spectrometry technology to identify the full-spectrum protein of complex biological samples (such as serum, urine, and cell lysates, etc.), and its purpose is to analyze and identify as many peptides and protein components as possible in mixture samples. Modern mass spectrometry techniques can better achieve full-spectrum identification of proteins due to their high sensitivity, high accuracy, and high throughput. Nowadays, shotgun proteomics is often used for protein profiling to identify samples, since the mixture samples contain intact proteins of different molecular weights, different solubility, and different modifications. The principle of shotgun identification is first the protein samples are digested into peptides, and separated by high performance liquid chromatography, then the separated peptides directly enter the mass spectrometer for analysis. Finally, the retrieval software can compare the mass spectrometry information with the corresponding data to obtain the exact sequence of the peptide segment, and then splicing the complete sequence of various proteins.
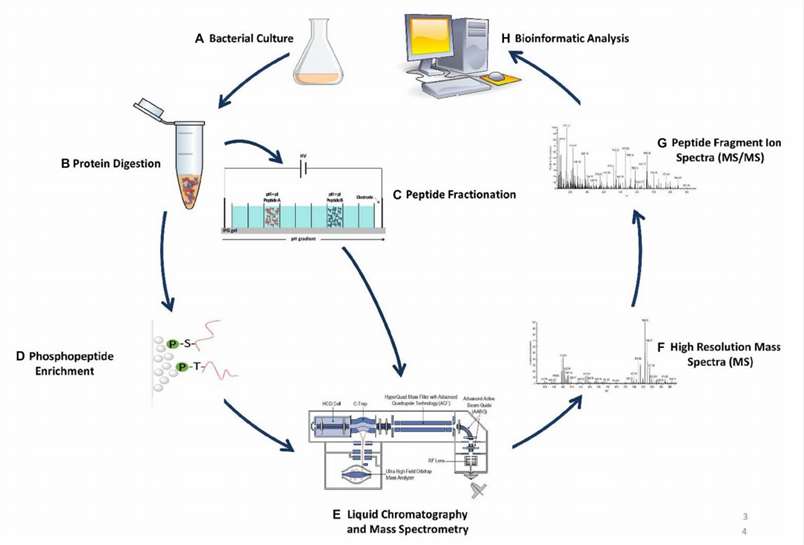 Fig. 1. Example of shotgun
proteomics workflow. (Soufi Y, et al., 2016)
Fig. 1. Example of shotgun
proteomics workflow. (Soufi Y, et al., 2016)
Creative Proteomics introduces a "bottom-up" approach to protein analysis, shotgun proteomics. Here, our highly qualified scientists utilize high-performance liquid chromatography (HPLC) techniques combined with mass spectrometry (MS) techniques to perform full-spectrum analysis of unknown proteins.
At Creative Proteomics, shotgun protein full-spectrum analysis is the use of shotgun technology for full-spectrum analysis of proteins, then protein identification is achieved using tandem mass spectrometry (MS/MS) to identify peptides. Our shotgun protein profiling allows for the identification of the entire protein spectrum of a sample, and hundreds to thousands of proteins can be identified simultaneously. The most striking feature is that it allows for the identification and quantification of multiple proteins with minimal protein separation.
With our powerful mass spectrometry sequencing platform, our shotgun proteomics full-spectrum analysis has been widely used by our customers for many different studies, including the following but not limited to:
Thanks to our powerful mass spectrometry sequencing platform, Creative Proteomics, as a first-class protein sequencing service provider, is committed to providing global customers with protein full spectrum analysis service to accelerate their project research journey. Our services guarantees accurate and reliable results, at quick turnaround time! Please contact us If you are interested in them.
References
For research use only, not intended for any clinical use.