Immunopeptide Affinity Screening in Epitope Discovery
Epitope discovery pipelines consistently generate vast libraries of candidate sequences, whether derived from computational predictions or upstream Immunopeptidomics Service workflows. However, identifying a peptide sequence is only the first step. To trigger a robust immune response, an antigenic peptide must physically bind to an HLA (Human Leukocyte Antigen) molecule with sufficient stability to be transported to the cell surface and recognized by a T cell.
Immunopeptide affinity screening bridges the critical gap between sequence identification and functional validation. By experimentally quantifying the binding strength between candidate peptides and specific MHC complexes, researchers can eliminate false positives predicted by in silico models. This targeted validation drastically reduces the number of candidates requiring expensive, labor-intensive downstream cellular assays, ensuring that research efforts focus exclusively on high-viability epitopes.
Peptide–HLA Binding and Immune Recognition
Antigen presentation is a highly selective mechanical process. Intracellularly processed peptides compete for limited binding grooves within MHC Class I and Class II molecules. The affinity of this interaction—defined by the equilibrium dissociation constant (KD) and the off-rate kinetics (koff)—directly dictates the half-life of the peptide-MHC complex on the cell surface.
High-affinity binding is a mandatory prerequisite for immunogenicity. Without a stable presentation platform, corresponding T cell receptors (TCRs) cannot securely engage the antigen, leading to failed CD8+ cytotoxic killing or CD4+ helper T cell activation. Consequently, quantifying peptide-HLA binding kinetics is the most reliable biochemical predictor of a peptide's true immunological potential.
Applications in Vaccine and Neoantigen Research
Our screening service is perfectly balanced to support both neoantigen validation for immuno-oncology applications and vaccine epitope discovery.
What We Offer in Immunopeptide Affinity Screening
We deliver specialized, modular solutions to validate your epitope candidates:
Workflow for Immunopeptide Affinity Screening
Our service is highly scalable, supporting everything from individual high-precision validations to library-scale screening. Clients may provide pre-synthesized peptides, or Creative Proteomics can handle end-to-end peptide synthesis and screening to ensure project flexibility.
Technology Platform for Peptide Interaction Analysis
Our screening platform integrates multiple state-of-the-art analytical methods, including Bio-Layer Interferometry (BLI), Surface Plasmon Resonance (SPR), and targeted biochemical assays, to ensure high-precision peptide-MHC binding analysis. These technologies offer real-time, label-free measurements for both kinetic constants and relative binding affinities. To support discovery-phase projects, our platform seamlessly integrates with Immune Peptide Library Screening workflows.
Our service provides broad coverage of both HLA Class I and Class II, ensuring comprehensive analysis for both CD8+ and CD4+ T cell epitopes. This pan-allelic capability supports the development of vaccines and immunotherapies targeting diverse patient populations. To guarantee analytical reliability, positive and negative controls, including high-affinity reference peptides and non-binding mutant controls, are rigorously used to establish assay baselines.
Affinity Screening Technology Comparison
| Feature | Surface Plasmon Resonance (SPR) | Bio-Layer Interferometry (BLI) | Biochemical / Competition Assays |
|---|---|---|---|
| Readout Type | Real-time kinetics (KD, kon, koff) | Real-time kinetics (KD, kon, koff) | End-point relative affinity (IC50) |
| Sensitivity | Ultra-high (ideal for small peptides) | High | Moderate to High |
| Throughput | Medium to High | Medium to High | Ultra-High (Library scale) |
| Sample Tolerance | Requires highly purified samples | Fluidics-free; tolerant to crude matrices | Requires standard buffer conditions |
| Primary Advantage | Industry gold standard for precise binding kinetics | Flexible, rapid setup with strong kinetic accuracy | Highly cost-effective for large-scale primary screening |
Affinity Screening vs Epitope Prediction
| Dimension | Epitope Prediction (In Silico) | Affinity Screening (Experimental) | T Cell Activation Assay |
|---|---|---|---|
| Method | Computational modeling | Biochemical physical interaction | Cellular co-culture |
| Evidence Type | Theoretical binding probability | Direct experimental binding strength | Functional immunological response |
| Throughput | Ultra-high (Whole genomes) | Scalable (Individual to Library) | Low to Medium |
| Validation Strength | Low (Prone to false positives) | High (Confirms physical viability) | Definitive (Confirms immune action) |
| Typical Use Stage | Wide-net candidate discovery | Candidate validation & ranking | Final preclinical confirmation |
Sample Requirements for Peptide Affinity Screening
To ensure kinetic accuracy, proper sample preparation is critical.
| Sample Type | Typical Input | Required or Optional | Purpose | Notes |
|---|---|---|---|---|
| Synthetic Peptides | Candidate epitopes | Required (if client provided) | Direct binding screening | >95% purity required for accurate kinetics |
| Peptide Libraries | Overlapping epitope pools | Optional | High-throughput screening | Library synthesis design varies |
| Neoantigen Peptides | Predicted tumor epitopes | Optional | Patient-specific validation | Sequence confirmation required |
Note: Exact sample quantities depend heavily on the desired analytical platform and number of target alleles. Please consult our team for verified project specifics.
Example Results from Immunopeptide Affinity Screening
Our deliverables translate raw biophysical interaction data into highly visual, actionable insights for epitope selection, ready for inclusion in research publications or regulatory filings.
Peptide Affinity Ranking Chart
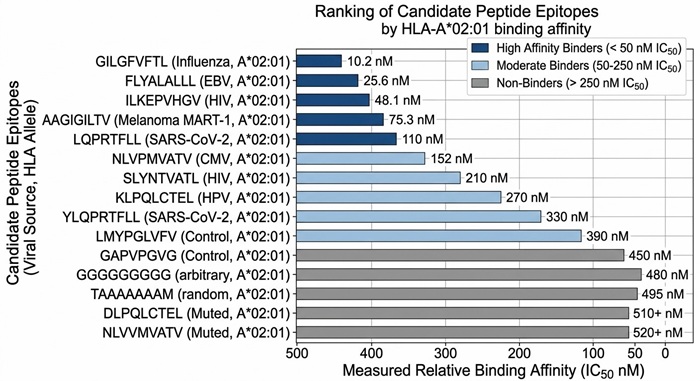
Quantitative ranking of candidate peptides by their relative binding affinity, effectively highlighting the strongest binders for downstream functional validation.
Peptide-HLA Interaction Heatmap
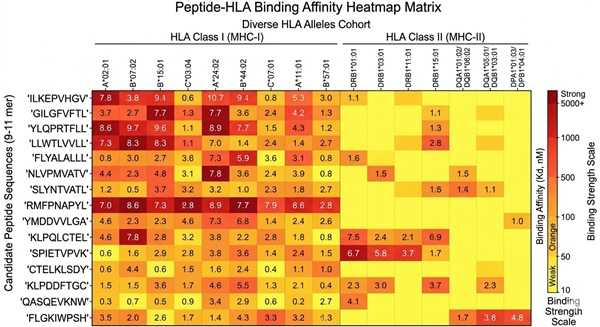
Comprehensive interaction heatmap illustrating binding strengths across candidate peptides and a panel of target HLA alleles, ideal for identifying broadly promiscuous vaccine epitopes.
Real-Time Kinetics Sensorgram
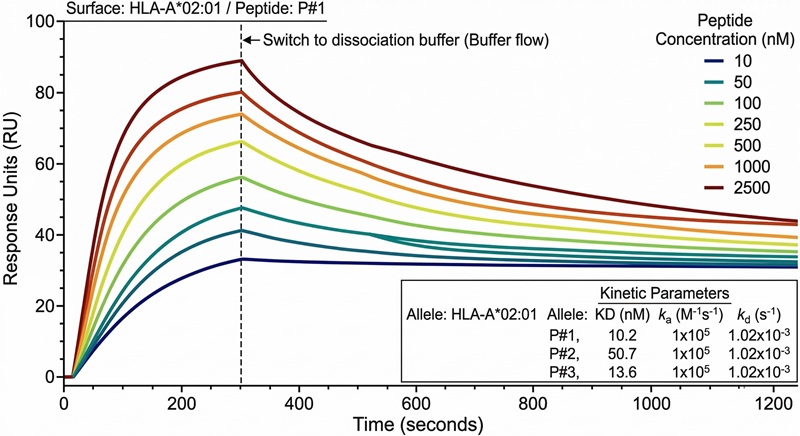
Real-time BLI/SPR binding curves capturing highly accurate kinetic association (kon) and dissociation (koff) rates for lead immunopeptide epitopes.
Epitope Prioritization Matrix
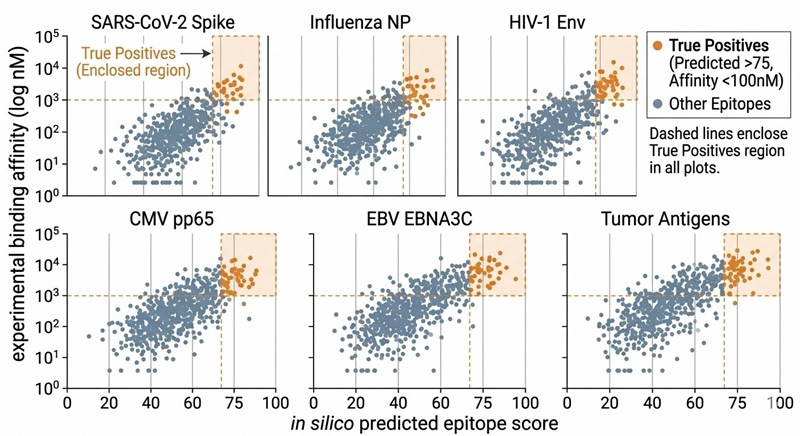
Scatter plot mapping theoretical prediction scores against actual experimental affinity, definitively separating biologically viable binders from computational false positives.
Deliverables
Our immunopeptide screening service provides comprehensive data packages designed to drive your immunological research forward:
- Kinetic Constants Report: Precise equilibrium dissociation constants (KD), association rates (kon), and dissociation rates (koff).
- Relative Affinity Rankings: Standardized IC50 values comparing your candidates against validated baseline reference peptides.
- Raw Interaction Data: Exported SPR or BLI sensorgrams for your internal records and publication needs.
- Candidate Shortlist: An expert-curated epitope prioritization matrix identifying the most promising peptides for downstream assays.
Frequently Asked Questions
HLA Class II Binding Landscape Mapping by Deep Mutational Scanning
Journal: Frontiers in Immunology
Published: 2023
Summary
To reduce unwanted immunogenicity in therapeutic proteins, researchers combined deep mutational scanning with HLA class II binding analysis to systematically evaluate how amino acid substitutions influence antigen presentation. By constructing mutation libraries spanning immunogenic sequence regions, the study generated a residue-level heatmap describing how substitutions altered predicted HLA class II binding across multiple alleles. This approach enabled the identification of sequence variants that reduce potential CD4⁺ T cell epitope presentation while preserving functional binding properties, providing a rational strategy for biologics de-immunization and sequence optimization.
Methods
This study developed an integrated experimental–computational workflow to characterize the HLA class II binding landscape of immunogenic peptide regions.
Mutation libraries containing single amino acid substitutions were generated for selected peptide sequences associated with potential immunogenicity. These variants were evaluated using HLA class II binding prediction models to estimate their relative binding affinities across relevant alleles. The resulting data were organized into a binding heatmap, allowing researchers to visualize how each residue contributes to antigen presentation.
In parallel, deep mutational scanning and yeast surface display assays were used to measure how substitutions affected protein functionality, including target binding capability. By combining functional screening with HLA binding analysis, the workflow enabled identification of substitutions that simultaneously reduce predicted immune recognition while maintaining biological activity.
Key Technical Features
Sample types: Mutational peptide libraries derived from immunogenic sequence regions
Mutation strategy: Systematic single-residue substitution scanning
Binding analysis: HLA class II binding prediction integrated with residue-level heatmap visualization
Functional screening: Deep mutational scanning combined with yeast surface display
Variant prioritization: Integrated analysis of immunogenicity risk and functional activity
Library design: Combinatorial variants constructed from beneficial substitutions identified during scanning
Creative Proteomics can offer advanced immunopeptidomics and peptide–HLA affinity screening services that support similar research pipelines, including:
- HLA class I and class II peptide binding affinity screening
- Epitope mapping and immunogenicity risk assessment for biologics
- Peptide library screening for epitope hotspot identification
- Integrated analysis combining experimental binding data with epitope prediction
- Custom peptide synthesis and kinetic binding validation using SPR or BLI
These services support vaccine development, neoantigen validation, and therapeutic protein de-immunization studies.
Results
Residue-Level Mapping of HLA Binding Sensitivity
The mutational scanning analysis produced a detailed map of how individual amino acid substitutions influence predicted HLA class II binding. The resulting heatmap revealed that certain positions strongly contribute to HLA binding, while other residues tolerate substitution without substantially affecting antigen presentation.
Identification of Substitutions That Reduce Predicted Immunogenicity
Several substitutions were identified that significantly lowered predicted HLA class II binding affinity. These variants represent potential strategies for reducing helper T cell epitope presentation in therapeutic protein sequences.
Balancing Immunogenicity and Functional Activity
By integrating HLA binding predictions with deep mutational scanning data, researchers were able to prioritize substitutions that reduced predicted immune recognition while preserving target binding functionality. This dual-filter strategy provides a practical framework for rational de-immunization in biologics engineering.
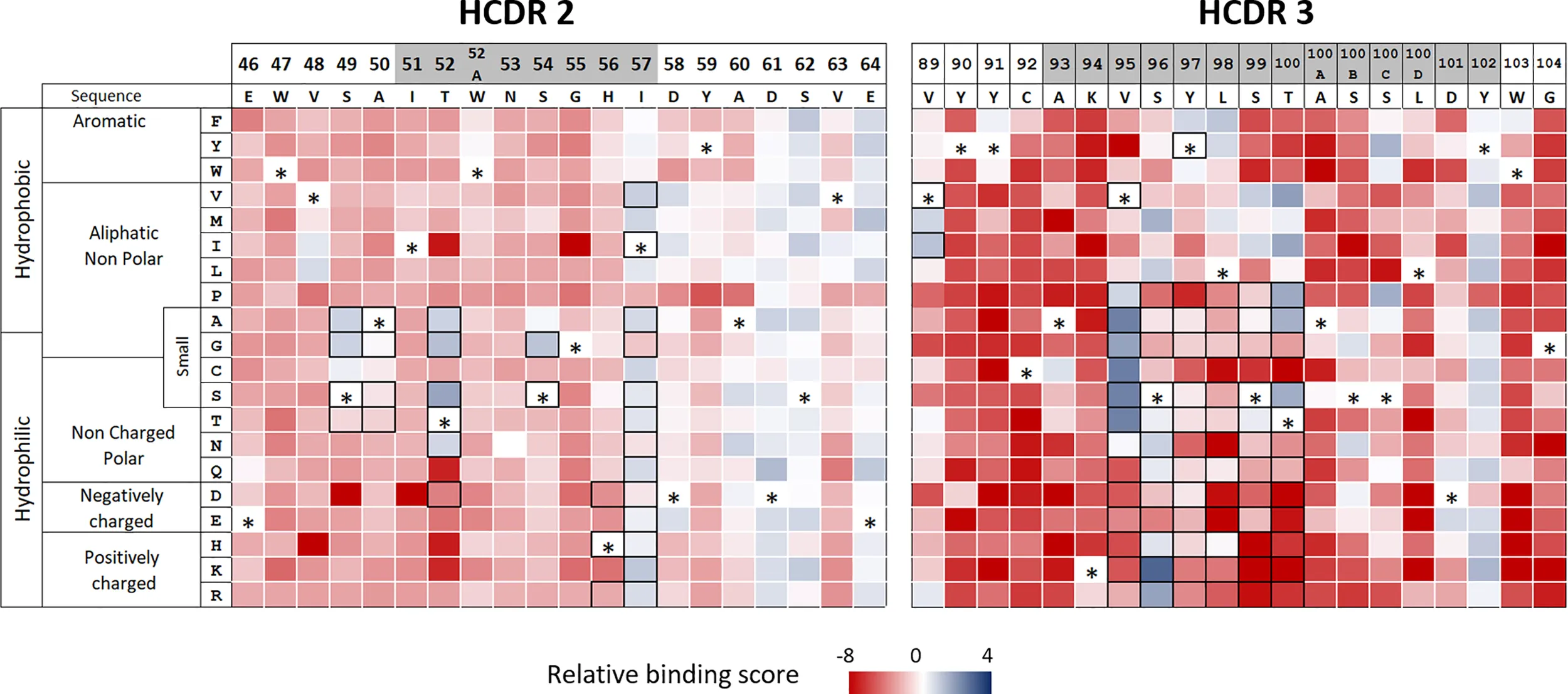
Figure 2 Heatmap visualization of mutation-dependent changes in predicted HLA class II binding, highlighting substitutions that may reduce antigen presentation while maintaining functional sequence integrity.
Reference
- Sivelle, Coline, et al. "Combining deep mutational scanning to heatmap of HLA class II binding of immunogenic sequences to preserve functionality and mitigate predicted immunogenicity." Frontiers in Immunology (2023). https://doi.org/10.3389/fimmu.2023.1197919