Resolving Brain Region Heterogeneity in Neuropeptide Research
Standard "bulk" tissue homogenization masks the subtle yet critical molecular changes occurring within specific brain structures. Neuropeptides function as potent neuromodulators within localized synapses and distinct neurocircuits; their expression, release, and degradation are highly spatially regulated. For instance, the expression of neuropeptide Y (NPY) or substance P can vary dramatically between the striatum, the hippocampus, and the prefrontal cortex. Grinding these distinct macro-structures together into a single lysate dilutes regional signals and entirely obliterates the anatomical context necessary for understanding neurobiological mechanisms.
When investigating complex phenomena—such as focal neuroinflammation, localized amyloid plaque accumulation, or distinct neurochemical alterations in specific brain nuclei—maintaining spatial context is essential. Spatial neuropeptidomics overcomes the limitations of bulk analysis by isolating region-specific peptide signatures. This targeted approach enables researchers to correlate molecular profiles with precise anatomical architecture. By doing so, it reveals the highly localized mechanisms driving neurodegenerative conditions, psychiatric disorders, and traumatic brain injuries, allowing for a much deeper understanding of how neuropeptidergic networks adapt or fail under pathological stress.
Our Service Offerings and Detection Types
We provide tailored analytical workflows to meet the diverse and exacting needs of translational CNS research. Our comprehensive spatial profiling solutions encompass the following core offerings and capabilities:
Our Technical Advantages
Integrated Platform: MALDI Imaging & LC-MS/MS Deep Discovery
To fully map the neuropeptidome, researchers must answer two fundamental questions: Where is the peptide located? and What is its precise molecular identity? Our integrated dual-platform approach answers both with uncompromising accuracy.
| Platform Component | Technical Function | Key Output |
|---|---|---|
| MALDI-TOF/TOF Imaging | Rapid visualization of peptide ion gradients across intact tissue sections via specialized matrix application and laser scanning. | 2D mass maps revealing anatomical clustering, spatial compartmentalization, and relative abundance of neuropeptides. |
| LCM + LC-MS/MS | Laser Capture Microdissection (LCM) of distinct regions of interest (ROIs) followed by DDA/DIA LC-MS/MS using high-resolution Orbitrap systems. | Definitive amino acid sequence identities and PTM mapping for spatially distributed masses, confirming biological identity. |
Strategic Choice: Selecting the Right Spatial Approach for Your Study
Selecting the appropriate analytical platform depends heavily on your specific research question, desired spatial resolution, and required identification depth. To assist in your project design, we have outlined the strategic differences below.
| Dimension | MALDI Mass Spectrometry Imaging | LCM coupled with LC-MS/MS |
|---|---|---|
| Primary Goal | Visualizing spatial distribution mapping across intact tissues. | Deep sequence identification and quantitation of discrete sub-regions. |
| Spatial Resolution | High (Sub-cellular to 10-50 micrometer resolution). | Dependent on LCM precision (typically defined by micro-dissected area). |
| Identification Depth | Lower (Focuses on highly abundant, highly ionizable peptides). | High (1,000 to thousands of peptides, dependent on sample input). |
| Best Suited For | Mapping known therapeutic peptides, abundant biomarkers, or broad screens. | Discovering novel neurocircuit markers and deep pathway analysis. |
Solution Selection Strategy:
- Choose MALDI Imaging if your objective is to visually trace the gradient of a known neuropeptide across different brain nuclei, or to assess the penetration and distribution of a peptide drug within a specific brain lesion.
- Choose LCM + LC-MS/MS if your goal is comprehensive discovery—for instance, identifying all neuropeptide alterations occurring exclusively within the substantia nigra of a Parkinson's disease model compared to healthy controls.
- Consider a Hybrid Approach if you need to visually map novel therapeutic targets and simultaneously validate their precise amino acid sequences to confirm biological activity.
Workflow: From Precision Sampling to Data Acquisition
Our laboratory follows a highly optimized, stringent standard operating procedure to ensure batch-to-batch reproducibility, minimal background noise, and maximal peptide recovery.
Sample Requirements and Preparation Guidelines
Proper sample preparation is the most critical variable in spatial neuropeptidomics. We provide detailed, step-by-step protocols to our clients upon project initiation.
| Sample Type | Recommended Input | Preparation Notes | Shipping Condition |
|---|---|---|---|
| Fresh Frozen Brain Tissue | Intact whole brain or specific macroscopic regions (>50 mg). | Snap-freeze immediately in liquid nitrogen. Avoid embedding media like OCT if conducting LC-MS. | Dry ice |
| Microdissected Tissue (LCM) | Approx. 1-2 cubic millimeters or >10,000 cells per specific region. | Collect directly into specialized lysis buffer. Ensure rapid processing to halt enzymatic activity. | Dry ice |
| Tissue Sections (for Imaging) | 10-12 micrometer thickness, mounted on ITO-coated glass slides. | Do not wash or fix prior to consultation. Follow specific mounting protocols. | Dry ice (slide mailers) |
Data Deliverables: Spatial Distribution and Quantitative Insights
We translate complex multidimensional mass spectrometry data into publication-ready visual insights, empowering researchers to directly address their biological hypotheses.
Regional Abundance Profiling
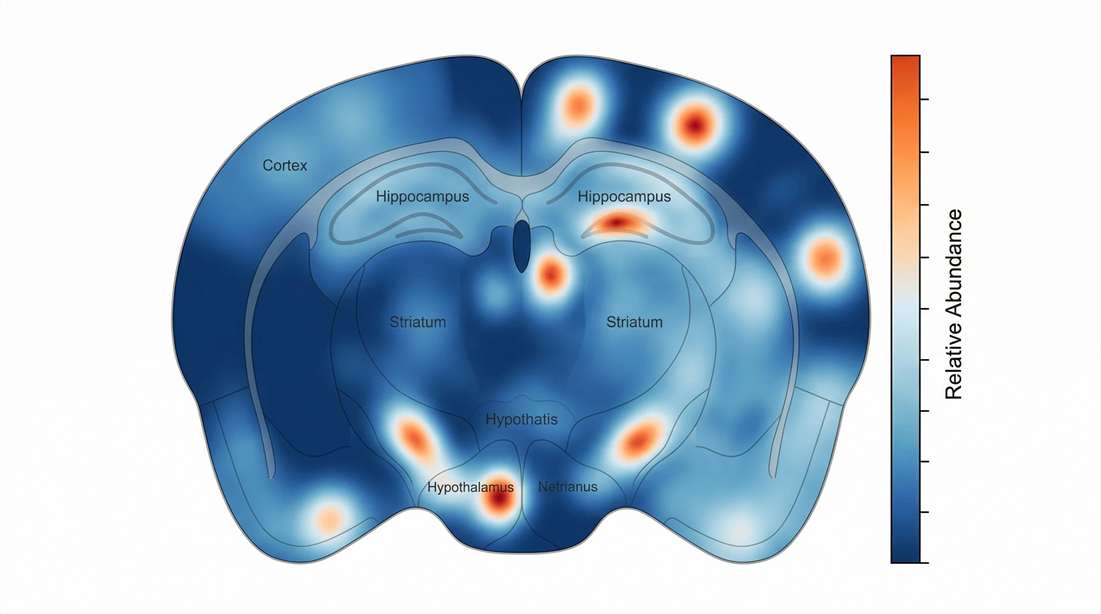
Visualizing spatial gradients of bioactive peptides directly on the tissue architecture.
Differential Expression by Location
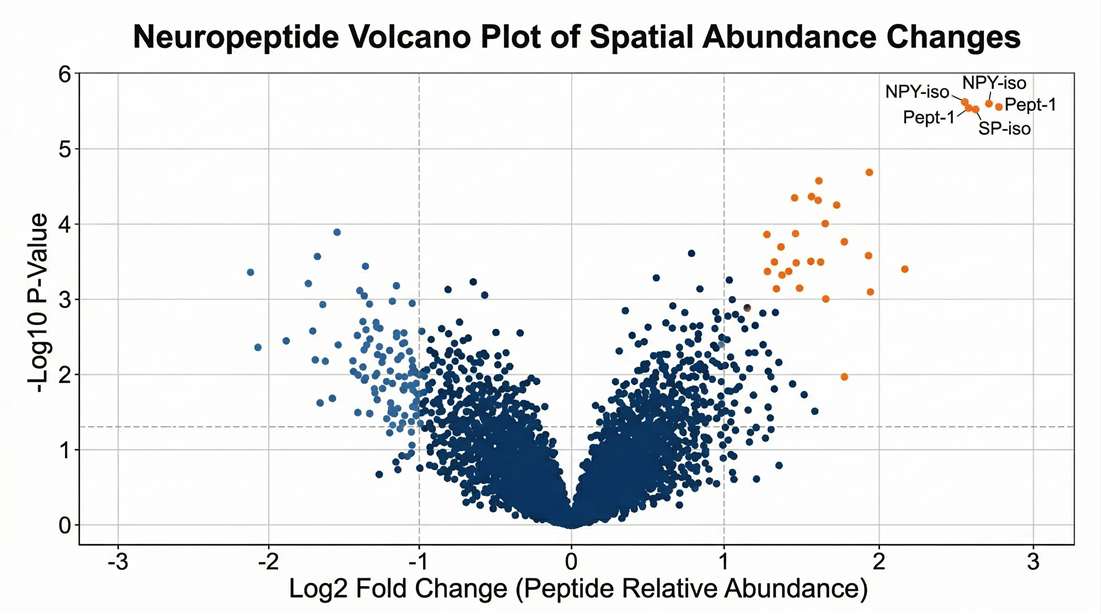
Statistically identifying significantly altered neuropeptides between distinct neurocircuits.
Multivariate Sample Segregation
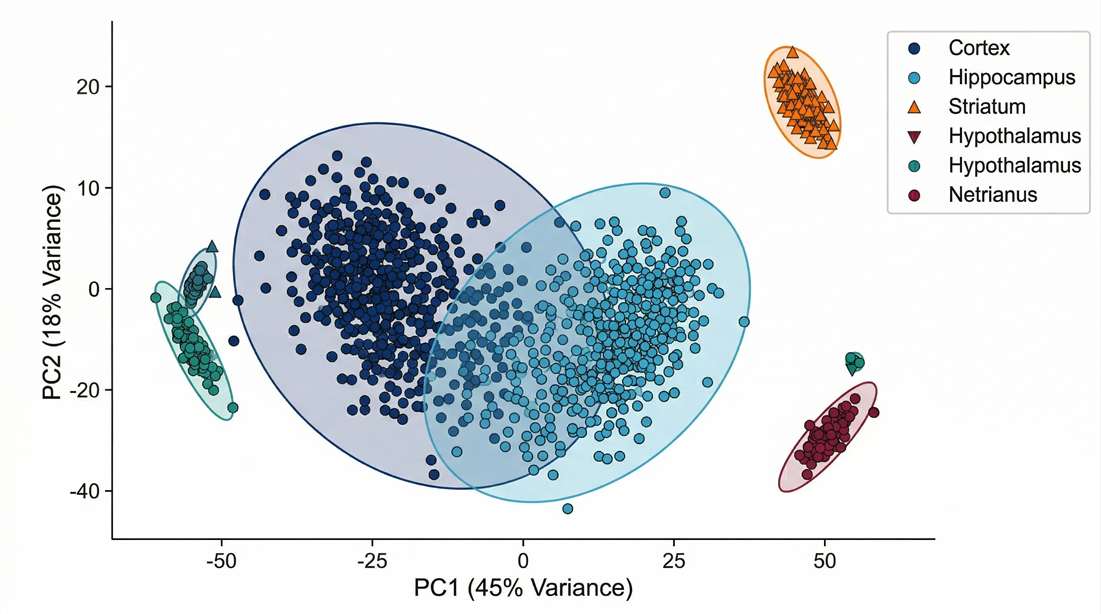
Validating molecular distinctness of targeted neurocircuits through dimensionality reduction.
Biological Mechanism Mapping
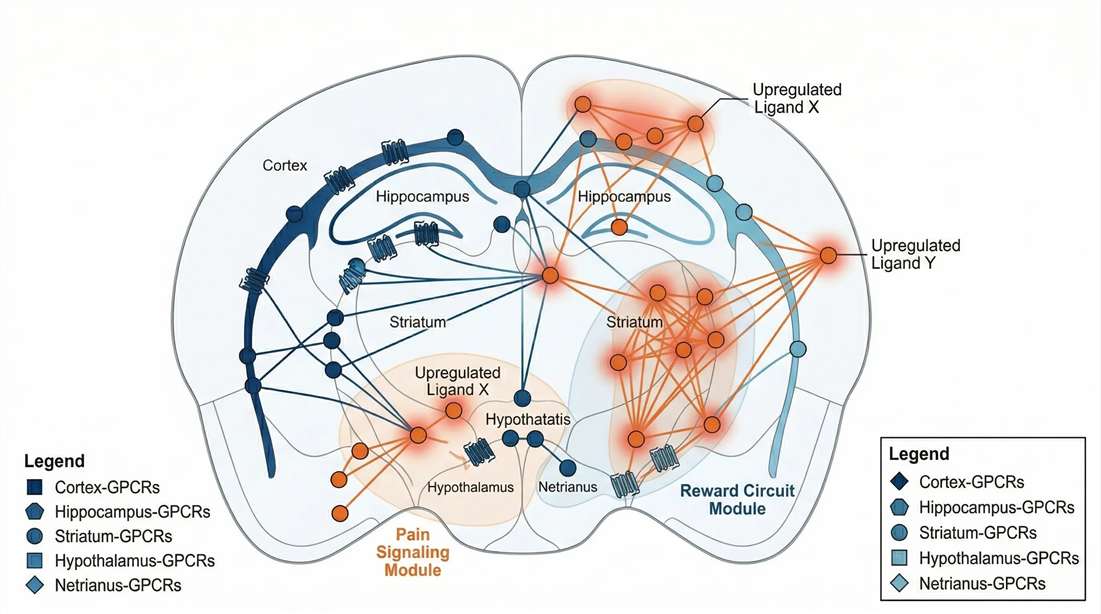
Decoding neuropeptide signaling mechanisms via receptor-ligand networks.
Comprehensive Bioinformatics Analysis
Transforming raw mass spectra into actionable biological insights requires robust, specialized bioinformatics pipelines. Our data analysis services are structured to provide both foundational peptidome profiling and advanced functional interpretation without requiring in-house computational expertise.
Minimum Deliverables (Included in standard projects)
- High-confidence peptide identification lists (strictly filtered by False Discovery Rate below 1 percent).
- Regional differential abundance matrices complete with rigorous normalization metrics.
- Basic statistical visualizations, including Principal Component Analysis (PCA), Volcano plots, and Hierarchical Clustering Heatmaps.
- Quality control (QC) summary reports outlining mass accuracy, retention time stability, and technical reproducibility.
Optional Add-ons (For advanced biological interpretation)
- Receptor-Ligand Interaction Mapping: Bioinformatics alignment of identified endogenous peptides with their known G-protein coupled receptors (GPCRs) to infer localized signaling networks and neuro-immune crosstalk.
- Pathway Enrichment for Spatial Domains: Gene ontology (GO) and biological pathway mapping specific to the neuropeptides upregulated or downregulated in defined brain regions.
- Precursor Protein Cleavage Analysis: Mapping detected peptides back to their pro-hormone precursors to identify potential alterations in localized processing enzyme activity.
Disclaimer: The services and products described herein are for Research Use Only (RUO). Not for use in diagnostic procedures.
Multiplexed Neuropeptide Mapping Integrating Three-Dimensional Mass Spectrometry Imaging
Focus Area: Spatial Neuroscience & Brain Microanatomy
Context: Based on methodologies described in advanced spatial neuropeptidomics literature.
Summary
Traditional immunohistochemical methods for localizing neuropeptides are limited by the need for specific antibodies and the brain's opacity. In this comprehensive methodological approach, researchers integrated Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry Imaging (MALDI-MSI) with microcomputed tomography (micro-CT) to achieve 3D spatial mapping of chemically diverse neuropeptides. By acquiring serial mass spectrometry images and co-registering them with high-resolution anatomical scans, the study successfully linked complex neuropeptide chemistry directly to distinct brain microanatomy, providing unprecedented multiplexed maps of intact neuroendocrine circuits without the need for labeling.
Methods
The study established a powerful multi-modal pipeline to resolve the spatial distribution of endogenous peptides:
Key Technical Workflows:
- Sample Preparation: Serial sectioning of intact brain tissues with immediate matrix application to preserve endogenous peptide stability and prevent rapid post-mortem degradation.
- MALDI-MSI Acquisition: High-resolution spatial scanning across multiple tissue slices to detect a wide array of neuropeptide masses simultaneously in a single measurement.
- Micro-CT Integration: Utilization of microcomputed tomography to create a structural 3D scaffold of the brain architecture.
- 3D Co-Registration: Advanced computational alignment of the 2D MALDI-MSI chemical maps onto the 3D anatomical micro-CT framework.
Creative Proteomics offers parallel spatial peptidomics solutions designed to support highly multiplexed brain mapping, including:
- High-resolution MALDI mass spectrometry imaging for intact tissue sections.
- Laser Capture Microdissection (LCM) coupled with LC-MS/MS for deep peptide sequencing of defined regions.
- Bioinformatics mapping of peptide distributions onto established anatomical brain atlases.
Results
Unprecedented Multiplexing Capability
- Demonstrated the ability to simultaneously detect and map dozens of distinct molecular species (including neuropeptides and related neurotransmitters) within a single analytical run, overcoming the multiplexing limitations of traditional antibody-based staining.
High-Fidelity 3D Anatomical Mapping
- Successfully reconstructed the 3D distribution of specific neuropeptidergic networks by aligning serial 2D MSI slices into a unified volumetric model.
- Correlated specific peptide mass signatures with precise anatomical substructures, revealing intricate region-specific expression gradients.
Methodological Robustness
- The integration of MS imaging with micro-CT provided a highly robust workflow for validating the precise spatial origin of neuroactive peptides, establishing a new benchmark for studying localized signaling at the molecular level.
Reference
- Dueck B, et al. "Multiplexed neuropeptide mapping in ant brains integrating microtomography and three-dimensional mass spectrometry imaging." PNAS Nexus (2023). https://pmc.ncbi.nlm.nih.gov/articles/PMC10194420/