Viral Epitope Discovery for Vaccine Development
The fundamental challenge in modern vaccine design—whether for pandemic preparedness or chronic viral infections—is identifying which viral fragments are actually displayed to the immune system. While a viral genome contains thousands of potential sequences, only a limited subset are processed and presented as HLA-bound peptides. These peptides serve as the primary targets for CD8+ and CD4+ T cell recognition, dictating the breadth and potency of cellular immunity.
Relying solely on prediction software often leads to high false-positive rates, as algorithms cannot fully simulate the biological intricacies of viral protein degradation, protease specificity, and MHC loading competition within the endosome. Our viral peptide discovery service provides direct, mass spectrometry-based evidence of presentation, allowing vaccine researchers to focus on immunodominant epitopes that are naturally displayed on the surface of target cells.
Immunopeptidomics for Viral Peptide Identification
Viral immunopeptidomics utilizes high-resolution liquid chromatography-tandem mass spectrometry (LC-MS/MS) to sequence peptides directly eluted from purified MHC molecules. This approach bypasses the limitations of functional validation assays, which often depend on access to suitable antigen-responsive T cells or donor immune material and require prior knowledge of the epitope sequence.
By analyzing the global landscape of peptides presented by infected cells for endogenous HLA-I presentation, and APC-based uptake/pulsing workflows for HLA-II and cross-presentation studies, we provide a high-resolution map of the viral peptidome. Our Immunopeptidomics Service is optimized to detect low-abundance viral ligands that may otherwise be masked by the vast background of endogenous host "self" peptides.
HLA-Presented Viral Peptides and T Cell Epitope Mapping
Precision T cell epitope mapping is the cornerstone of designing effective subunit and mRNA vaccines. Our workflow identifies both HLA Class I (CD8+ T cell) and HLA Class II (CD4+ T cell) ligands, providing a holistic view of the immune target landscape. This dual-class analysis is critical for vaccines requiring both cytotoxic killing and long-term helper-driven memory.
Once viral peptides are identified via mass spectrometry, we integrate NetMHCpan (Class I) and NetMHCIIpan (Class II) binding predictions to support peptide ranking. This integrated approach—combining experimental identification with computational modeling—provides a rigorous foundation for selecting candidate epitopes for downstream validation, such as ELISpot, Intracellular Cytokine Staining (ICS), or targeted Peptidomics-Based Antigen Discovery.
What We Offer in Viral Peptide and Epitope Discovery
We provide modular and integrated solutions to support viral antigen discovery at various stages of vaccine research:
Advantages of Our Viral Peptide Discovery Platform
Creative Proteomics offers a best-in-class platform for viral antigen discovery, providing several key advantages over standard epitope mapping techniques:
Workflow for Viral Epitope Discovery by Immunopeptidomics
Our standardized workflow ensures reproducible and high-quality results for complex viral samples:
Technology Platform for Viral Peptide Discovery
Our platform is engineered to meet the extreme sensitivity requirements of viral research. We utilize state-of-the-art Orbitrap high-resolution platforms, achieving sub-ppm mass accuracy and deep sequencing depth. This is supported by our comprehensive Immune Peptide Mass Spectrometry Analysis workflows.
Mass Spectrometry Platform Comparison for Immunopeptidomics
| Instrument Feature | Orbitrap Astral™ | Orbitrap Exploris™ 480 | Orbitrap Eclipse™ Tribrid™ |
|---|---|---|---|
| Sensitivity | Ultra-high sensitivity suitable for low-input and low-abundance peptide discovery | High (Robust discovery) | High (Intelligent acquisition) |
| Scan Speed | Up to 200 Hz | Up to 40 Hz | Up to 40 Hz |
| Recommended Applications | Rare viral peptides in limited samples | High-throughput viral peptidome mapping | Complex PTM-modified epitopes |
| Resolution | 80,000 at m/z 200 | Up to 480,000 at m/z 200 | Up to 500,000 at m/z 200 |
| Quantitative Mode | Label-free or Multiplexed | Label-free or TMT-multiplexed | TMT-optimized / Real-time search |
Viral Epitope Discovery vs. Epitope Prediction
| Dimension | Immunopeptidomics (Direct Discovery) | Computational Epitope Prediction |
|---|---|---|
| Discovery Approach | Experimental identification of physical peptides | Algorithmic probability of MHC binding |
| Primary Output | Sequenced and verified viral ligands | Predicted binding sequences |
| HLA-Presented Evidence | Direct biological proof of presentation | Theoretical affinity only |
| Strengths | Substantially reduces false positives from prediction-only workflows | Fast; high-throughput; no samples needed |
| Limitations | Requires biological samples/lysates | High false-positive rate; misses processing bias |
| Best-fit Project Stage | Candidate validation and lead prioritization | Broad early screening and library design |
Sample Requirements for Viral Peptide Analysis
To ensure successful discovery, samples must be prepared according to strict biosafety and quality guidelines.
| Sample Type | Typical Input | Required or Optional | Why It Matters | Notes |
|---|---|---|---|---|
| Infected Cell Lines | Inactivated lysates | Required (depending on design) | Enables naturally presented peptide discovery | Client must provide fully inactivated lysates |
| Recombinant Viral Proteins | Purified antigens | Optional | Supports antigen processing/pulsing workflows | Confirm expression system (e.g., HEK293) |
| Vaccine Candidates | Purified candidate proteins | Optional | Supports pre-vaccine epitope evaluation | Confirm formulation context |
| Antigen-Presenting Cells | Cell-based system | Required (depending on design) | Necessary for HLA-presented peptide generation | Confirm HLA matching/typing |
Note: Required quantities are dependent on viral titer and cell type; please consult our experts for project-specific guidelines.
Example Results from Viral Peptide Identification
Our deliverables translate raw mass spectrometry data into actionable vaccine research insights using Nature-standard visualizations.
Viral Peptide Length Distribution
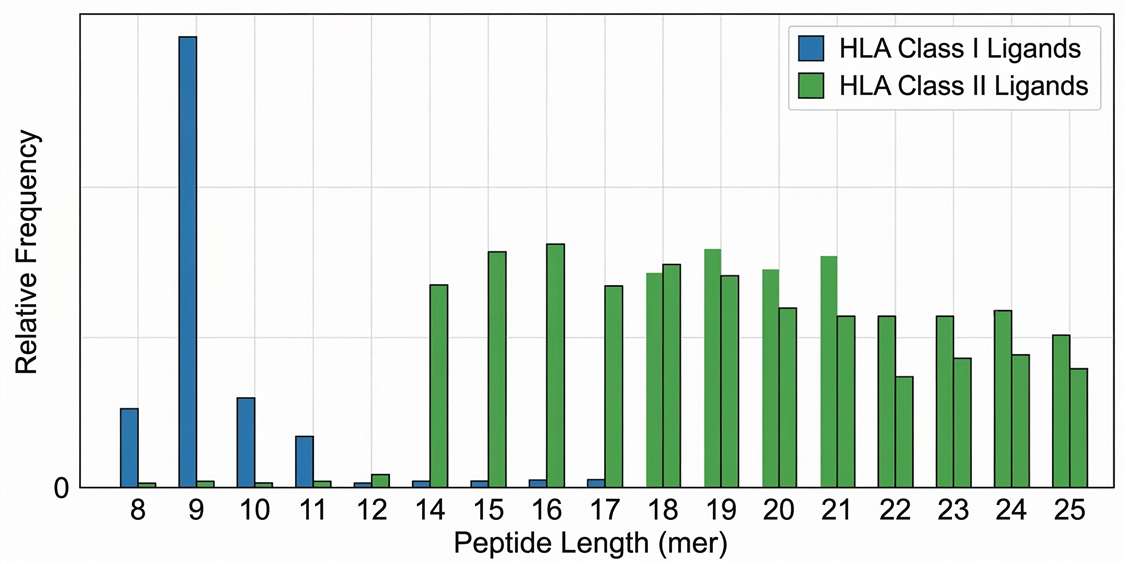
Distribution of peptide lengths for identified viral HLA ligands, a critical QC step confirming the characteristic presentation profiles for HLA Class I (9-mers) and Class II (12-25mers).
Viral Protein Coverage and Epitope Hotspots
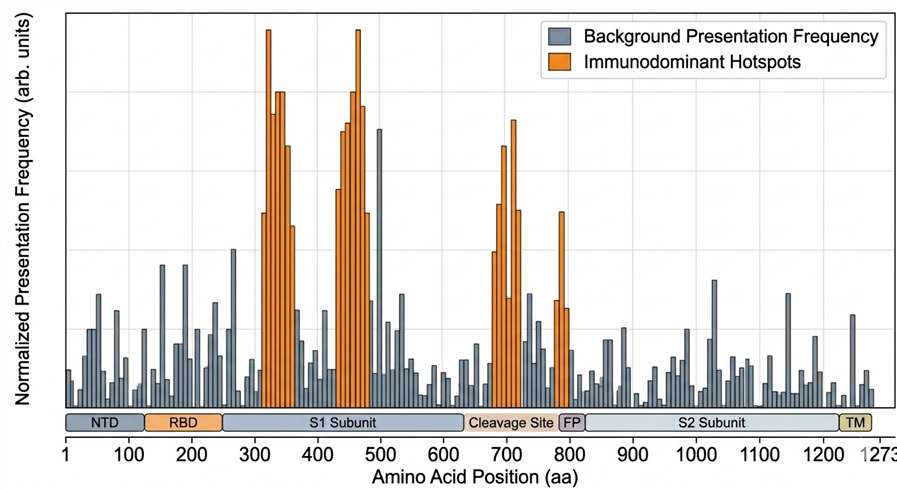
Comprehensive mapping of identified viral immunopeptides across the Spike protein sequence, pinpointing experimental presentation hotspots that represent immunodominant T cell epitope candidates for vaccine targeting.
Allele-Specific Binding Motif Logos
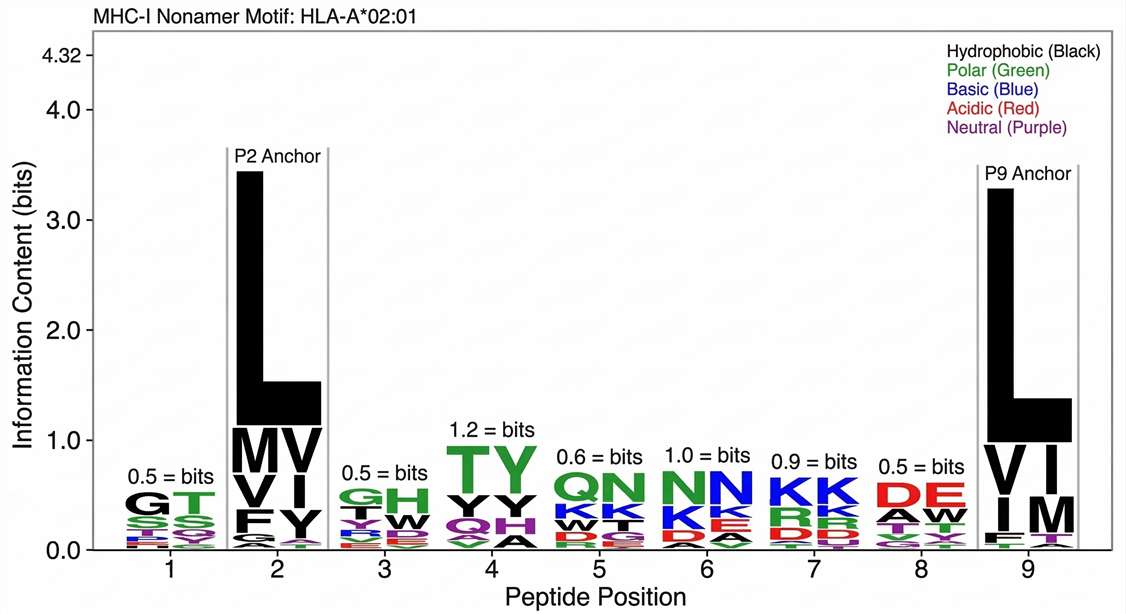
Bioinformatics-based WebLogo visualization of identified viral peptides, confirming specific amino acid conservation patterns and anchor residue preferences for the target HLA allele, such as Leu at position 2 for HLA-A*02:01.
HLA Allele Presentation Heatmap
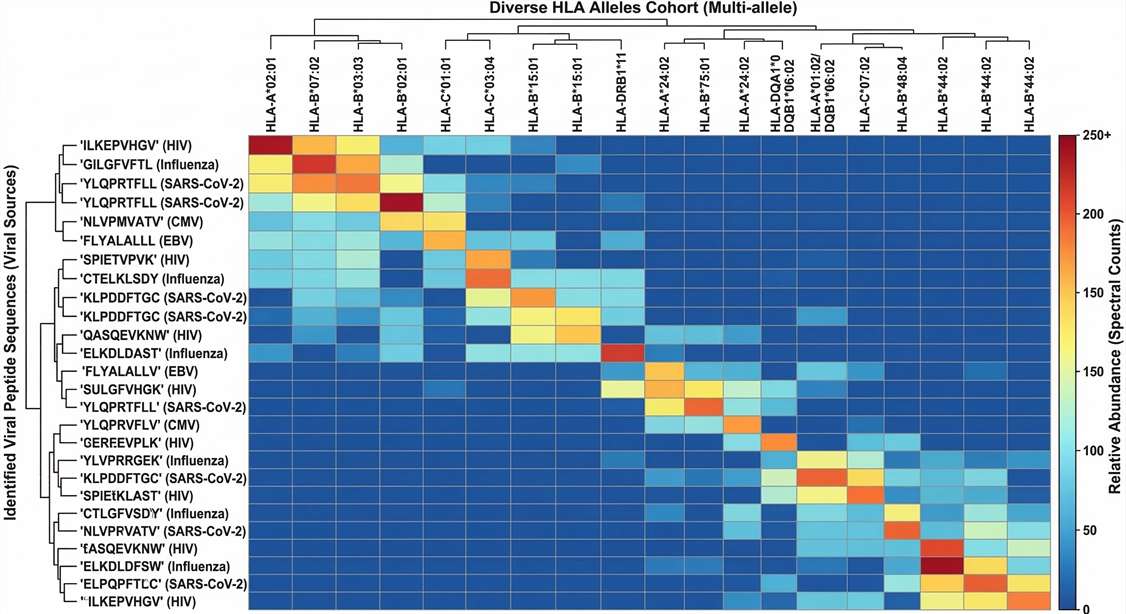
Multi-parameter heatmap illustrating the distribution and semi-quantitative abundance of identified viral peptides across a diverse donor HLA allele cohort, facilitating the selection of promiscuous vaccine epitope candidates.
Deliverables for Vaccine Epitope Discovery Projects
Creative Proteomics provides a structured data package ready for integration into your research reports:
- Full Analytical Report: Detailed methodology, QC metrics, and internal control performance.
- Viral Peptide Inventory: Comprehensive list including sequence, source protein, HLA allele assignment, and spectral abundance data.
- NetMHCpan/NetMHCIIpan Validation Report: Computational binding affinity predictions for all identified viral candidates.
- Epitope Prioritization Matrix: Expert analysis ranking candidates by presentation intensity and population coverage.
Frequently Asked Questions
Identification of Presented SARS-CoV-2 HLA Class I and HLA Class II Peptides Using HLA Peptidomics
Journal: Cell Reports
Published: Volume 35, Issue 13, 2021
Summary
Using HLA peptidomics, Nagler et al. directly identified naturally presented SARS-CoV-2 peptides from both HLA class I and HLA class II molecules. Rather than relying on prediction alone, the study combined infected-cell and viral gene expression models with LC-MS/MS-based immunopeptidomics to capture viral antigens that were genuinely processed and presented. In total, the authors identified 26 HLA-I peptides and 36 HLA-II peptides, including peptides derived from both canonical and non-canonical open reading frames (ORFs). Several peptides were shared across different cell types, and multiple identified ligands showed immunogenicity, supporting their value as experimentally grounded candidates for T cell epitope discovery and next-generation vaccine design.
Methods
This study used a combined HLA peptidomics + bioinformatics workflow to identify naturally presented SARS-CoV-2 ligands across prevalent HLA backgrounds.
Key Technical Features:
- Experimental systems: Cells infected with SARS-CoV-2 as well as cells transduced to express SARS-CoV-2 genes
- HLA context: Highly prevalent HLA class I alleles were prioritized to improve population relevance; both HLA-I and HLA-II peptidomes were analyzed
- Discovery platform: HLA-bound peptides were isolated and analyzed by LC-MS/MS-based HLA peptidomics
- Antigen scope: Analysis included peptides derived from both canonical viral proteins and out-of-frame / non-canonical ORFs
- Cross-cell comparison: Shared peptides presented across different cell types were identified to strengthen confidence in biological relevance
- Immunogenicity follow-up: HLA multimer staining was used to identify additional immunoreactive peptides beyond those already reported in prior studies
Creative Proteomics can support and extend this type of workflow through:
- High-sensitivity HLA class I and HLA class II immunopeptidomics by LC-MS/MS
- Viral peptide discovery from infected cell lysates or antigen-expression systems
- Canonical and non-canonical viral ORF-aware database searching
- Comparative ligandome analysis across cell types, infection conditions, or HLA backgrounds
- Experimental viral epitope prioritization supported by peptide annotation and binding prediction
- Custom bioinformatics for viral source mapping, hotspot analysis, and candidate ranking
These capabilities support vaccine epitope discovery, T cell target prioritization, and deeper characterization of viral antigen presentation landscapes.
Results
The study identified dozens of SARS-CoV-2-derived HLA-I and HLA-II peptides. A substantial portion of these peptides originated from viral proteins beyond Spike, and some were derived from non-canonical or out-of-frame open reading frames (ORFs).
Figure 1. Distribution of SARS-CoV-2 peptide lengths confirms standard HLA Class I and Class II presentation characteristics.
Naturally Presented SARS-CoV-2 Ligands Were Directly Identified
The authors identified 26 SARS-CoV-2-derived HLA-I peptides and 36 SARS-CoV-2-derived HLA-II peptides, providing direct evidence of viral antigen presentation.
Presented Peptides Originated from Multiple Viral Regions
Identified ligands were not limited to a single structural protein. Peptides were derived from both canonical viral proteins and non-canonical / out-of-frame ORFs, expanding the set of potential T cell targets beyond standard prediction-first approaches.
Shared Peptides Were Observed Across Different Cells
Some viral peptides were shared between distinct cell systems, strengthening confidence that these ligands reflect reproducible processing and presentation rather than a cell-line-specific artifact.
Immunogenic Peptides Were Identified
Among the detected ligands, seven peptides had been previously shown to be immunogenic, and the study identified two additional immunoreactive peptides using HLA multimer staining. This supports the translational relevance of direct HLA peptidomics for downstream T cell epitope validation.
Implications for Vaccine Design
Because the identified peptides span several viral genes and include naturally presented antigens beyond a spike-only focus, the results provide a stronger experimental basis for selecting T cell epitope candidates for next-generation SARS-CoV-2 vaccines.
Reference
- Nagler, Adi, et al. "Identification of Presented SARS-CoV-2 HLA Class I and HLA Class II Peptides Using HLA Peptidomics." Cell Reports 35.13 (2021): 109305. DOI: 10.1016/j.celrep.2021.109305