Immune Peptide Library Screening Overview
Immune peptide library screening is a structured method for testing many peptides in parallel to identify sequences associated with immune recognition. In epitope discovery workflows, it is commonly used to evaluate overlapping peptide libraries, focused candidate panels, viral peptide sets, tumor antigen panels, or custom peptide collections generated from computational prediction or immunopeptidomics.
Instead of testing every peptide one by one at the start of a project, researchers can screen peptide pools in a scalable format, detect positive immune responses, and then deconvolute those responses to identify the most relevant candidate peptides for downstream confirmation.
This makes peptide library screening especially useful when:
- the candidate list is too large for single-peptide testing
- sample volume is limited
- antigen-wide coverage is required
- predicted candidates need biological prioritization
- immunopeptidomics or in silico results need functional follow-up
When to Use Immune Peptide Library Screening for Epitope Discovery
Peptide library screening is often the right choice when researchers need to narrow a broad candidate space into a manageable shortlist.
Typical use cases
- T cell epitope mapping across full-length antigens
- Overlapping peptide screening for viral or bacterial proteins
- Screening predicted HLA Class I or HLA Class II binders
- Neoantigen panel prioritization in immuno-oncology
- PBMC-based immune monitoring against custom peptide pools
- Functional follow-up after immunopeptidomics-based antigen discovery
When another approach may be better
- If you already have only a few high-confidence peptides, single-peptide validation may be more efficient.
- If your main question is which peptides are naturally presented in vivo, immunopeptidomics may be the better starting point.
- If your main goal is to quantify binding kinetics, SPR or BLI is usually more appropriate after shortlist generation.
Immune Peptide Library Screening for T Cell Epitope Mapping
T cell epitope mapping depends on both library design and assay strategy. Our immune peptide library screening platform supports projects based on:
- HLA Class I-associated peptide screening for CD8+ T cell epitope discovery
- HLA Class II-associated peptide screening for CD4+ T cell epitope discovery
- Overlapping peptide libraries for broad antigen coverage
- Focused peptide panels derived from prediction, immunopeptidomics, or prior literature
By pairing structured peptide pooling with functional immune assays, researchers can identify responsive pools and then localize likely hit peptides through matrix deconvolution or follow-up confirmation.
Applications of Peptide Library Screening in Vaccine, Infection, and Immuno-Oncology Research
Immune peptide library screening supports a wide range of translational and discovery programs.
What We Offer in Immune Peptide Library Screening
Our peptide library screening service is designed to support both broad discovery and focused validation workflows.
Advantages of Our Immune Peptide Library Screening Service
Our platform is built to help researchers move from large peptide sets to decision-ready epitope candidates with greater efficiency, flexibility, and downstream usability.
Peptide Library Design and Screening Options
Project design can be tailored based on antigen type, candidate source, assay format, and sample constraints.
| Library Format | Target Application | Key Design & Architecture Considerations |
|---|---|---|
| Overlapping Peptide Libraries | Antigen-wide coverage | Peptide length (e.g., 15-mers), overlap strategy (e.g., 11 aa), total antigen size |
| Focused Predicted Epitope Panels | Shortlist refinement | HLA Class I vs Class II focus, total candidate count, algorithm thresholds |
| Viral Peptide Libraries | Pathogen antigen screening | Strain coverage, inclusion of conserved vs. hypervariable regions |
| Tumor/Neoantigen Panels | Immuno-oncology workflows | Mutation inclusion strategy, patient-specific vs. shared TAA screening |
| Custom Peptide Collections | Flexible study design | Peptide purity requirements, readout platform compatibility, solubility limits |
Exact design parameters should be confirmed case by case based on project goals and assay compatibility.
Immune Peptide Library Screening Workflow
Our workflow is built to move efficiently from peptide design to prioritized epitope shortlist.
Immune Peptide Screening Technology Platform
Our immune peptide library screening workflow can support a range of functional and analytical approaches depending on the scientific question.
| Platform Module | Core Technologies & Assays | Primary Objective |
|---|---|---|
| Upstream Data Integration | Computational epitope prediction, MS-based immunopeptidomics datasets, literature mining | Inform logical library design and refine initial candidate selection logic |
| Functional Immune Readouts | IFN-γ ELISpot, FluoroSpot, Flow cytometry (ICS / T cell activation markers), PBMC-based assays | Quantify T cell activation, map cellular responses, and detect active pools |
| Downstream Validation & Integration | SPR / BLI affinity characterization, HLA-binding evaluation, single-peptide confirmation | Provide biophysical confirmation of shortlisted hits and rank candidates |
Sample Requirements for Immune Peptide Library Screening
Input requirements depend on whether the project uses a newly designed peptide library or a pre-existing peptide panel.
| Project Type | Typical Input | Required or Optional | Purpose | Notes |
|---|---|---|---|---|
| Overlapping peptide library project | Target antigen or protein sequence | Required | Full-sequence epitope mapping | Library design defined per project |
| Focused peptide screening | Candidate peptide list | Required | Targeted prioritization | Sequence list should be provided |
| Viral library screening | Pathogen antigen list or sequence scope | Required | Vaccine or infection-related studies | Antigen boundaries should be confirmed |
| Tumor antigen or neoantigen screening | Candidate peptide panel and source context | Required | Immuno-oncology screening | Mutation annotation may be useful |
| Client-provided peptide collection | Synthetic peptide panel | Optional | Custom screening | Quality and format review recommended |
Note: Exact peptide counts, pool sizes, sample requirements, and assay compatibility should be defined case by case.
Immune Peptide Library Screening vs Other Epitope Discovery Methods
| Method | Discovery Breadth | Throughput | Typical Output | Best Fit Stage | Main Value |
|---|---|---|---|---|---|
| Immune Peptide Library Screening | Broad to targeted | High | Positive pools, deconvoluted hits, ranked candidates | Early to mid-stage | Efficient narrowing of large candidate sets |
| Single-Peptide Validation | Narrow | Low to medium | Confirmed function or binding for selected peptides | Mid to late-stage | Confirmation of top candidates |
| Immunopeptidomics Discovery | Unbiased endogenous landscape | Medium | Naturally presented peptide sequences | Discovery stage | Identification of physiologically presented ligands |
| SPR / BLI Affinity Validation | Very narrow | Low | Quantitative binding kinetics | Late-stage validation | Biophysical confirmation of shortlisted hits |
Selection guidance: Use immune peptide library screening when you need to efficiently reduce a large peptide space into a smaller set of actionable candidates. Use single-peptide assays when your shortlist is already small. Use immunopeptidomics when the key discovery question is natural peptide presentation. Use SPR or BLI when affinity characterization is the primary objective.
Example Results from Immune Peptide Library Screening
Functional response readout (ELISpot array)
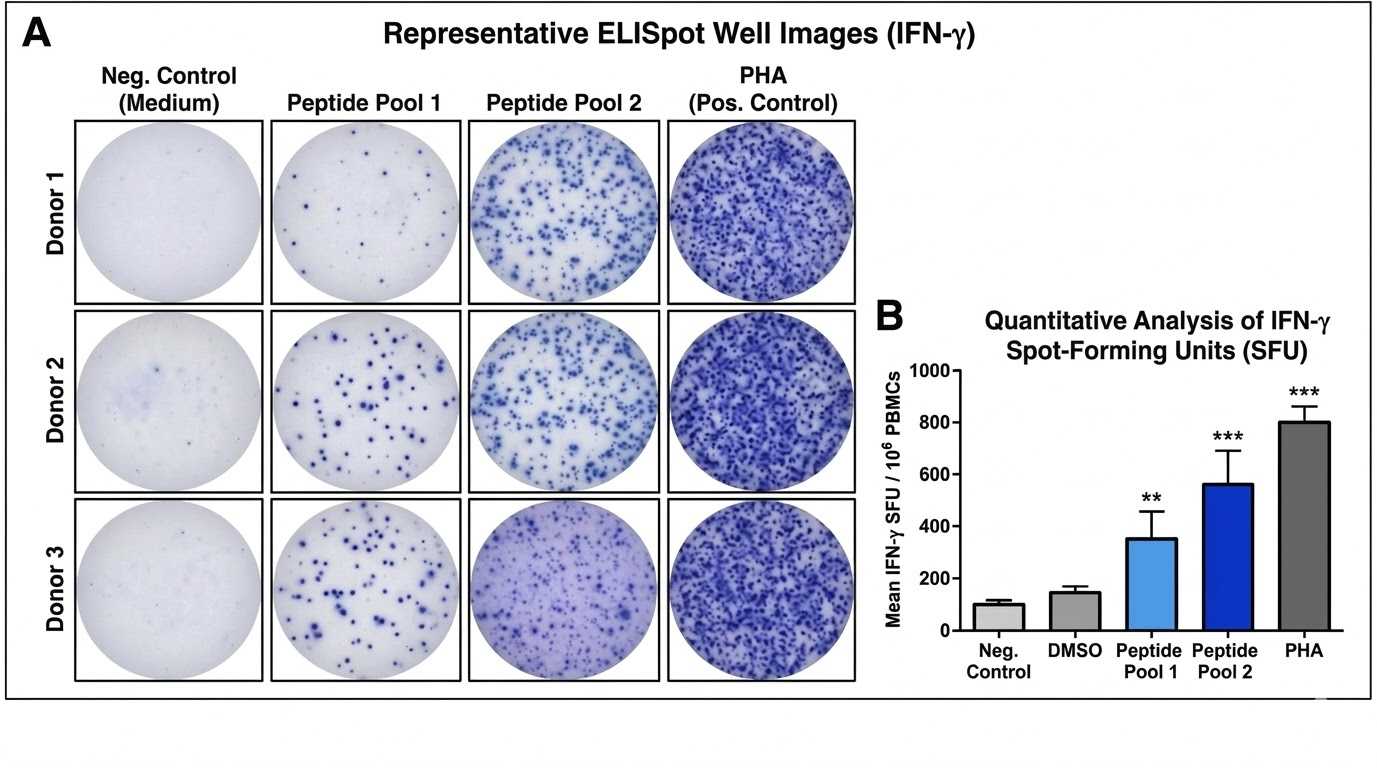
Representative ELISpot results showing differential T cell activation across peptide pools and identifying pools associated with elevated IFN-gamma responses.
Matrix pool deconvolution heatmap
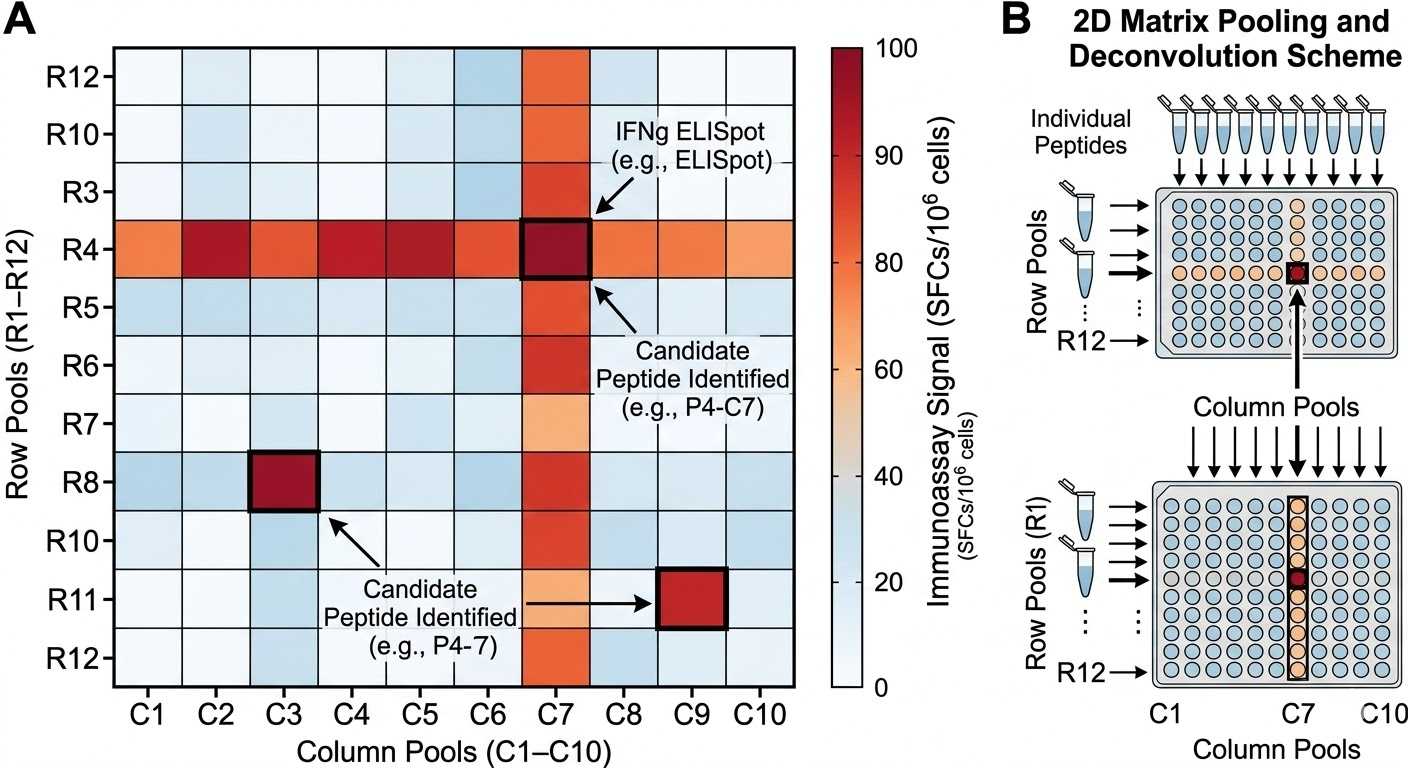
Matrix pool deconvolution heatmap illustrating how positive pool intersections can be used to localize likely hit peptides within a larger screening library.
Antigen coverage and epitope hotspot map
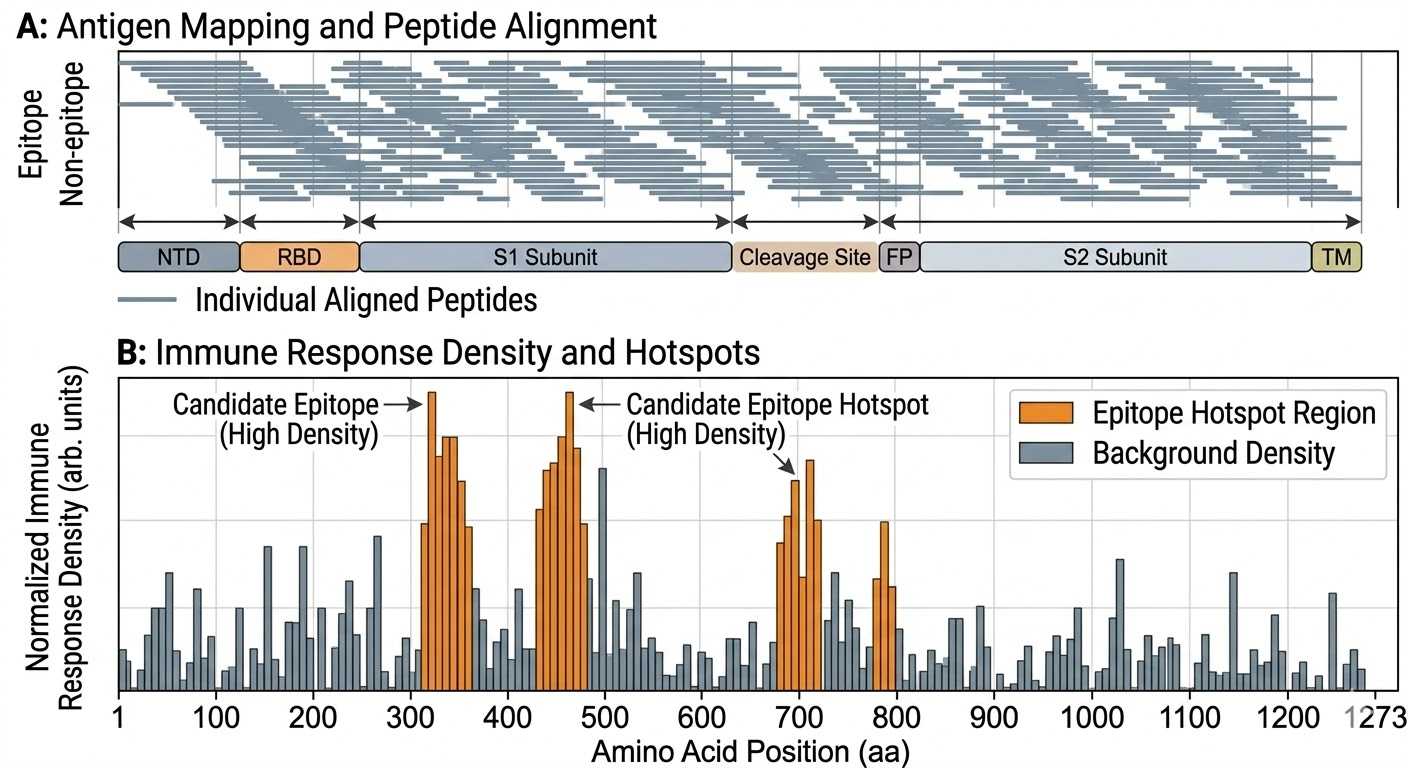
Antigen-wide coverage view showing overlapping peptide distribution and response-enriched regions for epitope hotspot identification.
Candidate prioritization plot
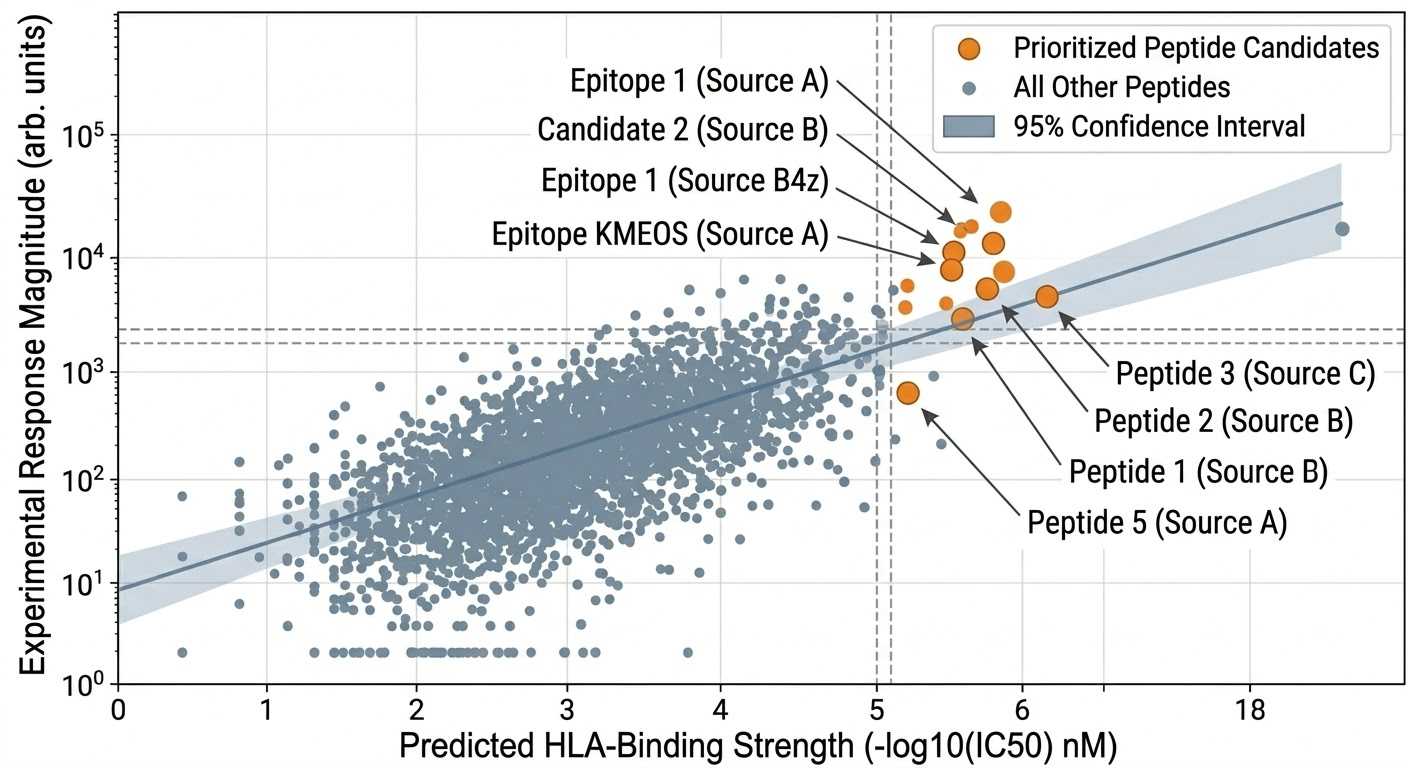
Candidate prioritization plot integrating functional response data with predicted HLA-binding information to guide downstream validation decisions.
Deliverables from Immune Peptide Library Screening Projects
Our deliverables are designed to help researchers move directly into follow-up validation and decision-making.
Standard deliverables
- Screening report with methodology summary, assay overview, and QC-related observations
- Raw or summarized assay readouts depending on project scope
- Pool response interpretation and deconvolution mapping where applicable
- Ranked candidate hit list with prioritized peptide sequences
- Next-step recommendations for single-peptide validation or downstream characterization
Optional downstream support
- single-peptide rescreening
- affinity validation planning
- candidate refinement for subsequent assay stages
- integration with related antigen discovery workflows
Frequently Asked Questions About Immune Peptide Library Screening
Case Study: Matrix Pool Screening and Deconvolution for Murine MHC Class I Epitope Discovery
Journal: Frontiers in Immunology
Published: 2022
Summary
Using a matrix-based peptide library screening workflow, researchers systematically identified murine MHC class I-restricted T cell epitopes from the recombinant oncolytic virus VSV-GP. Rather than testing each candidate peptide individually from the outset, the study combined epitope prediction, matrix pool design, IFN-γ ELISpot screening, and follow-up deconvolution to efficiently narrow a large candidate set into validated epitope hits. This approach enabled the discovery of multiple H2-Db- and H2-Kb-presented viral epitopes and provides a strong literature-backed example of how peptide library screening can reduce assay burden while preserving resolution for downstream validation.
Methods
This study used an integrated prediction-to-validation workflow for murine MHC class I epitope discovery.
Researchers first ranked candidate epitopes computationally and arranged them into a peptide candidate matrix consisting of horizontal and vertical pools. This matrix design enabled the screening of 50 H2-Db and 50 H2-Kb predicted epitopes following VSV-GP immunization in C57BL/6J mice.
One week after immunization, mouse splenocytes were harvested and stimulated with peptide pools in an IFN-γ ELISpot assay to identify response-positive pools. Matrix deconvolution was then applied to trace positive pool signals back to likely individual peptide candidates. These selected peptides were subsequently tested one by one, and intracellular cytokine staining (ICS) was used as an additional validation step to confirm peptide-induced IFN-γ+ CD8+ T cell responses.
Key Technical Features
- Sample types: Splenocytes isolated from VSV-GP-immunized and mock-treated C57BL/6J mice
- Candidate design: Computationally ranked murine MHC class I peptide candidates
- Pooling strategy: Matrix pool design with horizontal and vertical peptide pools
- Screening scale: 50 H2-Db and 50 H2-Kb predicted epitopes screened through pooled testing
- Functional readout: IFN-γ ELISpot for pool screening and individual peptide follow-up
- Hit deconvolution: Matrix-based elimination of non-significant pools to identify likely single-peptide hits
- Orthogonal validation: Intracellular cytokine staining to confirm IFN-γ+ CD8+ T cell responses
Creative Proteomics can offer immune peptide library screening services that support similar research workflows, including:
- Overlapping or focused peptide library design for epitope discovery
- Matrix pooling strategies to reduce assay burden and preserve deconvolution logic
- ELISpot or FluoroSpot-based functional screening of peptide pools
- Single-peptide follow-up testing for hit confirmation
- Downstream ICS, HLA-binding, or affinity validation planning
- Integration of prediction, screening, and prioritization into one structured discovery workflow
These services support vaccine epitope mapping, viral antigen screening, immuno-oncology research, and broader antigen-specific T cell studies.
Results
Matrix Pool Screening Reduced the Complexity of Large Candidate Sets: By organizing predicted epitopes into structured peptide matrices, the researchers were able to screen 100 candidate peptides efficiently without testing each sequence individually at the start. This demonstrates the practical value of peptide pooling for large-scale epitope discovery workflows.
ELISpot Screening Identified Response-Positive Peptide Pools: Following VSV-GP immunization, IFN-γ ELISpot analysis revealed significant T cell activation in multiple H2-Db and H2-Kb peptide pools compared with mock controls. These positive pools provided the basis for matrix deconvolution and candidate narrowing.
Deconvolution Enabled Identification of Individual Epitope Hits: After deconvolution and single-peptide testing, the study identified a total of 20 VSV-GP T cell epitopes presented by murine MHC-I alleles H2-Db and H2-Kb, including 5 H2-Db epitopes and 15 H2-Kb epitopes. This illustrates how pooled screening can move from broad candidate coverage to actionable peptide-level results.
ICS Added an Orthogonal Validation Layer: Intracellular cytokine staining further confirmed that a subset of the identified peptides induced significant IFN-γ+ CD8+ T cell responses. This strengthened confidence that the screening hits were not only pool-positive but also biologically relevant at the single-peptide level.
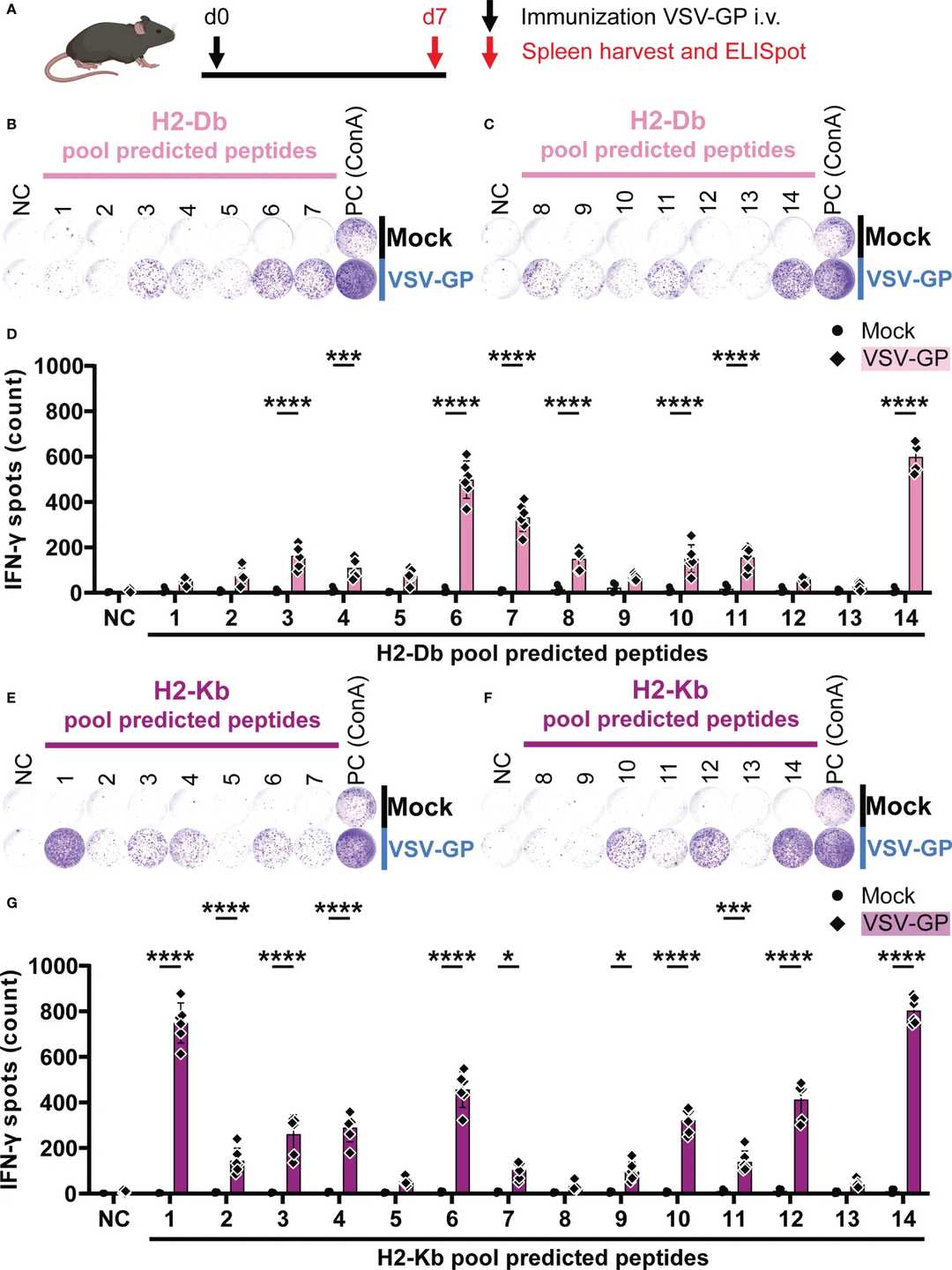
IFN-γ ELISpot-based peptide pool screening identifies response-positive H2-Db and H2-Kb peptide pools and supports matrix-guided deconvolution of candidate murine MHC class I epitopes.
Reference
- Vijver, Saskia V., et al. "Prediction and validation of murine MHC class I epitopes of the recombinant virus VSV-GP." Frontiers in Immunology 13 (2022). https://doi.org/10.3389/fimmu.2022.1100730