Challenges in Orphan GPCR Ligand Discovery
G protein-coupled receptors (GPCRs) represent the most successful class of therapeutic targets in modern medicine, yet over 100 "orphan" GPCRs (oGPCRs) remain without known endogenous ligands. Unlocking this "dark GPCRome" is a critical frontier for neurobiology and metabolic drug discovery.
However, identifying the true physiological ligands is immensely challenging. While oGPCR ligands can range from lipids to small metabolites, endogenous neuropeptides represent the largest, most structurally complex, and therapeutically promising class of undiscovered ligands. Yet, these neuropeptides exist in sub-picomolar abundances, are rapidly degraded by tissue proteases, and are often masked by high-abundance proteins in biofluids or secretomes. While traditional high-throughput screening of synthetic peptide libraries is common, it frequently derails discovery programs. Synthetic libraries inherently lack physiological context, omit complex native post-translational modifications (PTMs) crucial for receptor activation, and consistently yield high false-positive rates driven by non-specific hydrophobic binding.
When to Use Orphan GPCR Ligand Discovery
This highly specialized, project-specific service is designed for R&D leaders, Directors of Drug Discovery, and Principal Scientists driving advanced target validation. It is highly recommended when:
Integrated Deorphanization Workflow for Endogenous Ligand Discovery
Our solution is not a generic untargeted profiling service. It is a bespoke, receptor-oriented pipeline that actively de-risks your R&D investment by bridging early discovery with downstream functional validation.
Phased Project Milestones & Go/No-Go Checkpoints
Because deorphanization inherently carries discovery risks, we structure our custom projects in risk-mitigated phases, ensuring your budget is only deployed when analytical milestones are met.
| Project Phase | Key Deliverable | Timeline | Go/No-Go Checkpoint |
|---|---|---|---|
| Phase 1: Feasibility & Extraction | Pilot tissue extraction & MS QC report | 2–3 Weeks | Go/No-Go: Are native peptides adequately preserved and detectable? |
| Phase 2: Deep Profiling | Global peptidome mapping (Orbitrap Astral™ / timsTOF Pro) | 4–6 Weeks | Go/No-Go: Is the depth sufficient to capture sub-picomolar candidates? |
| Phase 3: Prioritization | Bioinformatics scoring & Top 5-20 ligand shortlist | 2–3 Weeks | Proceed to synthesis and cellular validation. |
Receptor-Centric Ligand Prioritization
Identifying thousands of native peptides is relatively straightforward; determining which specific peptide activates your orphan receptor is the true bottleneck. Our core differentiator is a proprietary prioritization algorithm that turns massive multi-omics datasets into a decision-grade candidate shortlist.
- Evolutionary Conservation & Co-expression: Prioritizing sequences that demonstrate high evolutionary conservation and exhibit spatial cross-tissue co-expression matching the orphan receptor's known distribution.
- PTM Activation Status: Up-weighting peptides possessing specific native modifications (such as C-terminal amidation or N-terminal pyro-glutamylation) that are fundamentally required for GPCR binding.
- Ligand-Receptor Interaction Likelihood: Leveraging advanced structural bioinformatics (including AI-driven peptide-protein docking and integration with global databases like GPCRdb) to evaluate the biochemical compatibility between the candidate peptide and the receptor's extracellular binding pocket.
- Cleavage Motif Recognition: Ensuring the candidate aligns with physiological prohormone convertase processing pathways.
Why Functional Assays and Synthetic Libraries Are Not Enough
Historically, "reverse pharmacology"—the process of identifying an endogenous ligand for a newly sequenced orphan receptor—relied heavily on screening massive libraries of known compounds or tissue homogenates. However, functional assays and synthetic libraries are effective primarily for screening known or engineered compounds. They fundamentally fail to answer the core question of deorphanization: "What does this receptor bind to in the natural biological system?"
| Comparison Dimension | Endogenous Peptidomics (Our Approach) | Functional Assays (Cell-Based) | Synthetic Peptide Libraries |
|---|---|---|---|
| Identify Unknown Endogenous Ligands | Excellent. Directly discovers novel native sequences. | Poor. Requires pre-existing ligand inputs to test. | Poor. Limited strictly to library contents. |
| Physiological Relevance | High. Reflects true tissue-specific biology. | Moderate. Often uses artificial overexpression. | Low. Context-free chemical interactions. |
| PTM Preservation | Complete. Detects native amidation/sulfation. | N/A. Depends entirely on what is added. | Limited. Complex PTMs are costly to synthesize. |
| False Positive Rate | Low. Mitigated by biological prioritization scoring. | High. Vulnerable to non-specific receptor activation. | High. Prone to off-target hydrophobic binding. |
| Receptor Specificity Inference | Strong. Supported by co-expression mapping. | Moderate. Confirms activation, not natural pairing. | Weak. Only indicates theoretical binding capacity. |
Technical Benchmarks and Quality Control Checkpoints
To ensure your functional validation budget is spent on true candidates, stringent quality control is embedded into every layer of our platform:
- Pre-Analytical Degradation Checks: Strict Go/No-Go checkpoints evaluating tissue integrity and rapid thermal stabilization efficiency before costly MS sequencing begins.
- High-Abundance Protein Depletion: Advanced biofluid and secretome handling protocols to prevent masking by serum albumin or cell culture media components.
- False Discovery Rate (FDR) Control: 1% FDR applied at both the peptide and PTM-localization levels to ensure the purity of the candidate pool.
Applications in Target Validation and GPCR Drug Discovery
- Orphan GPCR Target Validation: Move from generic genetic associations to concrete molecular pathways.
- Endogenous Ligand Identification: Establish the native pharmacophore to guide subsequent small-molecule or peptide drug design.
- Mechanistic Elucidation: Uncover novel neuroendocrine signaling axes regulating metabolism, pain, immunology, or cognition.
- Pipeline Acceleration: De-risk pharma pipelines by securing the biological "ground truth" before committing to synthetic screening campaigns.
- Bridging Genotype and Phenotype: Provide the missing biochemical link explaining observed knockout model phenotypes.
Demo Results and Deliverables for Orphan GPCR Ligand Discovery
We translate multidimensional omics data into highly visual, decision-ready reports designed to guide your next laboratory steps.
Prioritization Ranking
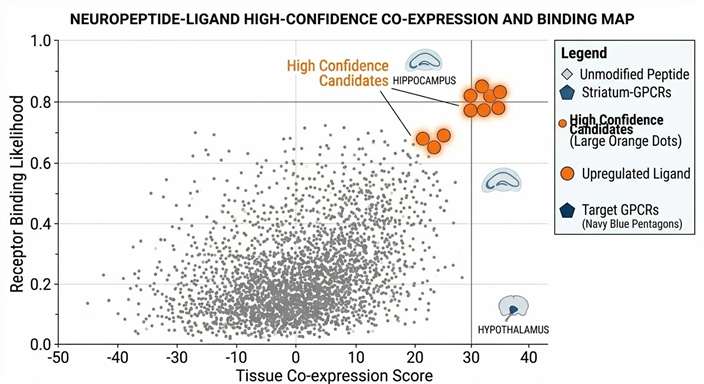
Algorithm-driven plots displaying top ligand candidates filtered by conservation and cleavage scores.
Multi-Dimensional Peptides
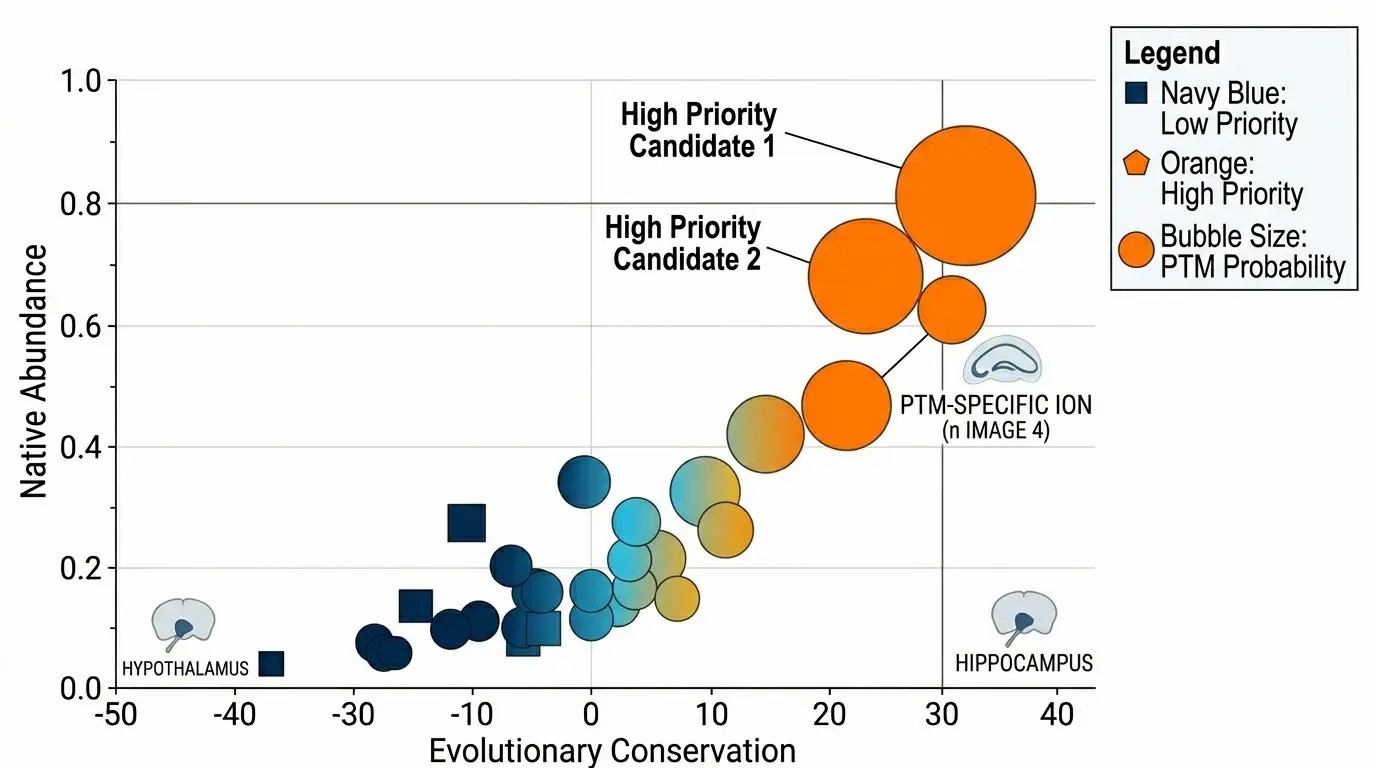
Bubble chart displaying candidate peptides where size indicates native abundance and color indicates PTM confidence.
Interaction Modeling
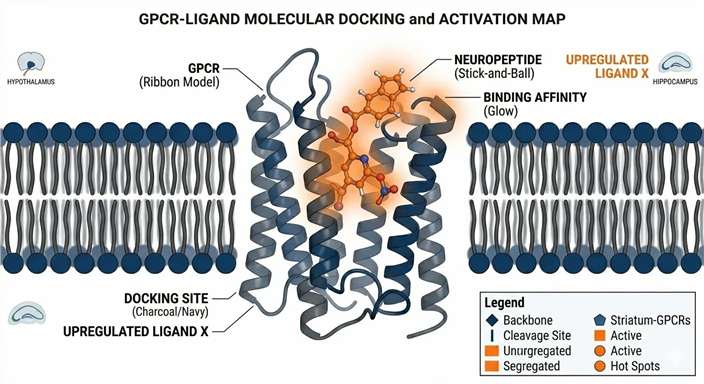
Structural hypotheses providing early insights into how the prioritized endogenous peptide interacts with the GPCR.
Candidate Heatmap
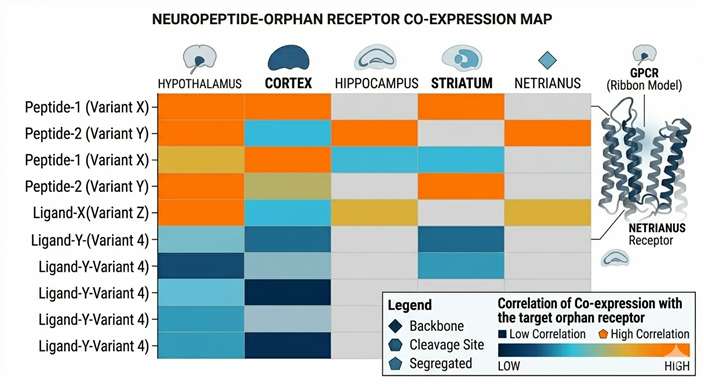
Tissue expression alignment cross-referencing native peptide abundance with spatial oGPCR distribution.
Comprehensive Project Deliverables
- Ranked Endogenous Ligand Shortlist: The top 5–20 highest-confidence candidates recommended for synthesis.
- Validation Strategy Guidance: Expert consultation on experimental design to validate the shortlisted hits.
- Ligand Prioritization Scoring Report: Detailed documentation of the exact rationale (co-expression, PTMs, docking) behind the ranking.
- Annotated Dataset: Complete PTM-aware characterization of the native peptidome.
Sample Requirements for Ligand Discovery Projects
| Sample Type | Recommended Use | Min. Input | Key Requirements |
|---|---|---|---|
| Brain regions / Microdissections | Neurological oGPCR ligand mapping | ≥10 mg | Immediate thermal stabilization or snap-freeze required. |
| Neuroendocrine tissues | Metabolic/hormonal receptor pairing | ≥20 mg | Optimal for identifying highly processed precursor pools. |
| Cell models (Receptor expressing) | Autocrine/paracrine signaling discovery | 107 cells | Secretome collection; requires rapid quenching of media proteases. |
| CSF / Plasma | Translational validation (Optional) | 200 µL | Low-binding materials; rigorous protease inhibition. |
Disclaimer: All services and platforms described are for Research Use Only (RUO). Not for use in diagnostic procedures.
Literature Demonstration of Endogenous Deorphanization
Journal: Nature Chemical Biology
Published: 2025
Background
Uncovering the physiological roles of highly conserved orphan GPCRs frequently stalls due to the inability to match them with native ligands. Standard synthetic libraries routinely miss complex, tissue-specific endogenous peptides, particularly those requiring highly specific post-translational modifications (PTMs) to achieve bioactivity.
Discovery Phase (Peptidomics)
A groundbreaking study published in Nature Chemical Biology (2025) perfectly exemplifies the power of the endogenous discovery methodology. Rather than utilizing synthetic screening, the research team employed advanced peptidomics to extract intact, fully modified endogenous peptides from relevant neuroendocrine tissues. By utilizing strict thermal stabilization and MWCO enrichment, they captured thousands of native peptides, including extremely low-abundance species.
The Prioritization Funnel
Generating a list of thousands of peptides is insufficient for downstream validation. The researchers applied a critical bioinformatics prioritization funnel—the core philosophy underlying our service. They filtered the massive dataset by scoring candidates based on three key pillars:
- Evolutionary Conservation: Selecting peptides highly conserved across mammalian species.
- PTM Profiling: Prioritizing peptides demonstrating C-terminal amidation, a classic hallmark of bioactive GPCR ligands.
- Spatial Co-expression: Cross-referencing peptide abundance specifically in tissue regions where the target orphan receptor is highly expressed.
This strict biological logic successfully distilled a massive pool of >5,000 native peptides down to a highly actionable shortlist of just 8 candidates.
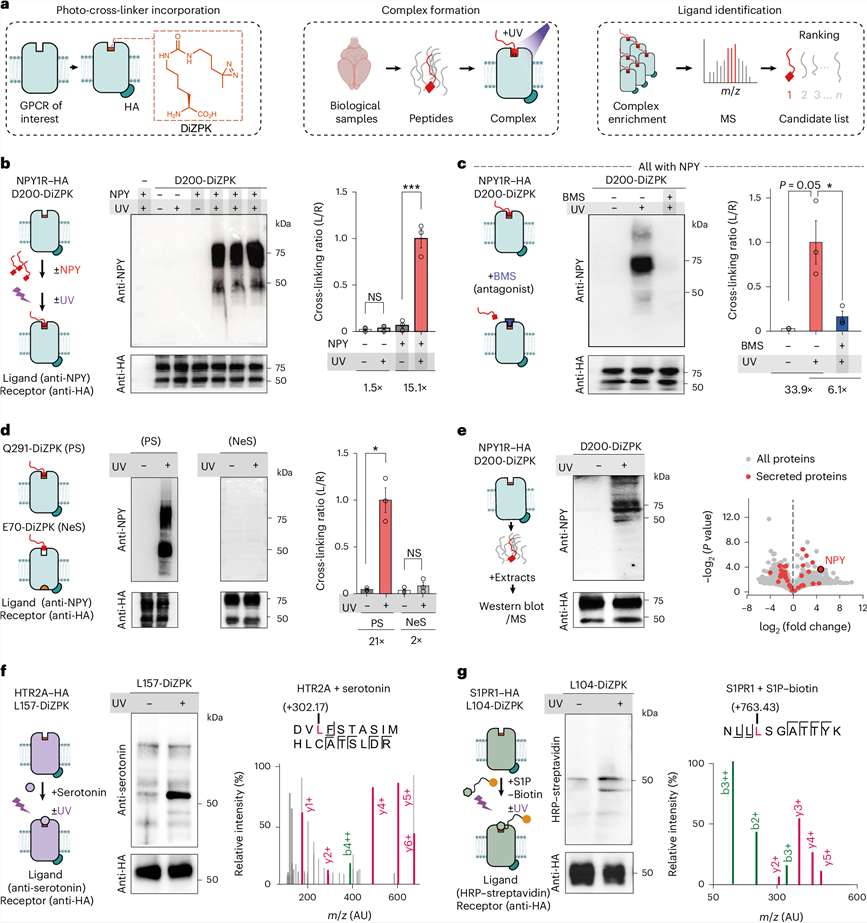
Functional Validation & Value
The top 8 shortlisted candidates were synthesized and evaluated in cell-based functional assays. The top-ranked native candidate successfully induced robust, dose-dependent receptor activation with an EC50 in the low nanomolar range. This published success validates our approach: leveraging native peptidomics coupled with stringent bioinformatic prioritization accurately pairs receptors with their true physiological ligands, saving months of trial-and-error screening and generating ready-to-use leads for drug discovery.
Reference
- "Photo-cross-linking-assisted deorphanization of GPCRs." Nature Chemical Biology (2025). https://doi.org/10.1038/s41589-025-02098-6