Preclinical Immunogenicity Risk Assessment and MAPPs Principles
Developing therapeutic proteins—including monoclonal antibodies, fusion proteins, and complex bispecifics—involves carefully evaluating the inherent risk of triggering unwanted immune responses. While in silico prediction algorithms offer rapid initial screening by calculating theoretical HLA binding affinities, they fundamentally fail to model the biological complexities of intracellular antigen processing, endosomal trafficking, and enzymatic cleavage competition. High-resolution identification of the actual presented ligands is required to understand the true risk profile of a biologic candidate.
To inform preclinical developability decisions accurately, scientists require presentation-based physical evidence. As a highly specialized application within our broader Immunopeptidomics Service, the MAPPs assay provides exactly this essential data. It reflects physiologically relevant antigen processing within a strictly controlled in vitro primary-cell system. Using primary human monocyte-derived dendritic cells (moDCs), the assay captures naturally processed HLA Class II ligands under optimal experimental conditions. By biochemically isolating these peptide-MHC complexes from the dendritic cells and sequencing the associated ligands via mass spectrometry, researchers can map the specific regions of the therapeutic protein that are physically displayed for potential CD4+ T-cell recognition, effectively bridging the gap between theoretical prediction algorithms and functional immunology.
T-Cell Epitope Mapping Applications in Biologics R&D
Understanding exactly which regions of a therapeutic molecule are successfully processed and presented by dendritic cells enables a wide array of critical biopharmaceutical research applications:
What We Offer: MAPPs Immunogenicity Services
We provide an end-to-end preclinical evaluation service designed to support your biologics pipeline through precise scientific deliverables and highly customized execution:
Advantages of Our MAPPs Immunogenicity Assay
MAPPs Assay Workflow for MHC Class II Peptide Sequencing
Our Integrated Technology Platform
To support this sophisticated workflow with sufficient sensitivity, reproducibility, and peptide recovery depth, the entire assay is executed on our fully integrated analytical platform:
- Ultra-Sensitive Mass Spectrometry: Supported by our Immune Peptide Mass Spectrometry Analysis expertise, we utilize state-of-the-art Orbitrap Eclipse and Exploris 480 instruments for high mass accuracy.
- Advanced Separation: Nano-flow UHPLC systems significantly improve peptide separation, minimizing ion suppression for complex immunopeptidomic samples and rare epitopes.
- Secure Bioinformatics Pipeline: Proprietary algorithms support nested peptide cluster reconstruction and accurate allele-specific association (1% FDR strict filtering).
- Environmental Control: All cell procedures are executed within highly controlled, ultra-clean facilities to eliminate environmental endotoxin interference and maintain moDC integrity.
MAPPs vs. In Silico Prediction vs. T-Cell Assays
| Dimension | In Silico Prediction | MAPPs Assay (LC-MS/MS) | T-Cell Proliferation Assay |
|---|---|---|---|
| Evidence Type | Theoretical probability | Physical in vitro identification | Functional cellular response |
| Primary R&D Value | High-throughput screening | Epitope mapping & ranking | Pre-clinical confirmation |
| False Positive Rate | High (No processing) | Low (Actual presentation) | Moderate (Donor background) |
| Localization | Predicted positions | Precisely mapped sequences | Not provided |
Sample Requirements for Therapeutic Proteins
| Parameter | Specification | Scientific Rationale |
|---|---|---|
| Recommended Amount | 2–5 mg | Ensures sufficient concentration to pulse multiple donor moDC cultures optimally. |
| Concentration | > 1.0 mg/mL | Prevents excessive dilution of cell culture media during pulsing phase. |
| Purity Limit | >95% | Prevents false identification of immunogenic host cell protein (HCP) contaminants. |
| Endotoxin Limit | <0.1 EU/mg | High endotoxin triggers artificial, non-specific maturation of APCs. |
Representative MAPPs Data
Transforming highly complex mass spectrometry raw data into actionable R&D insights. Once regions are identified, you can transition to Immunopeptide Affinity Screening to evaluate binding strengths.
Peptide Sequence Alignment Map
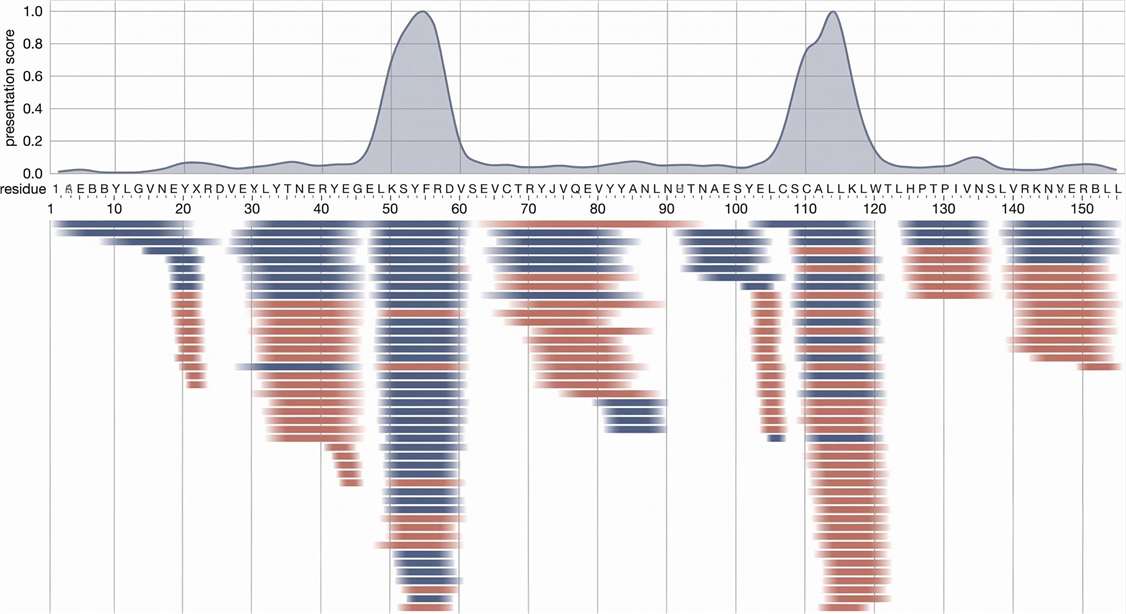
Sequence alignment mapping to pinpoint immunogenic hotspots on a therapeutic protein.
Donor HLA Presentation Matrix
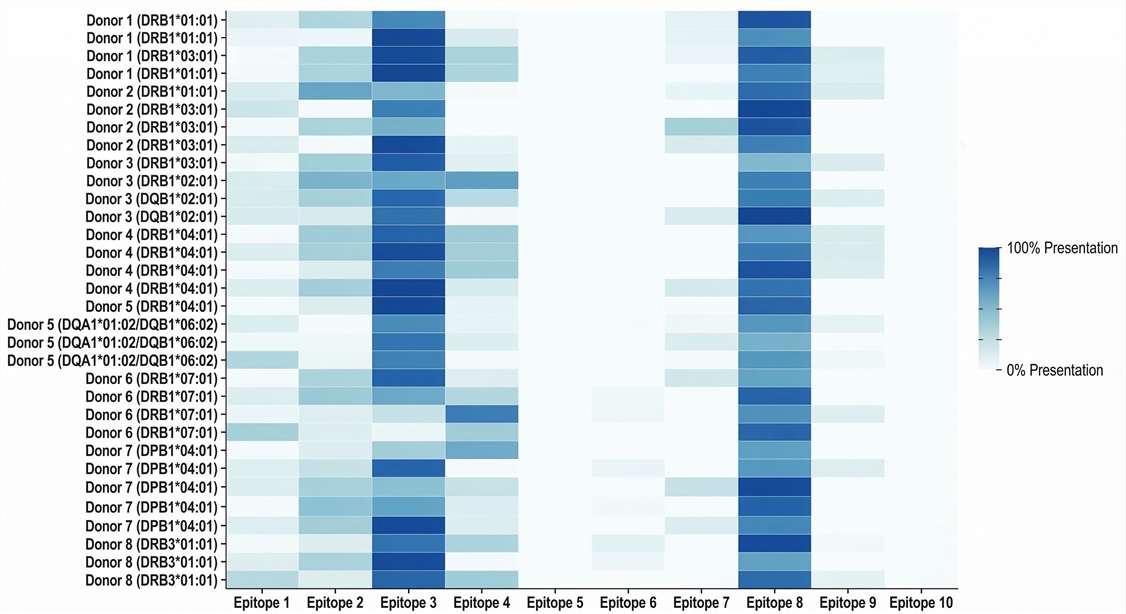
Heatmap illustrating in vitro population-level frequency of identified T-cell epitopes.
Relative Peptide Abundance Chart
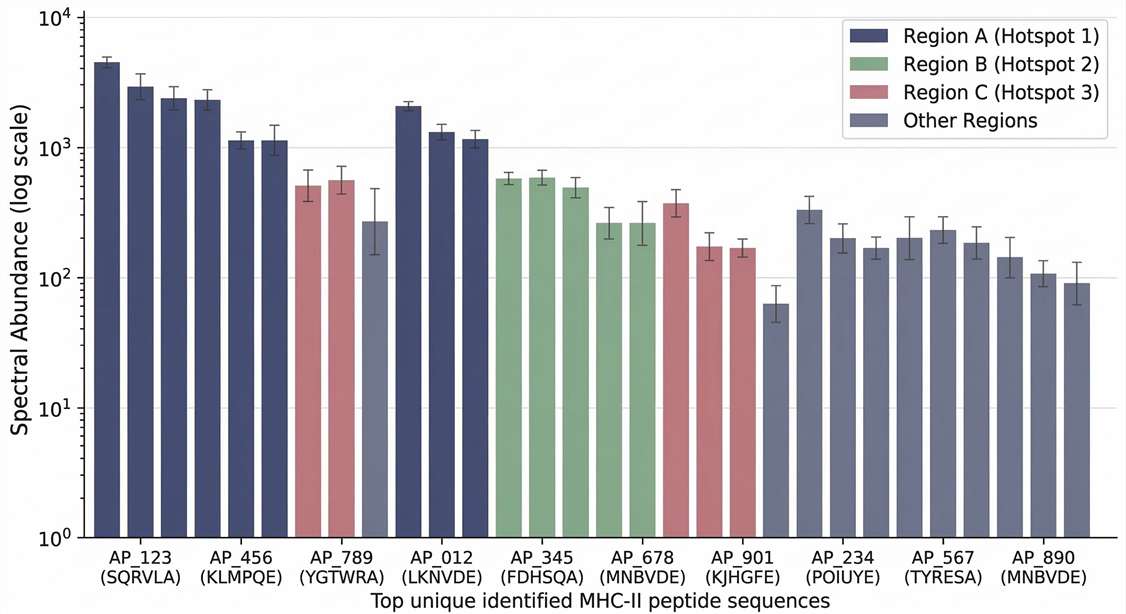
Spectral abundance analysis to evaluate dominant presented immunogenic regions.
Assay Control Validation Plot
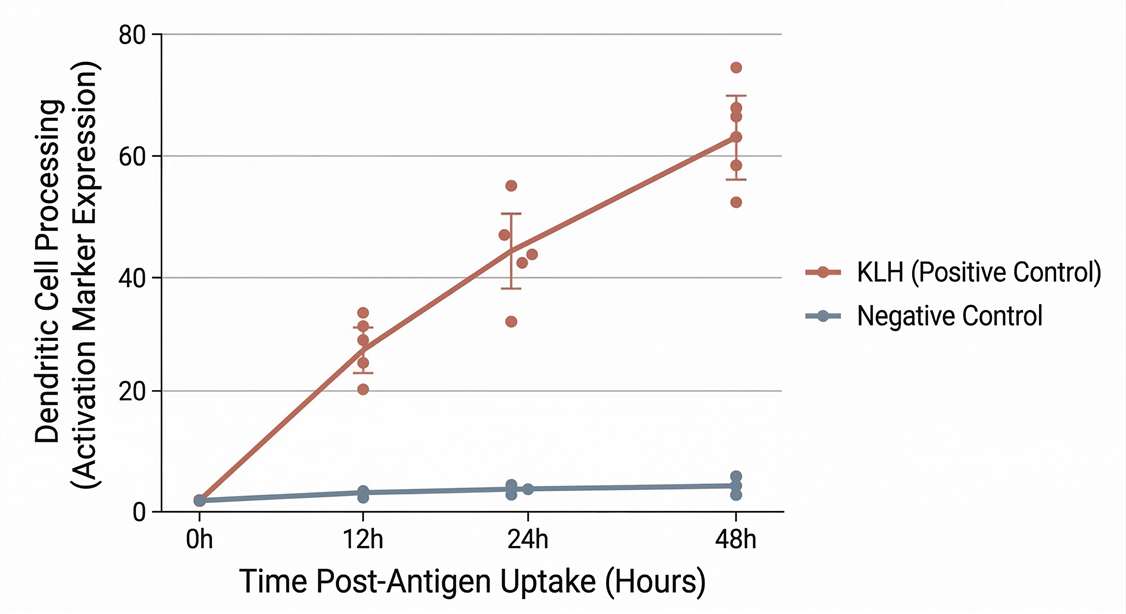
Confirmation of robust dendritic cell processing using the KLH positive reference.
Bioinformatics Analysis
Raw mass spectrometry data requires highly sophisticated interpretation to actively inform protein engineering and safety assessments:
- Peptide Source Annotation: Sequence mapping of HLA Class II peptides back to the therapeutic protein, filtering the endogenous "self" peptidome.
- Hotspot Clustering: Identification and alignment of overlapping fragments to define core minimal T-cell binding regions.
- Relative Presentation Scoring: Integrated evaluation considering spectral abundance and frequency across the donor pool to reliably rank candidates.
Comprehensive Deliverables for Preclinical Evaluation
- Annotated Peptide List: Full inventory of identified peptides including sequences, modifications, and mass spectrometry scores.
- Presentation Hotspot Mapping: Visual identification of therapeutic protein regions physically presented by dendritic cells.
- Donor HLA Metadata: Summary report detailing the HLA-typed donor cohort utilized for the specific assay.
- Executive Summary: Thorough analytical interpretation to directly support internal candidate ranking and de-immunization strategies.
Reference
- Applying MAPPs assays to assess drug immunogenicity. Frontiers in Immunology, 2020. https://doi.org/10.3389/fimmu.2020.00698
- The MHC-associated peptide proteomics (MAPPs) assay is a useful tool to evaluate the immunogenicity of therapeutic proteins. Frontiers in Immunology, 2023. https://doi.org/10.3389/fimmu.2023.1271120
- Assessing MAPPs assay as a tool to predict the immunogenicity potential of protein therapeutics. Life Science Alliance, 2024. https://doi.org/10.26508/lsa.202302095