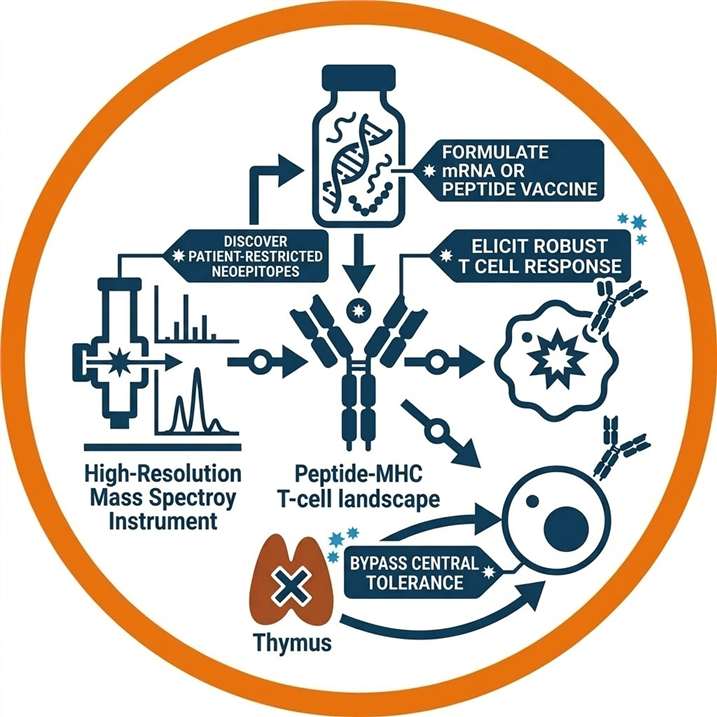
Overcoming In Silico Limitations in Tumor Antigen Discovery
The field of immuno-oncology has rapidly evolved beyond tracking simple tumor mutational burden (TMB). While next-generation sequencing (NGS) allows for the rapid identification of somatic mutations, relying solely on genomic data and in silico binding algorithms presents significant limitations.
Algorithms cannot accurately model the highly complex intracellular antigen processing machinery—including proteasomal cleavage, TAP transport, and ERAP trimming. This biological disconnect frequently leads to high false-positive rates, funneling resources into predicted neoantigens that are never naturally presented on the tumor cell surface.
To overcome this, modern neoantigen discovery demands an evidence-based approach. Whether it involves mapping the wild-type landscape via baseline HLA Peptidomics Analysis or actively hunting for tumor-specific mutations via high-resolution mass spectrometry, identifying true neoepitopes requires integrating physical peptide detection with deep genomic and transcriptomic context.
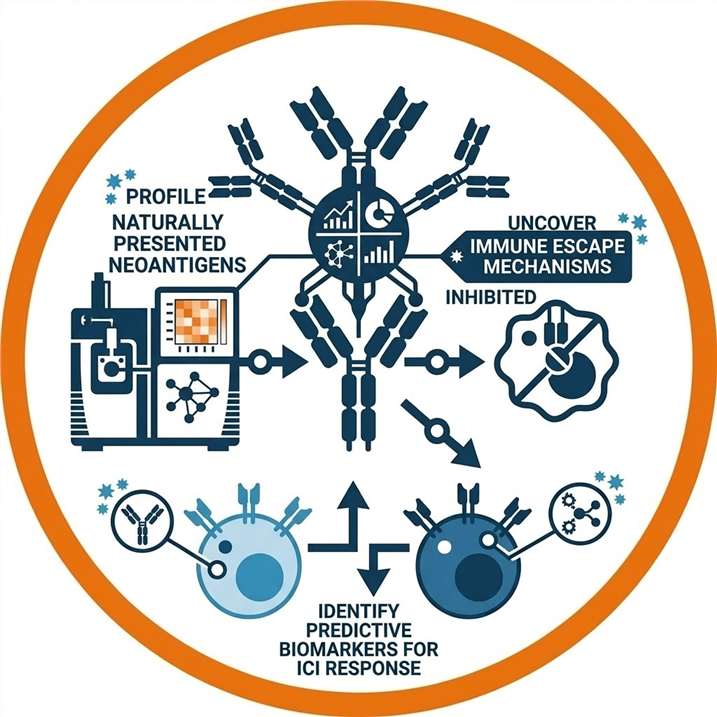
Immuno-Oncology Applications for Neoantigen Identification
Our neoantigen target identification hub is designed to support diverse translational research and therapeutic development pipelines:
Comprehensive Neoantigen Discovery Solutions
As a fully integrated CRO, Creative Proteomics offers a complete suite of neoantigen discovery and prioritization services. We help you select the optimal route based on your project goals, integrating our specialized capabilities into a seamless workflow.
Route Selection: Comparing Neoantigen Discovery Methodologies
Selecting the right discovery route depends heavily on your current research phase and sample availability. Our consultative approach ensures you invest in the appropriate level of evidence.
| Discovery Route | Primary Methodology | Output Evidence Level | Best Suited For |
|---|---|---|---|
| In Silico Prediction | WES + RNA-seq + Algorithms | Low: Theoretical binding scores and expression support. | Early target screening; establishing mutation burden. |
| MS-Based Discovery | HLA Immunoaffinity + LC-MS/MS | Medium-High: Physical detection of tumor ligandomes. | Direct discovery without sequencing; profiling cell lines. |
| Proteogenomics | WES/RNA-seq + LC-MS/MS | High: Custom database matching for unbiased discovery. | Comprehensive novel target discovery from clinical biopsies. |
| Targeted Validation | Pre-existing List + Targeted MS | Highest: Absolute confirmation using synthetic standards. | Late-stage pre-clinical validation; finalizing IND targets. |
Detectable Tumor-Specific Antigen Types
Integrating RNA-seq with high-resolution mass spectrometry unlocks the ability to discover non-canonical targets that traditional DNA-centric predictions overlook.
| Antigen Type | Biological Origin | Discovery Strategy |
|---|---|---|
| SNV-Derived Neoantigens | Point mutations in protein-coding exons. | Direct mapping of MS/MS spectra to variant databases. |
| Indel-Derived Neoantigens | Frameshift mutations altering the reading frame. | Identification of novel sequences via custom FASTA search. |
| Fusion-Derived Neoantigens | Chromosomal translocations producing chimeric transcripts. | Detection of peptides physically spanning the fusion junction. |
| Splice-Derived Neoantigens | Dysregulated splicing or splice site mutations. | RNA-seq guided mapping of intron-retention events. |
| Cryptic / Non-Canonical | Translation of non-coding regions (UTRs, ncRNAs). | Searching spectra against 6-frame translated RNA-seq data. |
Core Platform Advantages
Creative Proteomics bridges the gap between genomics and proteomics, offering unique technical advantages that ensure your neoantigen discovery is accurate, reproducible, and ready for clinical translation.
End-to-End Neoantigen Discovery Workflow
Regardless of the selected route, our operations follow a rigorous, quality-controlled wet-lab and dry-lab workflow to ensure data integrity from sample preparation to final reporting.
High-Resolution Mass Spectrometry Platforms for Immunopeptidomics
Detecting neoantigens is exceptionally challenging due to their incredibly low abundance among thousands of endogenous wild-type peptides. Following the complex enrichment workflow, Creative Proteomics relies on industry-leading mass spectrometry hardware to ensure unmatched sensitivity, resolution, and scanning speed.
MS Platform Comparison
| Platform | Core Technology | Key Advantage | Best Suited For |
|---|---|---|---|
| Orbitrap Astral™ | Asymmetric Track Lossless (Astral) analyzer | Unparalleled scanning speed (>200 Hz) and high dynamic range. | Deep proteogenomic ligandome discovery from limited samples. |
| timsTOF Ultra | Trapped Ion Mobility Spectrometry (TIMS) & PASEF® | 4D-Proteomics with extreme sensitivity and isobaric resolution. | Ultra-low input biopsies and complex HLA ligand mixtures. |
| Orbitrap Exploris™ 480 | High-Field Orbitrap | Robust, reproducible high-resolution mass accuracy. | Standard baseline immunopeptidomics and large cohort profiling. |
| TSQ Altis / SureQuant™ | Triple Quadrupole / Targeted Orbitrap | Absolute heavy-isotope quantification and multiplexing. | Targeted pre-clinical validation of shortlisted neoantigens. |
Comprehensive Sample Requirements for Neoantigen Profiling
Our flexible pipelines accommodate a wide variety of starting materials. Because HLA immunopeptidomics relies on extracting intact, non-crosslinked protein complexes, Formalin-Fixed Paraffin-Embedded (FFPE) tissues are strictly incompatible with MS-based discovery.
| Sample Type | Minimum Input Requirement | Ideal Input for Deep Profiling | Notes & Preservation |
|---|---|---|---|
| Fresh Frozen Tumor Tissue | ≥ 50 mg | 100 - 200 mg | Must be snap-frozen immediately in liquid N2. Do NOT use OCT embedding. |
| Matched Normal Tissue | ≥ 30 mg | 50 mg | Essential for proteogenomics to filter out patient-specific germline variants. |
| Cell Lines / Cultured Cells | ≥ 1×10⁸ cells | 5×10⁸ cells | Wash thoroughly with cold PBS before snap-freezing the cell pellet. |
| Blood / PBMCs | ≥ 1×10⁸ viable cells | ≥ 1×10⁸ viable cells | Suitable for baseline profiling or characterizing antigen-presenting cells (APCs). |
| Nucleic Acids (For In Silico) | 1 µg DNA / 2 µg RNA | 2 µg DNA / 5 µg RNA | Required if starting from physical samples rather than raw sequencing data. |
| Raw Sequencing Data | N/A | N/A | FastQ, BAM, or VCF formats. Acceptable for pure bioinformatic prediction. |
Multi-Omics Bioinformatics & Data Visualization
Our bioinformatics pipelines generate publication-ready, Nature-grade data visualizations, providing transparent insights into the discovered neoantigen landscape.
Differential Neoantigen Presentation (Volcano Plot)
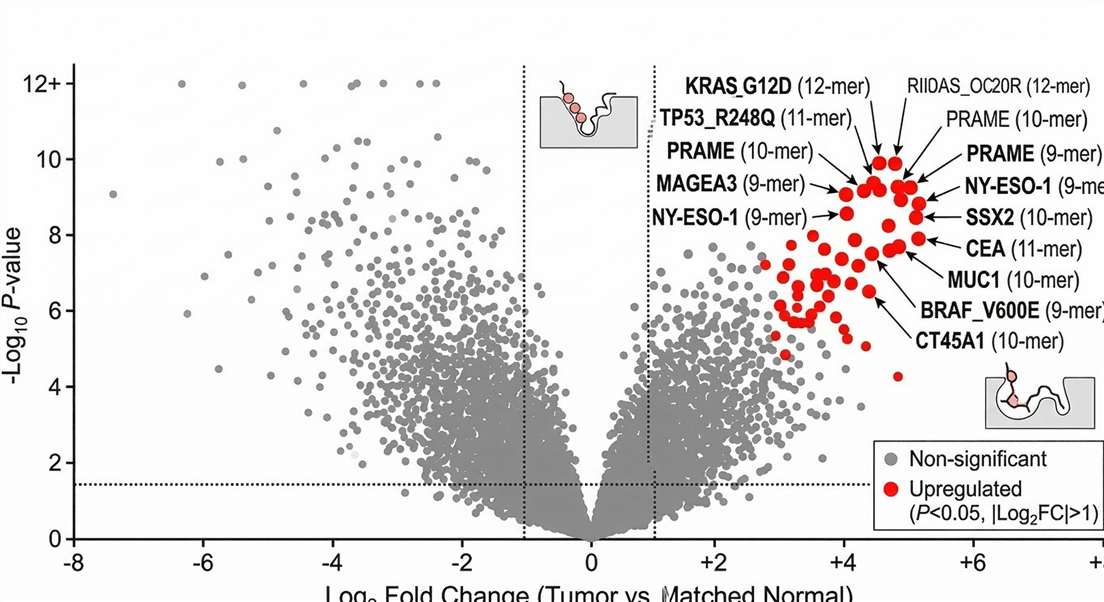
Differential Ligandome Analysis: Volcano plot highlighting significantly over-presented tumor-associated antigens and uniquely detected somatic neoepitopes compared to matched normal tissues.
HLA-Restricted Ligandome Motif Analysis
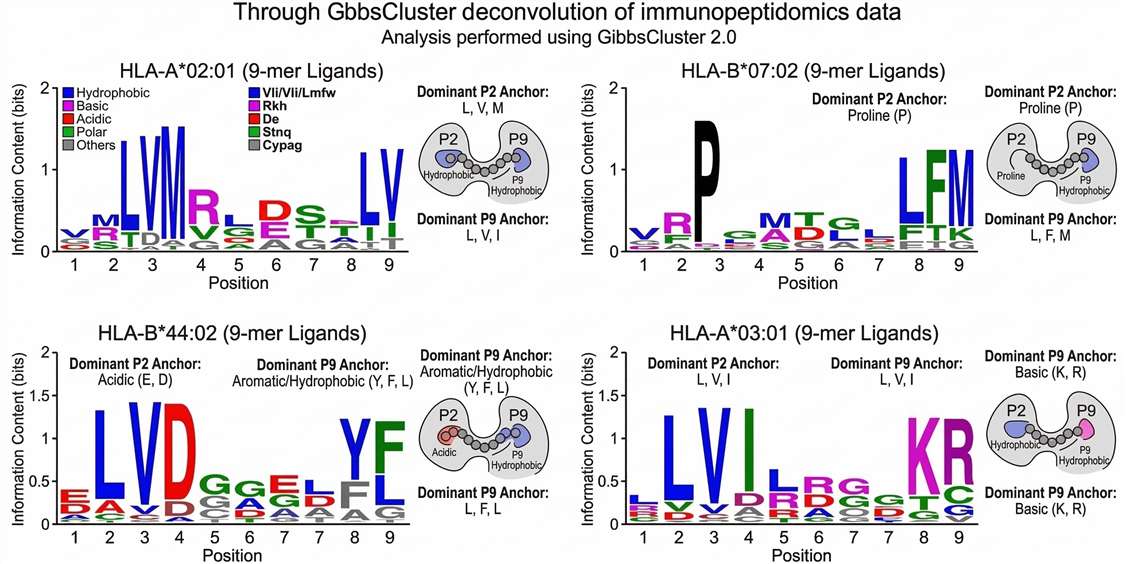
HLA Binding Motif Verification: Sequence logo clustering confirms that physically detected peptides adhere to the anchor residue motifs of the patient's HLA allotypes, validating immunoprecipitation quality.
Extracted Ion Chromatogram (XIC) Quantification
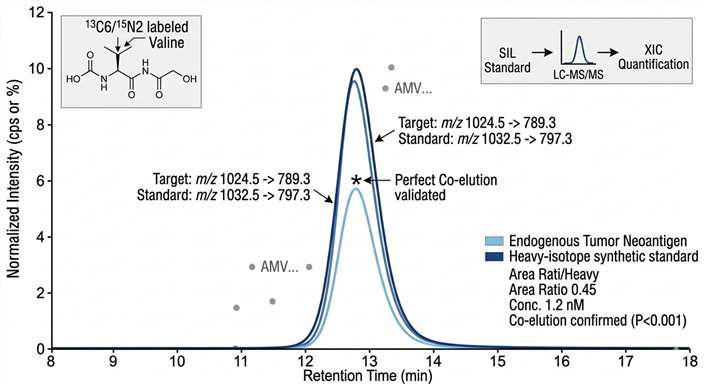
Quantitative Physical Validation: Extracted Ion Chromatograms (XIC) demonstrating perfect retention time alignment between the endogenous neoantigen and a synthetic standard, ensuring high-confidence identification.
High-Resolution MS/MS Sequence Tagging
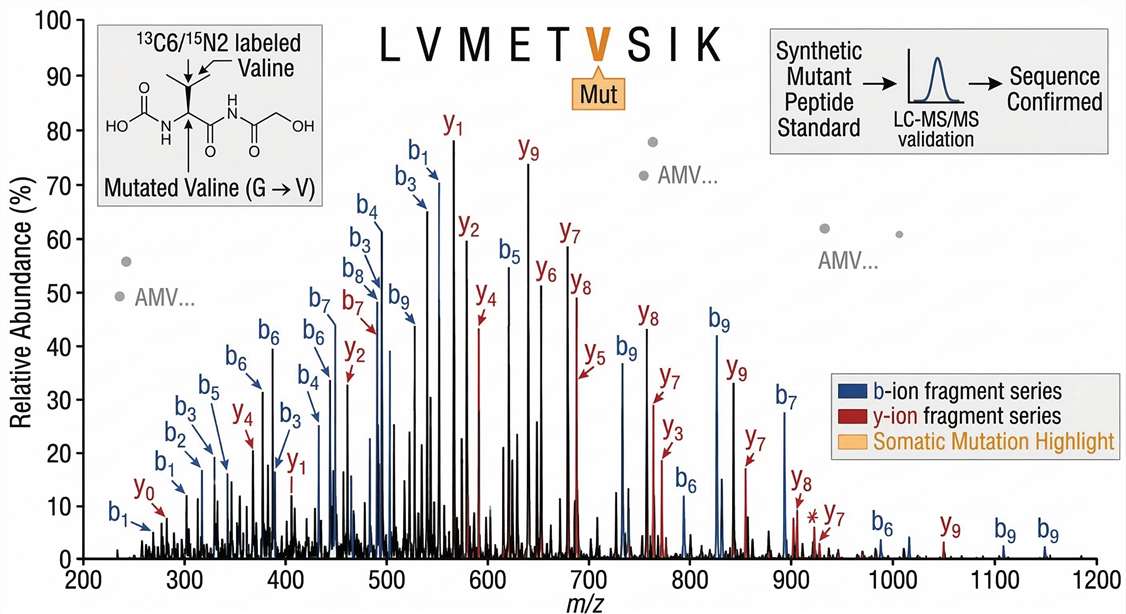
Definitive Sequence Confirmation: Annotated tandem mass spectrum (MS2) providing incontrovertible physical proof of the neoantigen's exact sequence, pinpointing the somatic mutation site.
Neoantigen Data Deliverables & Downstream Validation
Creative Proteomics delivers highly structured, actionable data packages designed to empower your downstream clinical and research decision-making.
- Target Prioritization Matrix: An integrated dataset ranking discovered neoantigens based on empirical MS abundance, transcriptomic expression (TPM), and HLA binding affinities (%Rank).
- Comprehensive Variant Annotation: Detailed genomic contextualization of every validated peptide, identifying source genes and variant types.
- Raw Multi-Omics Data: Full access to generated FastQ files, BAM alignments, and high-resolution mass spectrometry RAW files for internal archiving.
- Executive Summary Reports: Methodological details, quality control metrics (FDR control, HLA yield), and advanced graphical visualizations.
Frequently Asked Questions

Case Study: Proteogenomic Identification of Melanoma Neoepitopes
Journal: Nature Communications
Published: Volume 7, 2016
Summary
Using a multi-omics proteogenomic approach, researchers integrated WES, transcriptomics, and high-resolution mass spectrometry to identify naturally presented neoantigens directly from melanoma tissues. The study proved that mass spectrometry significantly reduces the false-positive rates of in silico predictions, leading to the discovery of physically validated neoepitopes capable of eliciting strong T cell responses.
Methods & CP Support
This study highlights a rigorous proteogenomic pipeline to physically validate algorithmic predictions. To support research following these principles, Creative Proteomics offers specialized services including:
- Custom variant database construction from patient-specific WES/RNA-seq.
- High-resolution HLA immunopeptidomics profiling on Orbitrap/timsTOF platforms.
- Multi-layer evidence integration to filter false positives for ranked candidate reporting.
Results
- MS Validation: Only a highly specific subset of computationally predicted mutations was physically detected by MS on the tumor cell surface.
- Novel Targets: The approach identified mutant neoepitopes (e.g., from NCAPG2 and SYTL4 genes) that were completely absent in matched normal tissues.
- Functional Activation: T-cell assays confirmed that MS-validated neoepitopes triggered robust IFN-γ secretion from the patient's own CD8+ T cells.
Reference
- Bassani-Sternberg, M., et al. "Direct identification of clinically relevant neoepitopes presented on native human melanoma tissue by mass spectrometry." Nature Communications 7, 13404 (2016). DOI: 10.1038/ncomms13404
For Research Use Only. Not for use in diagnostic procedures.