Neuropeptide PTM and Processing Characterization Strategy
In neurobiology and peptide therapeutics, the primary sequence is merely the starting point. The true biological activity, stability, and receptor affinity of a peptide are dictated by its precise maturation from a larger pro-hormone and the subsequent addition of post-translational modifications (PTMs) such as C-terminal amidation, pyro-glutamylation, or sulfation.
Standard tryptic proteomics inherently erases these endogenous signatures. As a dedicated partner in translational research, our integrated analysis of neuropeptidomics is built on a "Mechanism-First" philosophy. By combining discovery-based profiling with site-specific targeted quantification, we empower pharmaceutical chemists and academic PIs to seamlessly track peptide maturation, accurately quantify PTM occupancy, and unravel the functional processing mechanisms critical for drug development.
When to Use Neuropeptide PTM and Processing Characterization
Our specialized characterization platform is strategically designed for researchers and R&D directors facing specific biological and technical roadblocks:
Neuropeptide Processing and PTM Characterization
The functional identity of a neuropeptide is defined by two integrated events: the precise proteolytic cleavage from a pro-hormone, and the addition of specific PTMs.
Precursor Cleavage Mapping
Mapping a fragmented peptide back to its exact cleavage site on a pro-hormone is a complex bioinformatics challenge. Generic algorithms often confuse ex vivo artificial degradation with true enzymatic maturation. Our peptide variant analysis service utilizes 1% FDR-controlled alignment to reconstruct the true processing pathway, enabling us to:
- Distinguish Cleavage Types: Accurately differentiate between physiological prohormone convertase (PC) cleavages (typically at basic residues like KR/RR) and non-specific sample degradation.
- Cross-Species Conservation: Evaluate the evolutionary conservation of processing sites across animal models and human cohorts to ensure translational relevance.
- Identify Maturation Intermediates: Track transient, low-abundance processing intermediates to reveal the step-by-step maturation dynamics of complex neuroendocrine peptides.
High-Confidence PTM Identification
Many critical neuropeptide modifications are challenging to detect because they are prone to neutral loss during MS fragmentation or exist in extremely low stoichiometry. Our platform utilizes PTM-aware identification algorithms combined with optimized fragmentation modes (HCD/ETD) to secure high-confidence neuropeptide PTM identification for:
- C-terminal Amidation: Essential for GPCR activation and protection against carboxypeptidases.
- N-terminal Pyro-glutamate: Prevents aminopeptidase-mediated degradation.
- Tyrosine Sulfation: Critical for modulating binding affinity in specific peptide families.
- Disulfide Bond Mapping: Resolving complex folding in disulfide-rich neuropeptides.
Workflow and Quantitative Characterization Capabilities
Our platform integrates stringent pre-analytical stabilization with ultra-high-resolution mass spectrometry to capture the intact peptidome state without relying on trypsin digestion.
Endogenous peptides have half-lives of mere minutes ex vivo, making standard extraction protocols obsolete. We prioritize the "Molecular Lockdown":
- Pre-Analytical Stabilization: Proprietary Rapid Thermal Inactivation and specific protease-quenching cocktails lock PTMs and prevent artifactual cleavage immediately upon collection.
- Go/No-Go QC Checkpoint: Every sample undergoes strict pre-assessment (including hemolysis index and degradation metrics) to protect your research budget.
- Low Molecular Weight Enrichment: Utilizing optimized Molecular Weight Cut-Off (MWCO) and Solid-Phase Extraction (SPE) to isolate the intact <10 kDa functional peptidome.
Data acquisition is performed on the Orbitrap Astral™ or timsTOF Pro platforms. These high-resolution mass spectrometers scan a vast dynamic range to capture trace neuropeptides, enabling us to calculate rigorous localization probabilities (e.g., A-score/P-score) to definitively pinpoint challenging modifications.
Identifying a PTM is necessary, but quantifying its biological prevalence is critical for translational decision-making. We offer distinct quantification strategies to measure the exact ratio of modified to unmodified peptides (PTM Occupancy):
- Label-free Discovery: For broad screening of modified vs. unmodified abundance in discovery cohorts.
- TMT Labeling: For high-precision relative quantification across multiplexed samples.
- Targeted PRM/MRM: For absolute occupancy quantification and validation of critical therapeutic leads using targeted mass spectrometry quantitation.
Platform Technical Benchmarks
- Micro-Peptide Capture: Proven capability to reliably detect and sequence <10 amino acid short functional peptides that traditional workflows miss.
- Low-Input Compatibility: Engineered for precious specimens, requiring as little as ≤10 mg of specific brain region microdissections.
- High Reproducibility: Automated liquid handling and internal standard normalization yield intra-batch CVs typically <15%.
Choosing the Right PTM and Processing Characterization Strategy
Selecting the appropriate analytical depth is critical for project success and budget efficiency. Use the following matrix to align your research goals with our specialized strategies:
| Research Scenario | Recommended Strategy | Primary Output Metric |
|---|---|---|
| Novel Precursor Discovery | Discovery-based PTM Profiling | Maturation Map & PTM Inventory |
| Validating Known Active PTMs | Targeted PTM Characterization | Absolute PTM Occupancy (%) |
| Processing Enzyme Screening | Cleavage Site & Intermediate Mapping | Processing Efficiency Index |
| Peptide Drug SAR Optimization | PTM & Stability Correlation | Stability-Activity Relationship Data |
Why Standard Proteomics and Immunoassays Miss Neuropeptide Processing
Relying on standard tryptic proteomics or commercial ELISAs for neuropeptide research often leads to incomplete or misleading biological conclusions.
| Feature | MS PTM & Processing Characterization | Standard Trypsin Proteomics | Immunoassays (ELISA) |
|---|---|---|---|
| Cleavage Site Resolution | Exact. Pinpoints physiological termini. | Lost. Trypsin erases maturation sites. | None. Blind to cleavage position. |
| PTM Occupancy | Direct. Measures % modified ratio. | Ignored. Sub-stoichiometric PTMs missed. | Poor. Cross-reacts with isoforms. |
| Short Peptide Capture | High. Optimized for <10 aa fragments. | Low. Small peptides lost in wash steps. | Variable. Limited by epitope. |
Applications in Peptide Drug Development and Enzyme Screening
Our characterization platform acts as a critical readout for biopharma pipelines, seamlessly bridging the gap between structural biology and in vivo efficacy.
- Peptide Stability & Half-Life Optimization: Mapping metabolic "soft spots" and evaluating the protective effects of engineered PTMs.
- Target Engagement & Receptor Affinity: Verifying that the mature, modified form of a peptide is correctly produced and stable in the target tissue.
- Processing Enzyme Screening: Evaluate the efficacy of novel inhibitors targeting processing enzymes (such as PAM, PC1/3) by directly quantifying the downstream blockade of neuropeptide maturation.
Demo Results and Deliverables for PTM and Processing Analysis
We translate multidimensional MS data into comprehensive, publication-ready visual deliverables that answer your core functional questions.
Precursor Cleavage Map
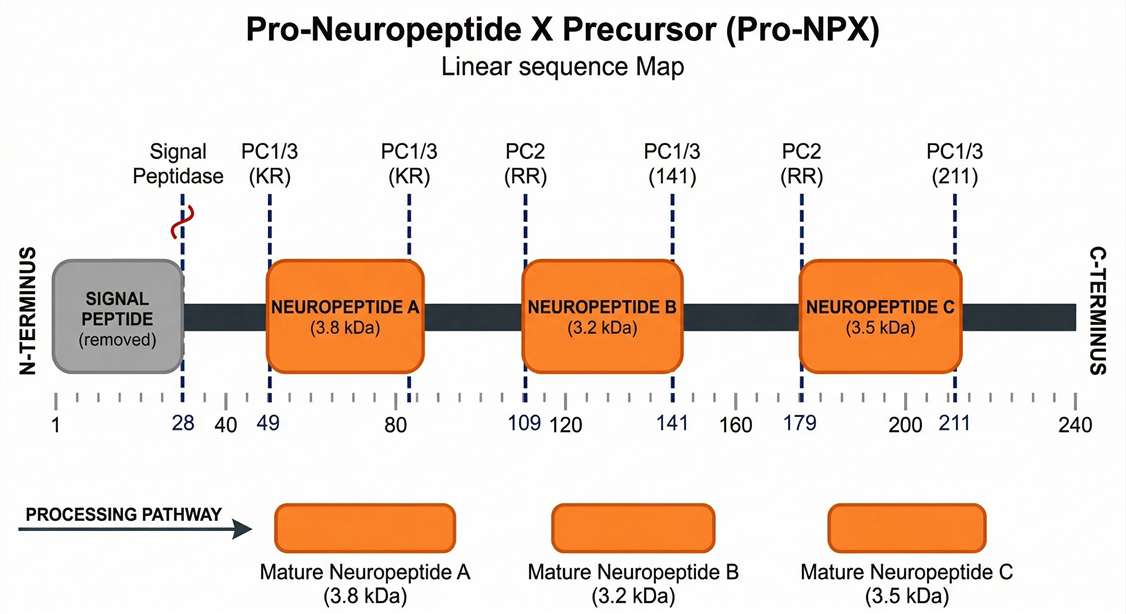
A horizontal pro-hormone alignment displaying all identified mature peptide segments and processing sites.
Processing Pathways
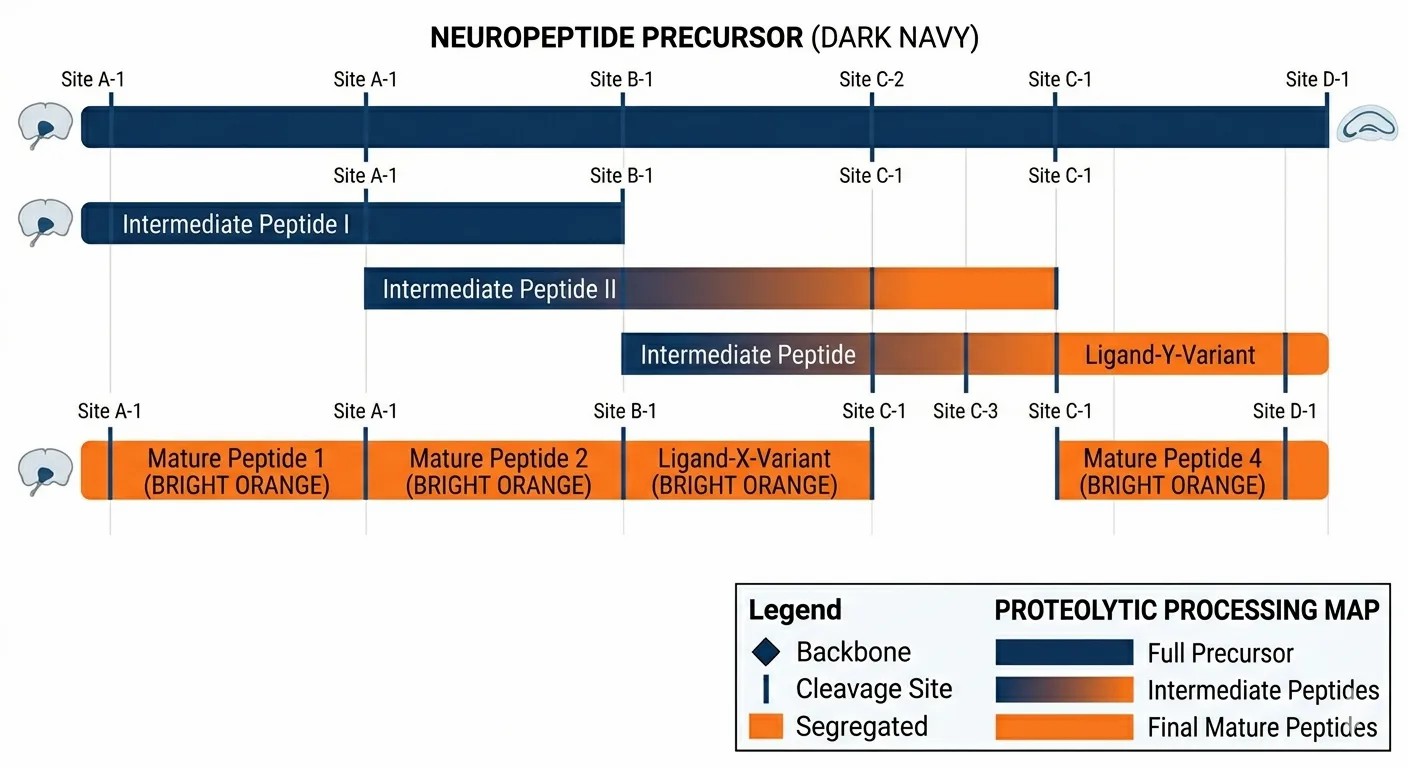
A nested visualization revealing the stepwise generation of mature peptides via intermediate precursors.
MS/MS PTM Validation
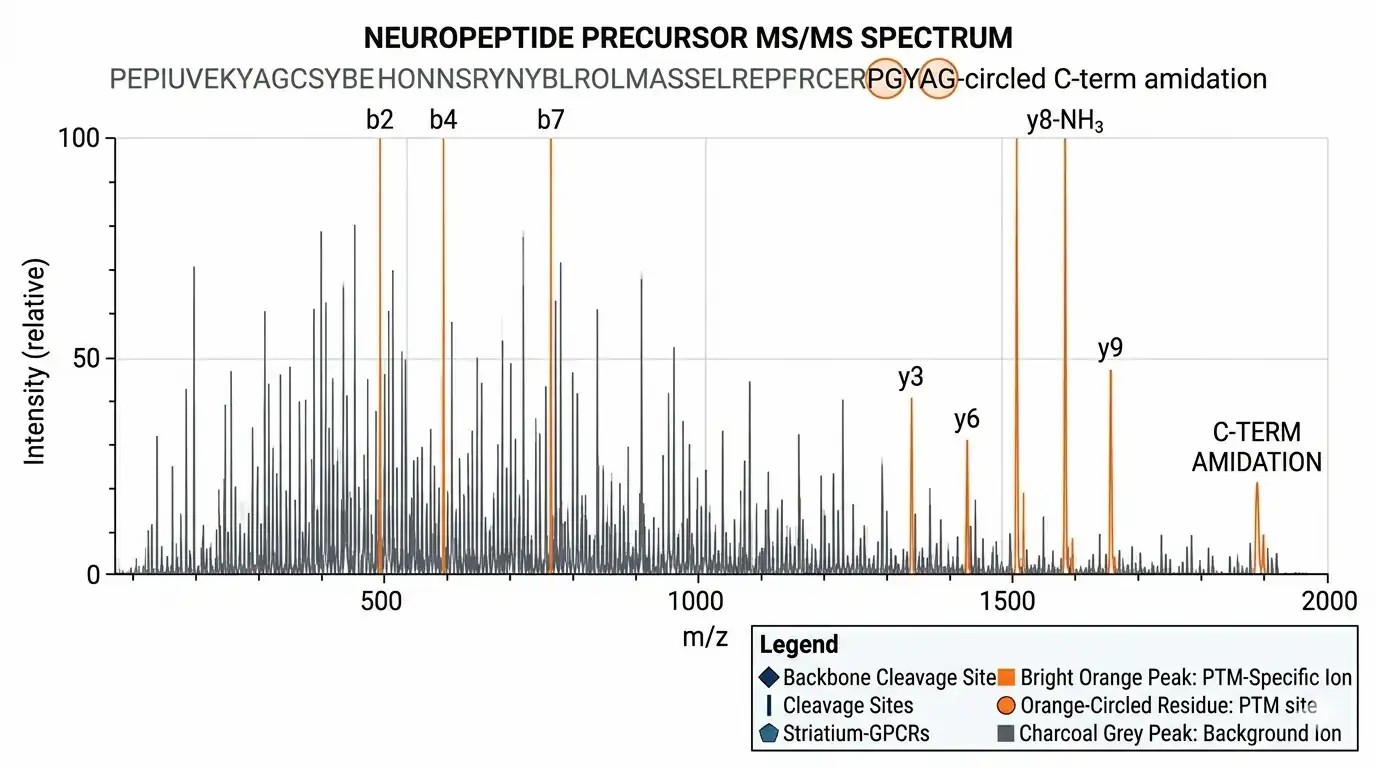
High-resolution fragment ion spectra proving high-confidence PTM localization (e.g., distinguishing specific b and y ion shifts).
PTM Occupancy
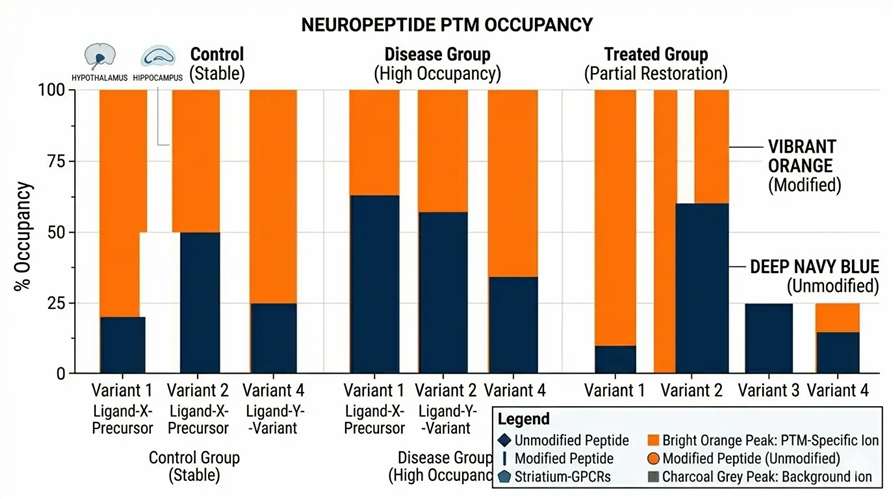
Statistical charts illustrating the dynamic shift in PTM occupancy across your experimental cohorts.
Data Deliverables
Clients receive comprehensive data packages designed to support both regulatory IND filings and top-tier academic publications:
- Candidate Biomarker Panel Shortlist (10–50 high-confidence targets).
- Annotated MS/MS spectra for PTM validation.
- PTM Occupancy Quantification Tables (% Modified vs Unmodified).
- Processing Intermediate Identification Logs.
- Final Peptide Variant List (Annotated with mature boundaries and specific PTMs).
Sample Requirements for Neuropeptide PTM and Processing Analysis
| Sample Type | Best Use Case | Min. Input | Critical Preservation Notes |
|---|---|---|---|
| Brain Tissue / Microdissections | Spatial mapping of processing enzymes | ≥10 mg | Require Rapid Thermal Inactivation or snap-freezing immediately upon collection. |
| Cerebrospinal Fluid (CSF) | Endogenous maturation profiling | 100 – 200 µL | Add specialized protease inhibitors; avoid any blood contamination. |
| Primary Neurons / Cell Culture | Phenotypic enzyme inhibitor screening | 106 cells | Quench processing rapidly; analyze secretome and intracellular fractions. |
Disclaimer: All services and platforms described are for Research Use Only (RUO). Not for use in diagnostic procedures.