Multiplex Profiling of Central Appetite and Peripheral Energy Metabolism
Appetite and energy balance are regulated by coordinated signaling networks across adipose tissue, pancreatic endocrine pathways, gut-derived peptides, and central hypothalamic circuits. Evaluating anti-obesity therapeutics or lipolytic agents requires a systemic assessment rather than the measurement of isolated biomarkers.
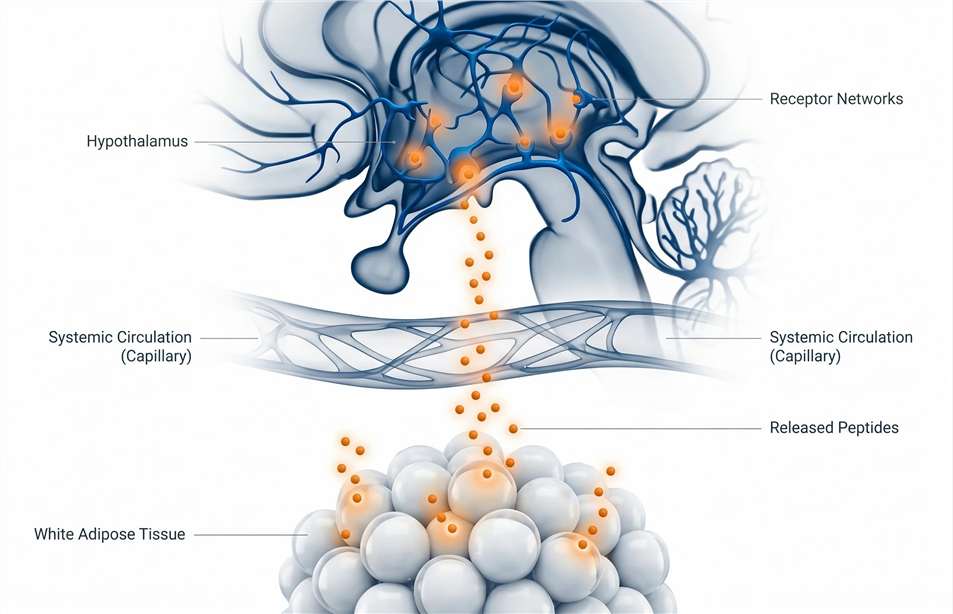
Our Appetite and Energy Metabolism Hormone Panel utilizes targeted liquid chromatography-tandem mass spectrometry (LC-MS/MS) to simultaneously quantify signals that drive hunger (orexigenic neuropeptides), promote satiety (anorexigenic peptides), and regulate basal energy expenditure (adipokines). This approach provides a direct, quantitative readout of the physiological cross-talk between peripheral energy stores and central behavioral control.
Preclinical Applications in Anti-Obesity Drug Discovery
This multiplex panel supports preclinical drug discovery and translational research by providing quantitative bioanalytical data across diverse and rigorous study designs:
Metabolic Hormone Panel Configurations
We offer customizable panel configurations tailored to specific mechanisms of action. Assays are structured into functional biological cores to streamline biomarker selection:
| Biological Core Focus | Targeted Multiplex Analytes | Physiological Role |
|---|---|---|
| Adipokine Core | Leptin, Adiponectin, Resistin, Visfatin | Signals long-term energy reserves; regulates systemic insulin sensitivity and lipid metabolism. |
| Central Appetite Core | AgRP, NPY, α-MSH, Orexin A/B | Hypothalamic neuropeptides driving feeding behavior (orexigenic) or energy expenditure (anorexigenic). |
| Metabolic Homeostasis | Insulin, Glucagon, Amylin, C-peptide, Somatostatin | Pancreatic regulation of glucose utilization, postprandial satiety, and counter-regulatory functions. |
| Peripheral Satiety | PYY (Total & 3-36), Ghrelin (Acylated & Des-acyl) | Gut-derived signals regulating acute meal initiation and termination (delayed gastric emptying). |
Custom Target Integration (PK/PD Co-Analysis)
Our LC-MS/MS assays can be customized to include proprietary anti-obesity peptide therapeutics (e.g., multi-receptor agonists or engineered adipokines). This capability enables the simultaneous monitoring of drug concentration (PK) and endogenous endocrine shifts (PD) within the same sample, reducing animal usage and improving data correlation.
Analytical Platforms: LC-MS/MS vs. Multiplex Immunoassays
Multiplex immunoassays (ELISA/Luminex) are suitable for routine screening of high-abundance markers like total Leptin or Insulin in standard plasma samples. However, targeted LC-MS/MS is required for studies involving low-abundance neuropeptides, active/inactive isoform resolution, complex tissue matrices, or translational cross-species comparisons.
| Analytical Capability | Targeted LC-MS/MS Hormone Panel | Multiplex Immunoassays (ELISA/Luminex) |
|---|---|---|
| Low-Abundance Sensitivity | Quantifies ultra-low abundance central neuropeptides (e.g., AgRP, NPY) via specific PRM/MRM mass transitions. | Often lacks the limits of detection required to capture subtle neuropeptide fluctuations in plasma. |
| Matrix Tolerance | Solid-Phase Extraction (SPE) removes lipid interference from adipose tissue and brain homogenates. | High lipid content (e.g., WAT) frequently causes background noise and antibody interference. |
| Translational Utility | Measures conserved amino acid sequences, directly translating across Mouse, Rat, Canine, and NHP models. | Requires purchasing and validating species-matched commercial kits for each animal model. |
| Isoform Resolution | Distinguishes closely related functional isoforms (e.g., acylated vs. unacylated Ghrelin) by exact molecular mass. | Antibodies frequently cross-react between intact active peptides and partially cleaved fragments. |
Peptide Extraction from Adipose Tissue and Hypothalamic Samples
While plasma provides a systemic overview, evaluating local paracrine mechanisms requires analyzing the source tissues. Complex biological matrices present significant analytical challenges for standard extraction methods.
- Adipose Tissue (WAT/BAT): High lipid content causes severe ion suppression in mass spectrometry. Our protocol utilizes specialized delipidation buffers and matrix-matched Solid-Phase Extraction (SPE) to isolate hydrophilic peptides from the hydrophobic bulk, ensuring reproducible recovery rates in DIO models.
- Hypothalamic Punches: Central regulators (AgRP, NPY) act locally within the arcuate nucleus at low concentrations. We deploy micro-extraction protocols to quench endogenous proteases and recover neuropeptides from milligram-level brain biopsies. This synergizes with our targeted neuropeptide quantification platform.
Cross-Species Translational Assays: Mouse to NHP
Commercial immunoassay kits often lack species-specific antibodies for non-human primate (NHP) or canine models, disrupting the continuity of preclinical obesity research. Targeted mass spectrometry circumvents this limitation by detecting the highly conserved amino acid sequences and exact molecular weights of the target peptides. This sequence-based approach allows researchers to maintain consistent, directly comparable analytical methods from early-stage rodent pharmacology through advanced NHP efficacy evaluations.
Bioanalytical Workflow and Quality Control
To ensure quantitative accuracy, our laboratory implements a standardized multi-step quality control workflow optimized for fragile metabolic peptides.
Sample Requirements for Metabolic Phenotyping
Energy metabolism hormones respond rapidly to nutritional cues. We provide guidance to ensure sampling strategies (e.g., light/dark cycles) align with experimental objectives without inducing artifactual stress responses.
| Matrix Type | Species & Model Context | Min Volume / Mass | Pre-Analytical Stabilization Notes |
|---|---|---|---|
| Plasma / Serum | DIO Mice, NHP, Clinical Cohorts. | 50 - 100 μL | Requires immediate cold centrifugation and protease/esterase inhibitors. |
| Adipose Tissue (WAT/BAT) | Thermogenesis, tissue-specific knockout models. | 30 - 50 mg | Snap-freeze in liquid N2. Do not use PBS wash. |
| Hypothalamic Punches | Central appetite regulation mapping. | 10 - 20 mg | Snap-freeze immediately upon regional brain extraction. |
| Cell Culture Media | Adipocyte cell lines, Hypothalamic slice cultures. | 100 - 200 μL | Add inhibitors prior to freezing for secretagogue screening. |
Data Deliverables and QC Metrics
Standard data deliverables are tailored to evaluate pharmacological target engagement, efficacy, and systemic metabolic shifts. Our comprehensive bioanalytical reports provide direct statistical insights:
- Validation & QC Report: Details analytical metrics including the Lower Limit of Quantification (LLOQ), assay dynamic range, intra-/inter-batch Coefficient of Variation (CV%), and spike-and-recovery rates for complex matrices.
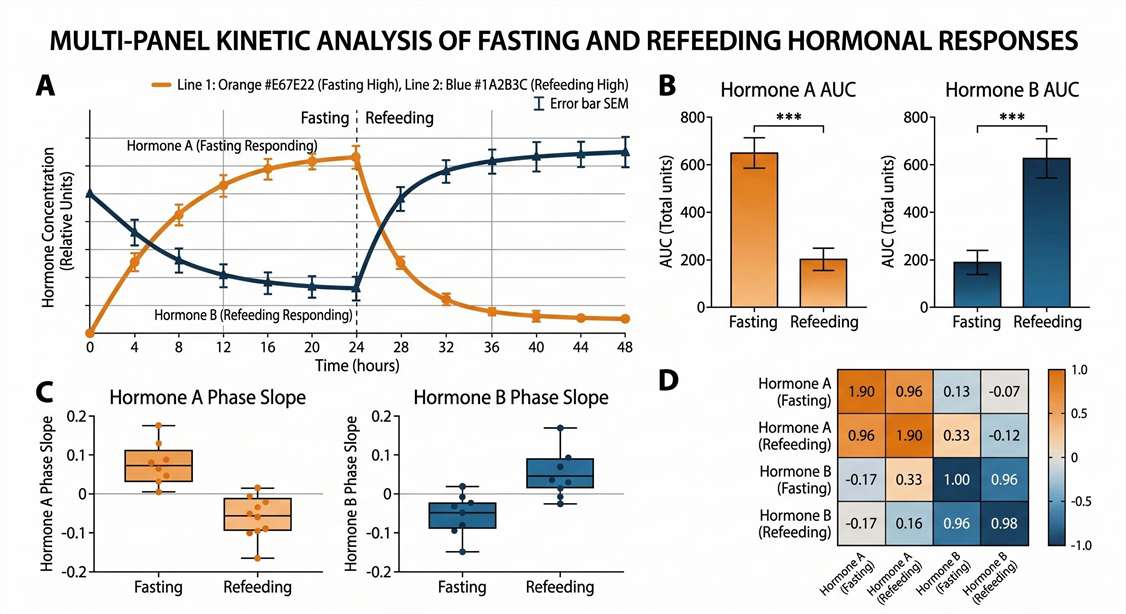
- Kinetic Curves (Feeding vs. Fasting): Visual representation of reciprocal shifts in orexigenic (e.g., active Ghrelin) and anorexigenic (e.g., Leptin) hormones over specific time courses.
- Tissue-to-Plasma Partitioning: Data matrices displaying concentration gradients between local tissue reservoirs and systemic circulation.
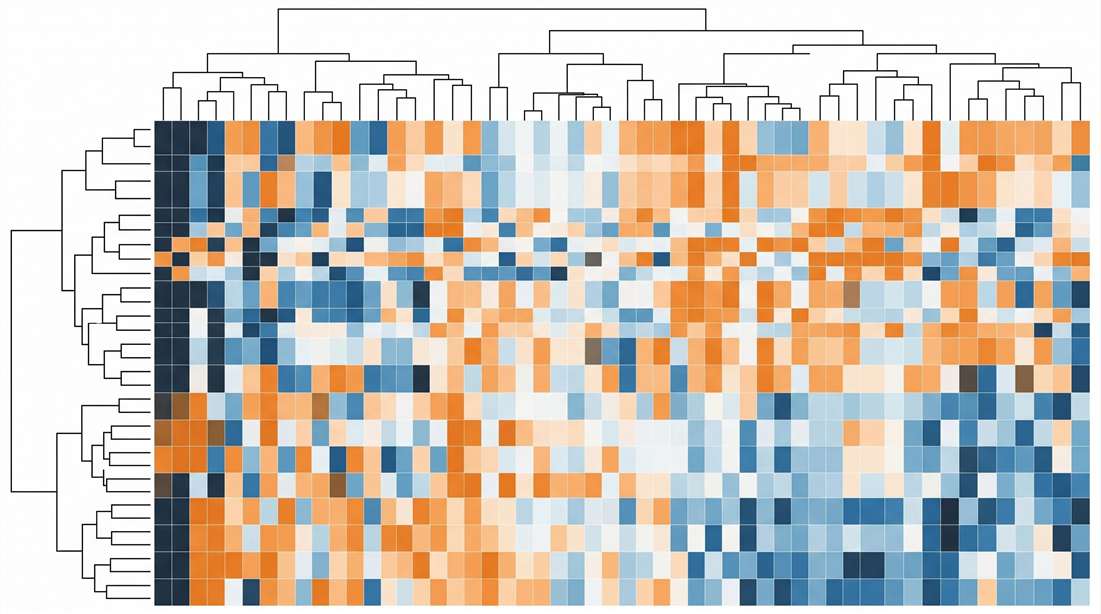
- Metabolic Heatmaps: Color-coded matrices comparing multi-hormone network regulation between vehicle-treated, drug-treated, and control groups.
- Absolute Concentration Matrix: Raw and normalized concentration tables (pg/mL or pM) formatted directly for biostatistical software import.
Demo Results
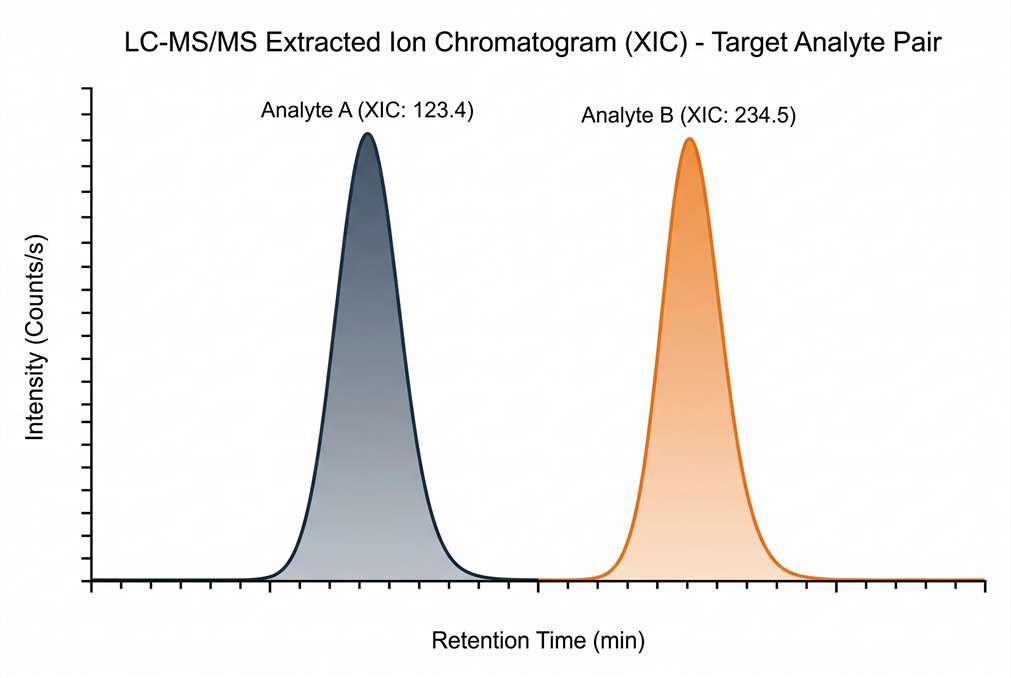
Absolute Quantification Standard Curve
Feeding vs. Fasting Hormone Kinetics
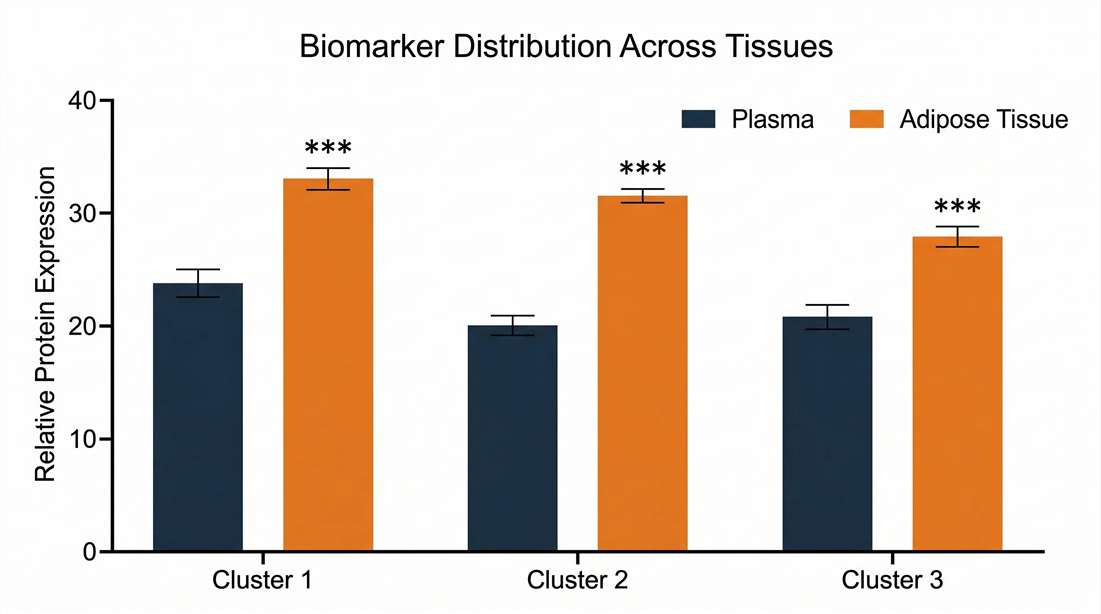
Adipose Tissue to Plasma Partitioning
Metabolic Hormone Heatmap (Lean vs. DIO)
Disclaimer: All services and analytical platforms described are intended for translational research and preclinical support. Research Use Only (RUO). Not for use in diagnostic procedures.