Decoding Peptide Signaling Across the Microbiota-Gut-Brain Axis
While much of the Microbiota-Gut-Brain Axis (MGBA) research has historically focused on small-molecule neurotransmitters or short-chain fatty acids (SCFAs), endogenous peptides represent the most direct and highly targetable signaling language between the gut and the brain.
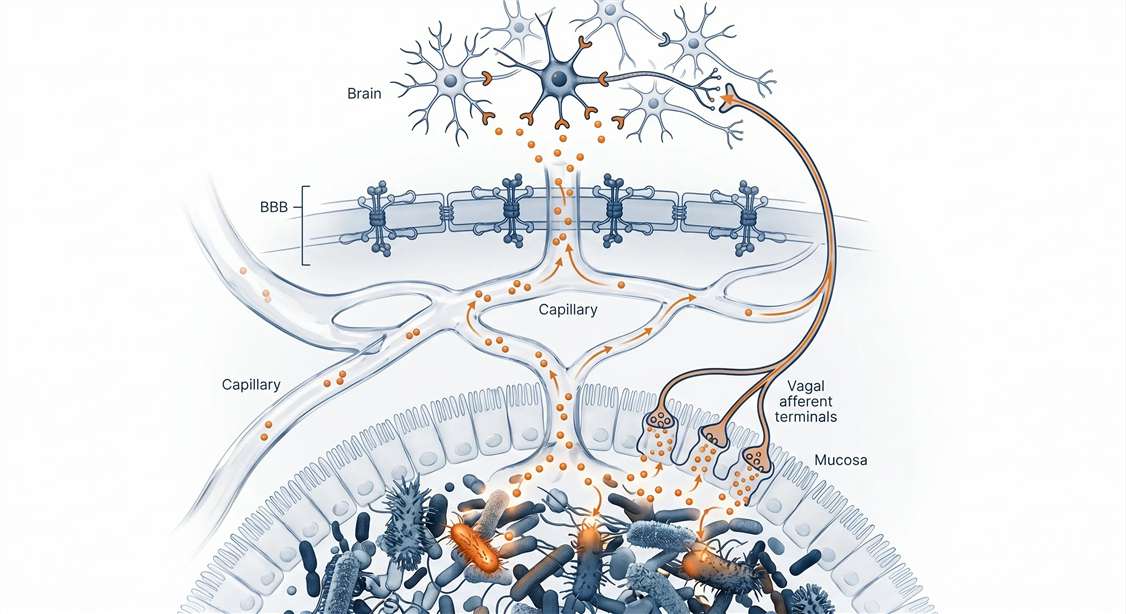
Gut-brain crosstalk is mediated by a vast array of signaling peptides, including host-derived enteroendocrine hormones (e.g., GLP-1, PYY, CCK, Ghrelin), central neuropeptides (e.g., NPY, Orexin, Substance P), and microbiome-derived bioactive peptides (microcins). Analyzing this dynamic peptidome requires moving beyond standard proteomics to capture the precise, functional peptide signals that actively mediate cross-kingdom communication.
Recommended Research Applications
Our specialized MGBA peptidomics platform is engineered to support translational medicine, microbiota-targeted drug development, and mechanistic neuroscience.
Overcoming Fecal Matrix and Low-Abundance Barriers in MGBA Profiling
Standard proteomics or metabolomics platforms routinely fail when applied to the MGBA due to two critical, opposing technical barriers:
- Extreme Complexity of the Fecal Matrix: Fecal and cecal contents are arguably the most challenging biological matrices, dense with bile acids, undigested lipids, and microbial debris. This matrix completely suppresses the mass spectrometry signal of low-abundance signaling peptides. We employ rigorous, proprietary solid-phase extraction (SPE) and fractionation workflows specifically optimized for fecal peptides.
- Low-Abundance Humoral & Vagal Pathways: We recognize that gut-brain communication occurs via dual pathways: humoral transit across the blood-brain barrier (BBB), and localized activation of vagal afferent terminals in the intestinal mucosa. The fraction of gut-derived peptides that enter circulation is present at extremely low concentrations. Our platform utilizes advanced enrichment and high-sensitivity mass spectrometry to capture these transient signals in plasma and brain tissues, regardless of their primary communication route.
Analytical Strategies: Host-Microbiome Peptidomics and Targeted Quantification
Untargeted Meta-Peptidomics for Discovery
For exploratory studies, we employ high-resolution untargeted neuropeptidomics coupled with specialized database searching. By utilizing combined host FASTA and microbiome Metagenome-Assembled Genomes (MAGs) databases, we perform meta-peptidomic analysis to discover novel microbial peptides and map the global host neuroendocrine response.
Targeted Quantification of Classic Gut-Brain Peptides
For preclinical drug evaluation and validation, we offer highly sensitive targeted neuropeptide quantification (PRM/MRM) multiplex panels tailored for classic gut-brain hormones, including GLP-1, GIP, PYY, CCK, Ghrelin, VIP, and central targets like NPY and AgRP.
Cross-Compartment Tracking Across Gut, Blood, and Brain
The ultimate proof of MGBA signaling is demonstrating a continuous pathway. We design multi-matrix analytical runs to track the abundance of specific peptides or functional clusters from the site of origin (feces/colon), through the transport route (plasma/serum), to the final destination (hypothalamus, hippocampus, or CSF).
Step-by-Step Workflow for Gut-Blood-Brain Peptide Tracking
Our end-to-end workflow incorporates rigorous quality control at every phase of the multi-compartment journey:
Typical Results: Translating Raw Data into Multi-Omics Mechanisms
A list of peptides is incomplete without understanding the microbial drivers behind them. We deliver publication-ready data visualizations that definitively prove cross-compartment MGBA signaling. Below are typical bioinformatics results demonstrating our meta-peptidomic capabilities:
Host vs. Microbiome Attribution
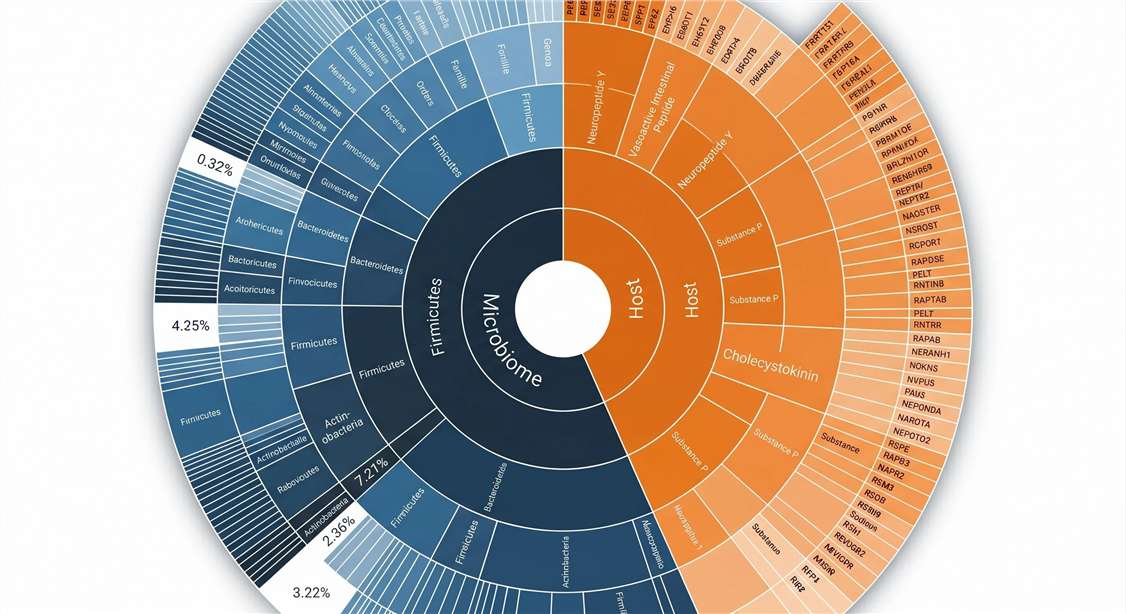
Taxonomic mapping via Meta-Database searching.
3-Compartment Tracking
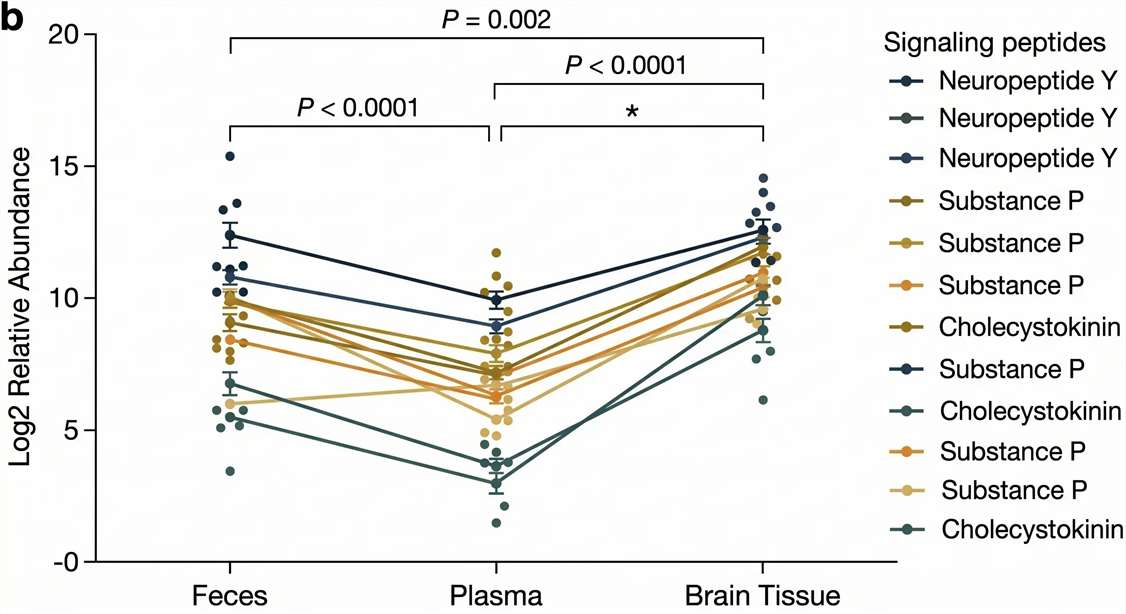
Tracking peptide transit across physical barriers.
Multi-Omics Correlation
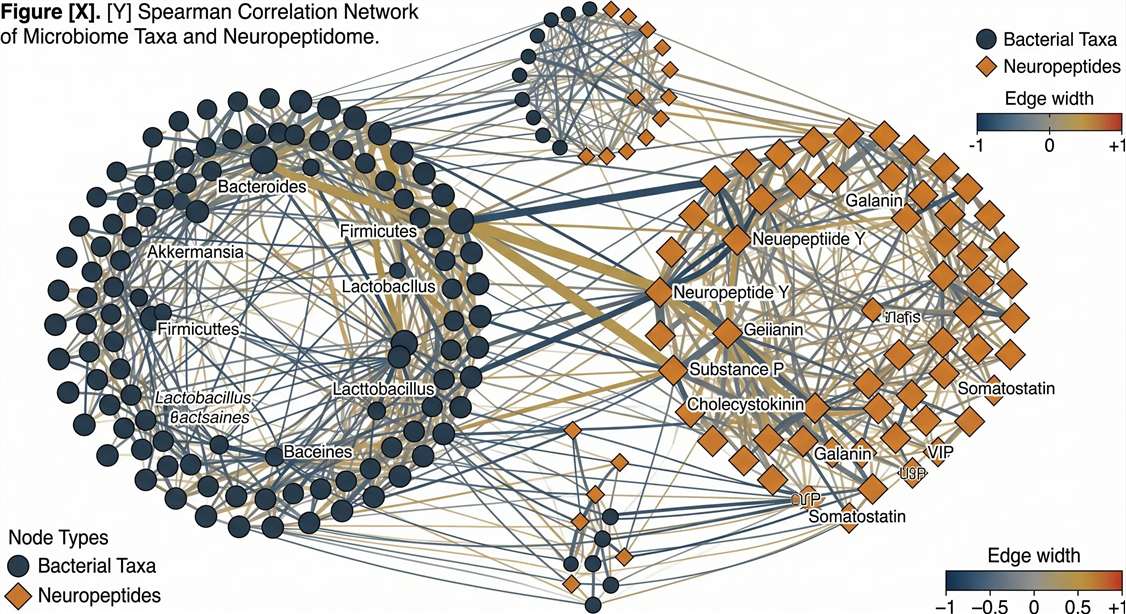
Linking bacterial taxa to CNS neuropeptide shifts.
Targeted GLP-1 Validation
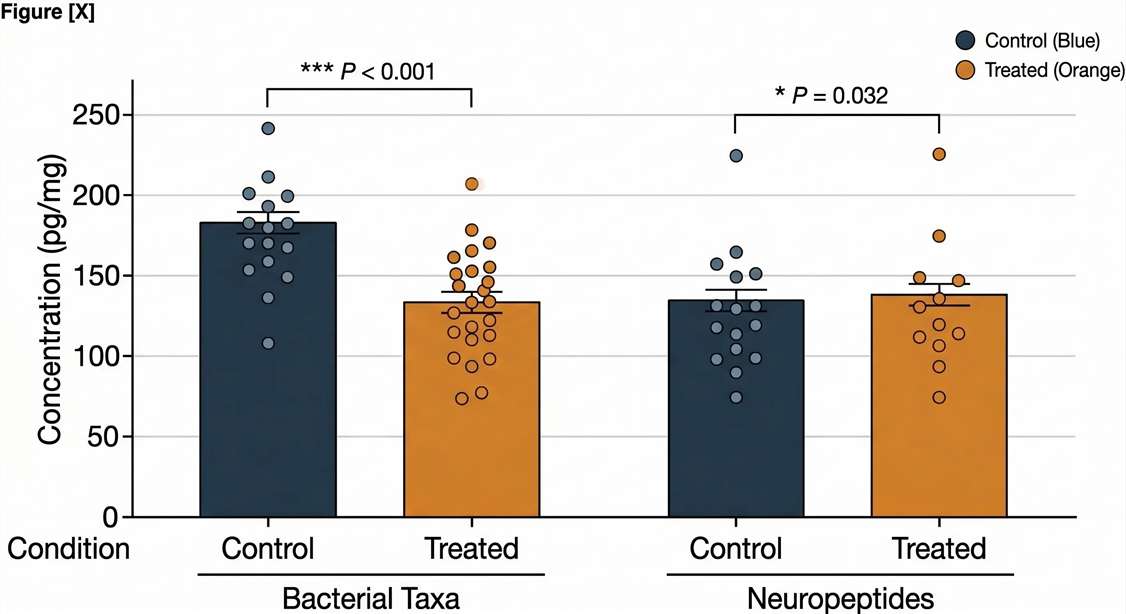
Absolute quantification via PRM/MRM.
Comprehensive Project Deliverables
Beyond visual results, your team will receive a complete, transparent data package ready for IND submission or high-impact publication:
- Taxonomic Attribution Report: Detailed lists of identified peptides with strict FDR-controlled host vs. microbiome assignments.
- Differential Expression Matrix: Statistically significant peptide abundance shifts across treatment groups and multi-matrix compartments.
- Multi-Omics Correlation Scripts: Ready-to-publish network models mapping 16S/Metagenomic features to the neuropeptidome.
- Targeted Quantification Summary: Absolute concentration values (e.g., pg/mg) for standard gut-brain peptides (GLP-1, PYY, CCK).
- Data Security & IP Ownership: 100% intellectual property ownership of your results. All raw MS files (.raw, .d), annotated sequences, and custom scripts are securely transferred via an encrypted cloud portal, ensuring absolute confidentiality for your drug development pipelines.
Multi-Omics Bioinformatics: Correlating the Microbiome with the Neuropeptidome
If you possess corresponding 16S rRNA or shotgun metagenomics data for your samples, we integrate this with our neuropeptide profiling data.
We generate Spearman Correlation Networks linking specific bacterial taxa (e.g., Bacteroides, Akkermansia) to the altered abundance of specific brain neuropeptides. This multi-omics integration provides the definitive mechanistic link needed to prove that altering the gut microbiome actively reshapes the brain's signaling architecture.
Standard Metabolomics vs. MGBA Neuropeptidomics
| Capability | Standard Metabolomics / Proteomics | Our MGBA Meta-Peptidomics |
|---|---|---|
| Signal Target | Global proteins, SCFAs, small molecules | Endogenous signaling peptides & microbial microcins |
| Fecal Matrix Handling | Basic homogenization (heavy suppression) | Specialized peptide SPE and lipid/bile depletion |
| Database Searching | Standard Host FASTA | Dual Host-Microbiome Meta-database searching |
| Multi-Compartment Tracking | Seldom correlated across physical barriers | Dedicated Gut ➔ Blood ➔ Brain abundance mapping |
| Biological Insight | General metabolic state | Direct neuroendocrine receptor signaling |
Sample Requirements and Preservation for MGBA Matrices
Endogenous peptides degrade within minutes due to ubiquitous proteases. Classic gut peptides like GLP-1 and PYY are rapidly cleaved by circulating enzymes such as DPP-4, resulting in ultra-short half-lives. Strict adherence to sample preparation guidelines is critical for gut-brain axis research.
| Matrix Type | Suggested Amount | Preservation | Key Experimental Considerations |
|---|---|---|---|
| Feces / Cecal Contents | 100 - 200 mg | Snap-freeze, -80°C | High heterogeneity; GF/Antibiotic models require strict sterility. |
| Plasma / Serum | 200 - 500 μL | Add Broad-spectrum + DPP-4 Inhibitors | Avoid hemolysis; collect into pre-chilled tubes to arrest active signaling states. |
| Brain Tissue (e.g., Hypothalamus) | 20 - 50 mg | Snap-freeze, -80°C | DO NOT wash in PBS; fast extraction is critical. |
Guidance on Biological Replicates
The microbiota-gut-brain axis is inherently noisy due to the extreme heterogeneity of the gut microbiome and individual metabolic differences. To achieve robust statistical significance in differential peptide expression, we strongly recommend:
- Inbred Animal Models (e.g., C57BL/6): n = 8 to 10 biological replicates per group.
- Clinical/Human Cohorts: n ≥ 30 per group (case-by-case evaluation recommended based on dietary/medication metadata).
- Matched Sampling: Whenever possible, collect feces, plasma, and brain tissue from the same individual subject to enable accurate intra-subject cross-compartment correlation tracking.
Shipping Guidelines
All MGBA samples must be shipped strictly on dry ice. Please contact our project managers to discuss custom protease inhibitor recommendations prior to your animal sacrifice or clinical collection phase.
Disclaimer: All services and analytical platforms described are intended for Research Use Only (RUO). Not for use in diagnostic procedures.