Why Connect Antigen Discovery with T Cell Response Analysis?
In modern cellular immunity research, no single technology fully explains antigen-specific T cell responses. While advanced mass spectrometry can identify thousands of naturally presented peptides on the cell surface, it provides no information about which of these peptides actually trigger a T cell response. Conversely, T cell receptor (TCR) sequencing expertly profiles clonal expansion and repertoire diversity, but a TCR sequence alone does not directly reveal its target antigen identity.
To build a stronger case for antigen-specific T cell immunity, researchers need an integrated workflow that connects antigen presentation, clonotype dynamics, and downstream validation. This integrated approach reduces reliance on purely predictive assumptions and helps prioritize biologically relevant candidates for downstream development.
Immunopeptidomics for Naturally Presented Antigen Discovery
The foundation of the workflow is our immunopeptidomics service, which directly sequences the peptides physically eluted from HLA molecules. By utilizing HLA peptidomics analysis, we provide direct, physical proof of which antigens are naturally processed and presented on the cell surface. To accommodate diverse sample types and research scales, we utilize multiple state-of-the-art mass spectrometry platforms.
Antigen Discovery Mass Spectrometry Platforms
| Parameter | Orbitrap Series (e.g., Astral, Exploris 480) | timsTOF Series (Pro/HT with PASEF) |
|---|---|---|
| Primary Strength | Deep, high-resolution profiling | Ultra-high sensitivity and speed |
| Mass Resolution | Ultra-high (sub-ppm mass accuracy) | High (enhanced by ion mobility separation) |
| Sample Input Flexibility | Ideal for standard to large tissue/cell inputs | Exceptional for low-input or limited clinical samples |
| Throughput | High | Ultra-high |
| Best Fit For | Comprehensive immunopeptidome mapping | High-throughput, sensitivity-constrained profiling |
TCR Sequencing for Immune Repertoire Profiling and Clonotype Tracking
To understand the immune effector side, TCR sequencing profiles the diversity and expansion of T cells. We offer flexible options supporting both bulk TCR repertoire profiling (profiling alpha and beta chains across a population) and single-cell immune profiling (scTCR-seq combined with scRNA-seq). This tracks which specific T cell clones are expanding in response to a disease state, vaccination, or immunotherapy treatment.
TCR-pMHC Interaction Validation via SPR and BLI Binding Assays
Identifying a peptide and a reactive T cell clone is powerful, but confirming their exact physical interaction requires downstream validation. We rely primarily on Surface Plasmon Resonance (SPR) to deliver high-precision kinetic data (KD, kon, koff) for highly selected "lead" TCR-pMHC pairs. While SPR is our gold standard for absolute kinetic quantification, we can also supplement with Bio-Layer Interferometry (BLI) for broader immunopeptide affinity screening.
Because TCR-pMHC interactions are often weak and technically challenging, SPR/BLI is typically reserved for highly prioritized candidates with suitable reagent quality and assay feasibility. Where appropriate, orthogonal validation strategies such as peptide-HLA stabilization assays, tetramer staining, or cell-based functional assays may also be recommended.
Key Applications in Immuno-Oncology, Neoantigen, and Vaccine Research
This integrated solution is designed with a balanced approach, providing equal utility for both immuno-oncology and infectious disease research:
End-to-End Workflow: From Antigen Discovery to TCR-pMHC Validation
Bridging multiple complex omics platforms requires sophisticated coordination. Our integrated workflow seamlessly moves your project from initial discovery to final confirmation.
Our Integrated Antigen Discovery and TCR Profiling Services
Mass Spectrometry and SPR Platforms for Immune Profiling
By managing the entire pipeline under one roof, we eliminate the variability and data-loss risks associated with using disjointed vendors for mass spectrometry, genomics, and structural biology.
- Comprehensive Bioinformatic Integration: We do not simply hand over isolated datasets. We perform comprehensive integration using public databases (e.g., VDJdb, IEDB), advanced structural predictions, and clonal frequency metrics to rank probable TCR-peptide associations.
- End-to-End Reagent Sourcing: For the physical validation phase, you do not need to source complex reagents. We provide end-to-end synthesis and expression, generating the required high-purity peptides, recombinant MHC complexes, and soluble TCRs entirely in-house.
- Targeted High-Precision Validation: Rather than wasting resources on massive, low-confidence screening, we strategically filter the data to select highly promising "lead" pairs for low-throughput, high-precision kinetic validation.
Multi-Omics Workflow vs. Single-Platform Analysis
| Dimension | Immunopeptidomics Alone | TCR Sequencing Alone | Interaction Validation Alone | Integrated Workflow |
|---|---|---|---|---|
| Question Answered | What is presented on the cell surface? | Which T cells are expanding? | How strongly does this TCR bind this pMHC? | Which presented peptides drive specific T cell expansion? |
| Primary Output | Peptide sequences | Clonotype frequencies | Binding kinetics (KD) | Verified, functional TCR-pMHC targets |
| Evidence Depth | Antigen presentation | Effector response | Physical biochemistry | Complete immunological loop |
| Strengths | Direct evidence of processing | Captures systemic response | Definitive proof of binding | Bridges discovery to biological mechanism |
| Limitations | Cannot prove T cell reactivity | Cannot identify the target antigen | Requires prior knowledge of the pair | Requires complex coordination and bioinformatics |
| Best-Fit Stage | Early discovery | Immune monitoring | Late-stage preclinical | End-to-end translational research |
Sample Requirements for Immunopeptidomics and TCR Sequencing
| Workflow Stage | Recommended Sample Type | Minimum Input / Quality Considerations | Preferred Format | Key Notes |
|---|---|---|---|---|
| Antigen Discovery | Tumor tissue, cell pellets, matched healthy tissue | Project-specific depending on tissue type and HLA abundance | Snap-frozen (highly preferred) | Sample suitability review available before project initiation. Avoid fixatives. |
| TCR Tracking | PBMCs, TILs, sorted T cells | Sufficient viable cells or high-quality RNA/DNA | Fresh, cryopreserved, or extracted nucleic acids | Bulk TCR and scTCR have distinct viability requirements. Matched samples to discovery input are highly recommended. |
| Interaction Validation | Sequence data only (TCR alpha/beta, Peptide, HLA allele) | High-confidence sequences from discovery phase | Digital sequence data | We synthesize physical reagents in vitro. Feasibility assessment required prior to recombinant expression. |
Example Data for Peptide Discovery and TCR Validation
Our final deliverables provide publication-ready visualizations that clearly communicate the journey from multi-omics discovery to biophysical validation.
Antigen Source Annotation Summary
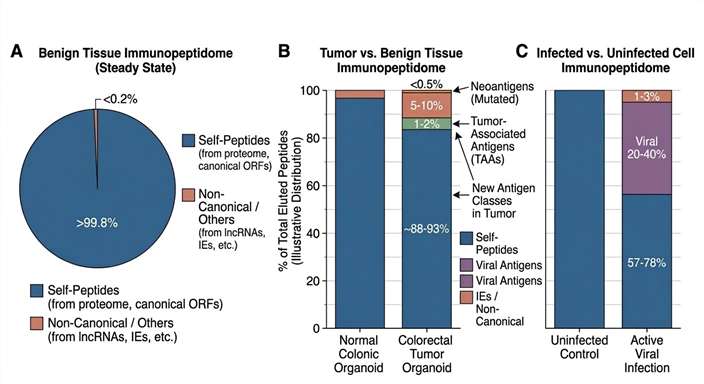
Summary of naturally presented peptides identified via MS-based immunopeptidomics, categorized by their source protein origins to highlight targetable disease-associated antigens.
TCR Clonotype Tracking Visualization
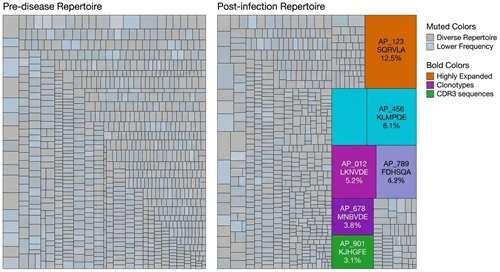
High-resolution TCR sequencing profiling reveals the selective expansion of specific T cell clonotypes, indicating a focused, antigen-driven immune response.
Integrated Interpretation Map
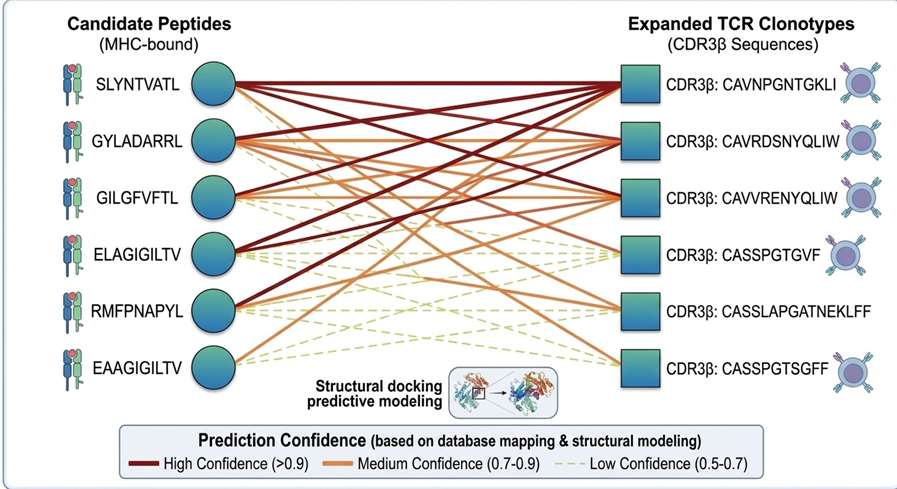
Comprehensive bioinformatic integration correlating identified presented peptides with expanded TCR clonotypes, establishing highly probable targets for downstream validation.
Receptor-Level Interaction Validation
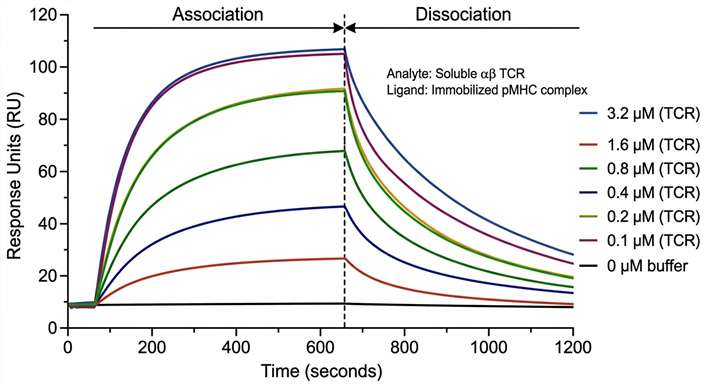
High-precision SPR kinetic validation of a lead candidate pair, confirming the physical binding affinity and stability of the TCR-pMHC interaction.
Deliverables for the Integrated Antigen and TCR Workflow
Because this is an integrated service, you will receive staged data packages culminating in a comprehensive final report:
- Immunopeptidomics Report: Complete list of naturally presented peptides with confidence scores, source protein annotations, and HLA restriction motifs.
- TCR Repertoire Report: Diversity metrics, V(D)J usage, and identification of significantly expanded clonotypes.
- Bioinformatic Integration Report: Cross-referencing of peptide and TCR data utilizing public databases and structural predictions to prioritize top interaction candidates.
- Interaction Validation Data: If the downstream module is selected, raw SPR sensorgrams and calculated kinetic constants (KD, kon, koff) for the selected lead TCR-pMHC pairs.
- Executive Summary: A synthesized research conclusion linking the discovery, tracking, and validation phases into actionable insights for your therapeutic or vaccine pipeline.
Frequently Asked Questions
Representative Literature Example Supporting the Antigen-Discovery-to-Immune-Follow-Up Workflow
Journal: Frontiers in Immunology
Published: 2025
Summary
Using MHC-associated peptide proteomics (MAPPs), researchers identified naturally presented AAV9 capsid peptides and then evaluated whether these peptides could drive measurable T cell responses in human donor PBMCs. Rather than relying only on overlapping peptide libraries, the study focused on peptides that were actually presented on HLA molecules, creating a more biologically relevant route for epitope discovery. By combining immunopeptidomics-derived peptide selection with downstream immune stimulation assays, the authors identified multiple immunodominant CD8 T cell epitopes of the AAV9 capsid, including several previously unreported epitopes. This example study illustrates how naturally presented peptides can guide downstream immune response mapping and candidate validation.
Methods
This study used an integrated antigen-to-response workflow centered on naturally presented peptide discovery.
Researchers first used MAPPs to define the HLA-presented peptide repertoire of the AAV9 capsid. They then compared two peptide strategies for downstream immune testing: conventional overlapping peptide pools and MAPPs-derived peptide pools. Because direct ex vivo detection of capsid-specific T cells in healthy donor PBMCs showed very low sensitivity, the team introduced an in vitro PBMC expansion step before FluoroSpot restimulation assays.
To identify immunodominant CD8 epitopes in greater detail, the HLA Class I MAPPs-derived peptides were further deconvoluted and tested across a donor cohort enriched for predicted binding alleles. Binding assays were then used to confirm whether selected novel epitopes bound stably to their predicted HLA alleles.
Key Technical Features
Sample types: Healthy donor PBMCs, including seropositive and seronegative donors for AAV9
Antigen discovery: MHC-associated peptide proteomics (MAPPs) to identify naturally presented AAV9 capsid peptides
Immune follow-up: PBMC expansion followed by IFN-γ FluoroSpot restimulation assays
Comparison strategy: Overlapping peptide pools versus MAPPs-derived peptide pools
Epitope deconvolution: Screening of 41 HLA Class I MAPPs peptides in matrix pools across 24 healthy donors
Validation: Binding assays to confirm stable interaction of selected epitopes with predicted HLA alleles
Biological insight: Naturally presented MAPPs peptides and overlapping peptides expanded largely distinct T cell populations
Creative Proteomics can offer integrated discovery and follow-up services that support similar research pipelines, including:
- HLA immunopeptidomics for naturally presented antigen discovery
- Peptide pool design and epitope deconvolution strategies
- T cell response profiling using donor PBMC-based functional assays
- TCR repertoire tracking to monitor responsive clonotype expansion
- Peptide–HLA binding validation and downstream interaction analysis
- Integrated interpretation linking presented peptides to immune response prioritization
These services support viral epitope discovery, gene therapy immunogenicity assessment, vaccine target screening, and broader antigen-specific T cell research.
Results
Naturally Presented Peptides Outperformed Generic Peptide Assumptions
The study showed that conventional overlapping peptides and MAPPs-derived peptides did not stimulate the same T cell populations. PBMCs expanded with overlapping peptides responded poorly to MAPPs-derived peptides, while PBMCs expanded with MAPPs-derived peptides responded preferentially to the MAPPs pools. This indicated that naturally presented peptides can reveal antigen-specific responses that are not captured well by standard overlapping peptide designs.
MAPPs-Derived Responses Were Dominated by CD8 T Cells
When donor PBMCs were expanded with the MAPPs-derived capsid peptides, the immune response was dominated by HLA Class I-derived peptides, consistent with a capsid-specific CD8 T cell response. The authors further confirmed this trend through CD4 depletion experiments, supporting the use of naturally presented class I peptides for identifying cytotoxic T cell-relevant targets.
Novel Immunodominant Epitopes Were Identified
By screening 41 HLA Class I MAPPs peptides in 13 matrix pools across 24 healthy donors, the researchers identified 10 epitopes that elicited IFN-γ release in at least one donor. Notably, 9 of these 10 epitopes were novel, and the identified peptides ranged from 9 to 13 amino acids in length.
Binding Validation Strengthened Candidate Confidence
The study reported that the newly identified epitopes displayed strong binding to their predicted HLA alleles, and binding assays confirmed stable interaction for several of the novel candidates. This added an extra validation layer beyond peptide discovery and T cell activation alone.
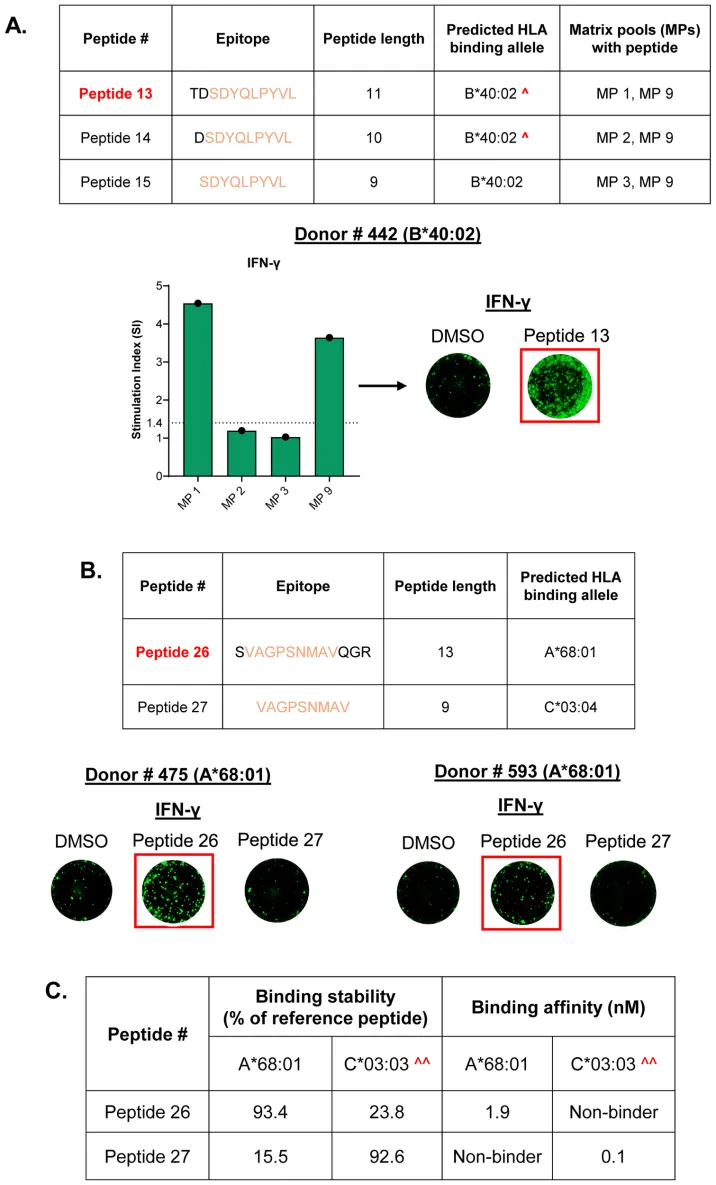
Matrix-based screening of HLA class I peptides derived from immunopeptidomics reveals immunodominant AAV9 capsid epitopes capable of inducing IFN-γ responses in donor PBMCs.
Reference
- Balasubramanian, Akhila, et al. "Epitope mapping using immunopeptidomics reveals novel immunodominant CD8 T cell epitopes of the AAV9 capsid." Frontiers in Immunology (2025). https://doi.org/10.3389/fimmu.2025.1641289