Neurodegenerative Disease Biomarker Discovery Strategy
CNS biomarker discovery in neurodegenerative diseases presents unique challenges for therapeutic development. In disorders such as Alzheimer's disease (AD), Parkinson's disease (PD), and Huntington's disease (HD), pathological changes often precede clinical symptoms by years. While static structural proteins have historically guided diagnosis, accelerating modern drug development requires dynamic, high-resolution molecular signatures for mechanistic biomarker discovery. These markers must accurately reflect disease staging, neuroinflammation, and intervention responses.
Finding these elusive signatures requires navigating the blood-brain barrier (BBB) and bridging the gap between localized brain pathology and accessible peripheral signals. As a dedicated partner in translational research, our CNS biomarker discovery service delivers integrated multi-omics biomarker discovery solutions (proteomics, peptidomics, and metabolomics), including cutting-edge neuropeptidomics for neurodegeneration. We empower pharmaceutical pipelines to transition from multi-matrix discovery to targeted validation, identifying actionable molecular signatures for biomarker development and clinical validation.
- Alzheimer's Disease (AD): Moving beyond traditional amyloid and tau measurements to profile synaptic dysfunction biomarkers and neuroinflammatory cascades that correlate with early cognitive decline.
- Parkinson's Disease (PD): Identifying dynamic multi-omics signatures that reflect dopaminergic neuron degeneration and mitochondrial dysfunction before motor symptoms appear.
- ALS & Huntington's Disease: Capturing stage-specific molecular changes and tracking disease progression through longitudinal biofluid sampling.
Multi-Matrix Profiling: Connecting Brain Pathology to Peripheral Signals
The greatest hurdle in translational CNS biomarker research is the "Brain-to-Blood" translation chasm. Pathological mechanisms occur within discrete brain regions, but clinically viable biomarkers must be detectable in accessible biofluids.
We overcome this barrier through a strategic **Multi-Matrix Profiling** approach tailored for neurodegenerative disease peptidomics and proteomics:
When to Use This CNS Biomarker Discovery Service
This service is strategically designed for researchers and clinical directors when:
- You need to discover novel biomarkers beyond amyloid/tau to gain a competitive edge in neurodegenerative research.
- You aim to translate brain tissue findings from animal models or post-mortem studies into accessible peripheral (CSF/plasma) biomarkers.
- You are initiating large-scale clinical cohorts that require a robust, reproducible pipeline with minimal batch effects.
- You require a seamless transition from discovery to targeted validation (PRM/MRM) without switching platforms or vendors.
Comprehensive Discovery-to-Validation Biomarker Workflow
Our integrated discovery-to-validation biomarker workflow ensures actionable results for large clinical cohorts with strict batch-effect control.
Assessment of brain tissues, CSF, and plasma with proprietary high-abundance protein depletion and stabilization QC.
Utilizing Orbitrap Astral or timsTOF Pro for deep proteomic and peptidomic profiling across biological replicates.
Advanced bioinformatics integration to identify overlapping signatures between brain pathology and peripheral biofluids.
Applying machine learning algorithms to define the most robust peptide biomarker discovery signature for patient stratification.
Rapid transition to absolute quantification (PRM/MRM) for shortlisted candidates across expanded clinical cohorts.
Platform Performance Benchmarks
- Deep Resolution: Identification of 1,000–3,000+ molecular features per sample depending on the matrix.
- Rigorous Precision: Intra-batch CV typically <15%, inter-batch CV <20% for large-scale cohort studies.
- Low-Input Compatibility: Optimized for precious samples (≥10 mg tissue / ≥100 µL CSF / ≥200 µL plasma).
- High Reproducibility: Proven consistency across multi-center cohort designs through automated sample preparation.
MS Peptidomics vs. Antibody-Based Panels for CNS Biomarker Discovery
To support robust CNS drug development biomarkers, multi-omics provides unparalleled depth compared to traditional isolated protein assays.
| Feature | Integrated Multi-Omics Profiling | Single-Target Assays (ELISA) |
|---|---|---|
| Disease Staging Capability | High. Identifies dynamic panels that shift across disease stages. | Limited. Often relies on late-stage structural markers. |
| Mechanistic Insight | Deep. Reveals neuroinflammation and autophagy pathways simultaneously. | Limited. Only measures a single predefined target. |
| Brain-to-Blood Translation | Seamless. Connects brain discovery directly to plasma validation. | Disjointed. Dependent on antibody availability. |
| Intervention Response | High. Captures off-target effects and systemic responses. | Limited. Blind to pathways outside the kit. |
Biomarker-Guided CNS Drug Development and Intervention Monitoring
Moving beyond discovery, our platform is a critical engine for CNS drug development biomarkers, supporting pharma R&D in evaluating therapeutic efficacy:
- Pathway Reversal Analysis: Quantify whether a candidate drug successfully reverses disease-associated pathways (e.g., restoring synaptic peptide levels).
- Target Engagement Evidence: Identify downstream molecular shifts proving the therapeutic agent has reached its intended CNS target.
- Longitudinal Monitoring: Track biomarker dynamics over time in clinical trial participants to correlate molecular changes with clinical outcomes.
Translational Visual Insights (Demo Results)
We transform high-dimensional data into clear narratives to support mechanistic understanding and clinical translation.
Pathway Enrichment
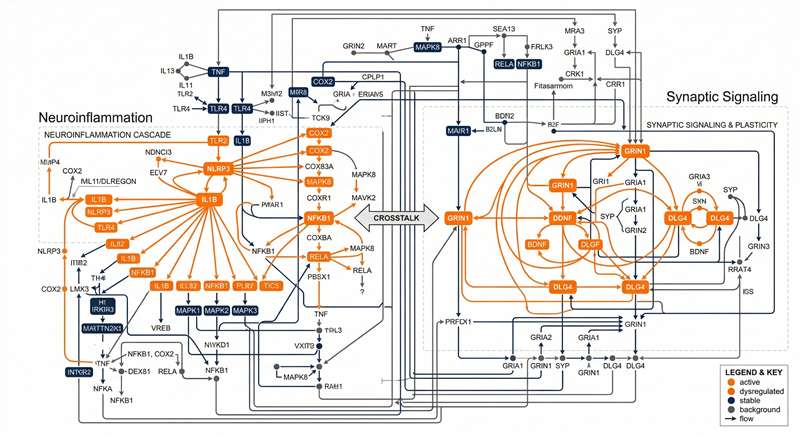
Mapping dysregulated targets into specific CNS pathways like neuroinflammation or synaptic signaling.
Candidate Shortlist
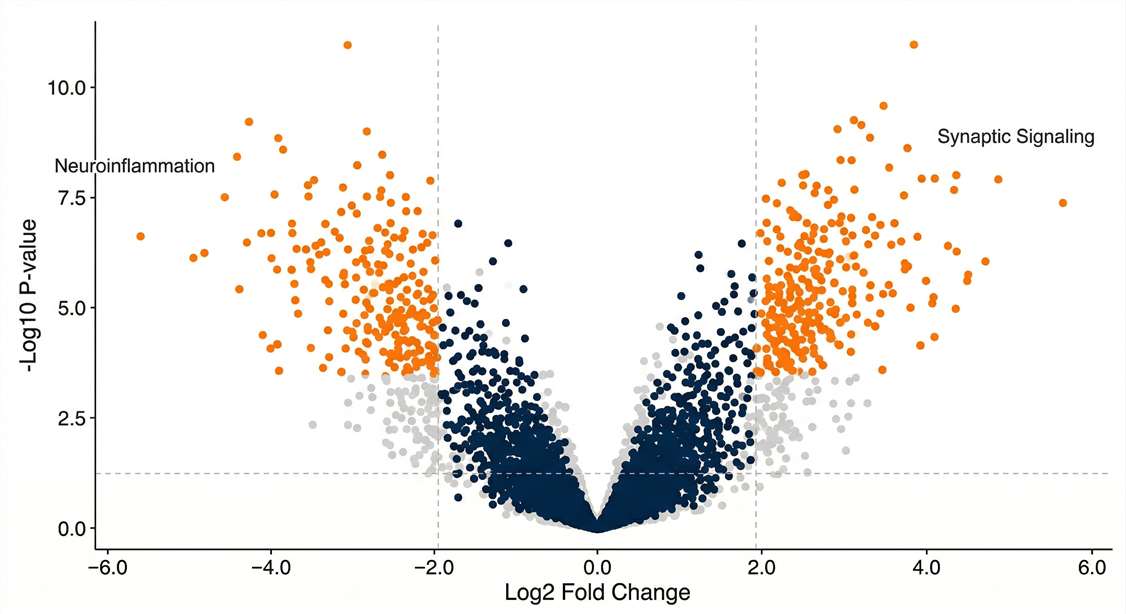
Visualization of significant upregulated and downregulated targets applying strict FDR thresholds.
Cleavage Mapping
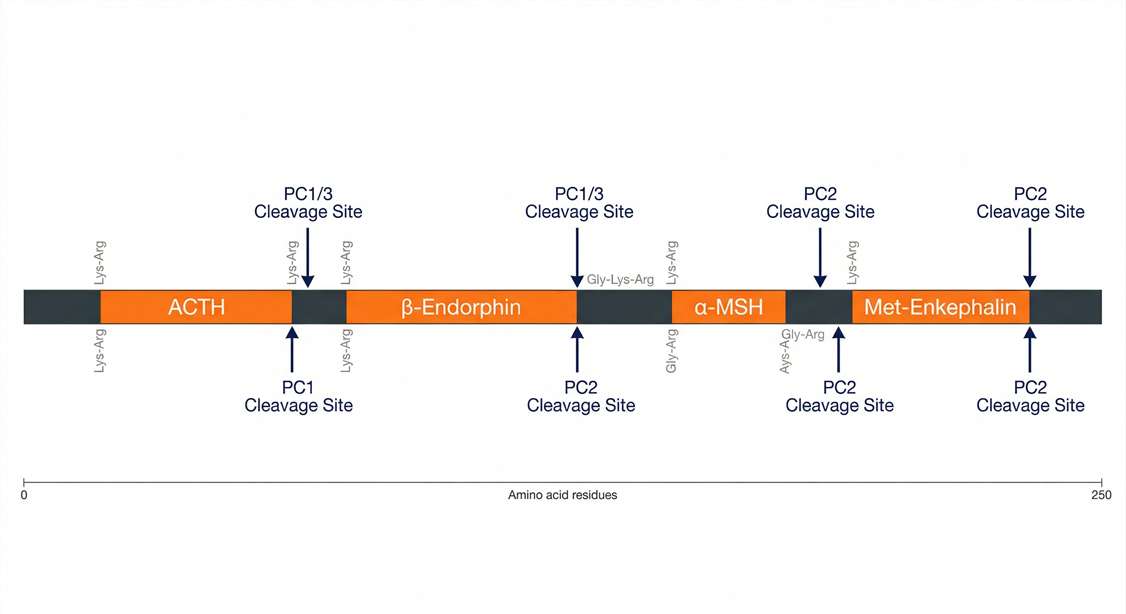
Advanced precursor cleavage mapping revealing altered proteolytic activity linked to disease progression.
Cohort Stratification
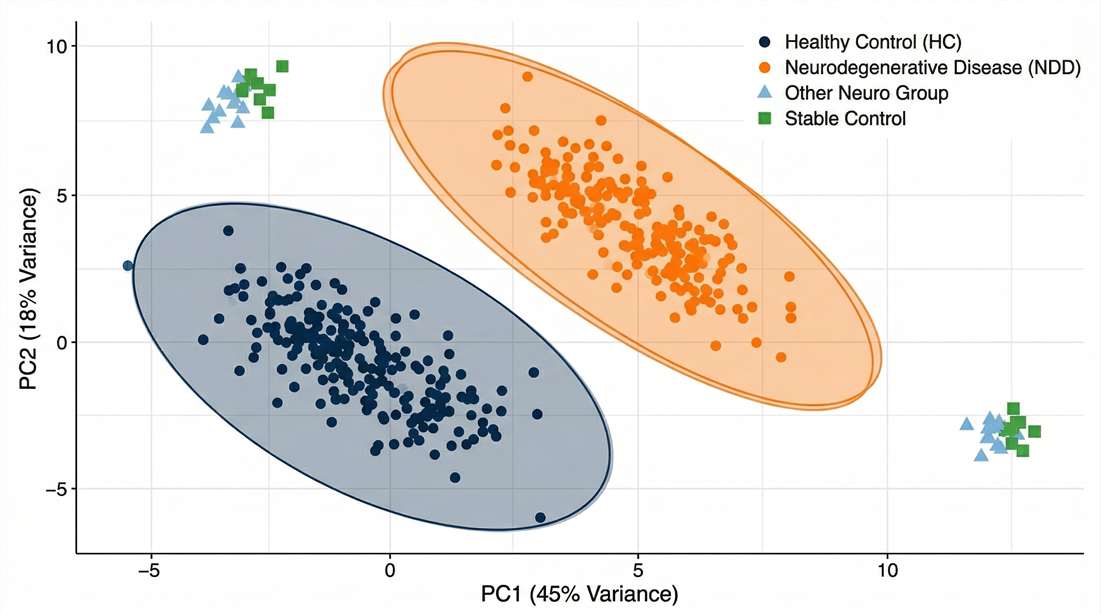
Multivariate analysis validating molecular distinctness, serving as robust patient stratification biomarkers.
Deliverables | What You Will Receive
- Candidate Biomarker Panel Shortlist: A refined list of 10–50 high-confidence targets ready for validation and clinical translation.
- Comprehensive Identification Matrix: Fully annotated lists of proteins/peptides with confidence scores and normalized intensities.
- Bioinformatics & Pathway Report: In-depth analysis of biological networks, protein-protein interactions, and functional clustering.
- Executive Statistical Summary (PDF): Visual highlights including PCA, Volcano plots, and machine learning models.
- Targeted Validation Data: Detailed absolute quantification tables and assay performance metrics for validation phases.
Sample Requirements for CNS Biomarker Discovery
| Sample Type | Best Use Case | Min. Input | Key Considerations |
|---|---|---|---|
| Brain Tissue | Mechanism & Spatial Discovery | ≥10 mg | Snap-freeze in LN2; store at -80°C. |
| CSF | Closest Proxy to CNS State | 100 – 200 µL | Use low-protein binding tubes. |
| Plasma / Serum | Clinical Validation | 200 – 500 µL | EDTA-plasma preferred. |
| Primary Cells | In Vitro Secretome | 10^6 cells | Wash thoroughly with PBS. |
Disclaimer: All services and platforms described are for Research Use Only (RUO). Not for use in diagnostic procedures.
Case Study: Stage-Specific Serum Biomarkers in Huntington's Disease
Journal: Cells
Published: 2025
Background
Huntington's Disease (HD) has a prolonged pre-manifest phase. Identifying accessible peripheral biomarkers that reflect continuous disease progression is critical for early intervention and monitoring clinical trial efficacy in CNS research.
Methods
The research utilized a multi-cohort serum discovery study. The workflow involved:
- Processing: High-Abundance Protein Depletion to uncover trace signaling molecules in patient serum.
- Discovery: Nano LC-MS/MS profiling across manifest HD, pre-manifest, and healthy controls.
- Bioinformatics: Identification of stage-specific serum proteomic signatures using advanced multivariate statistics.
Results
- The MS profiling successfully distinguished pre-manifest HD from active manifest stages with high accuracy.
- Early disease stages correlated with dysregulated immune response and complement cascades; later stages showed cytoskeletal shifts.
- Several peripheral biomarker candidates (e.g., CFH, CAPZB) were validated as non-invasive tools for tracking disease progression.
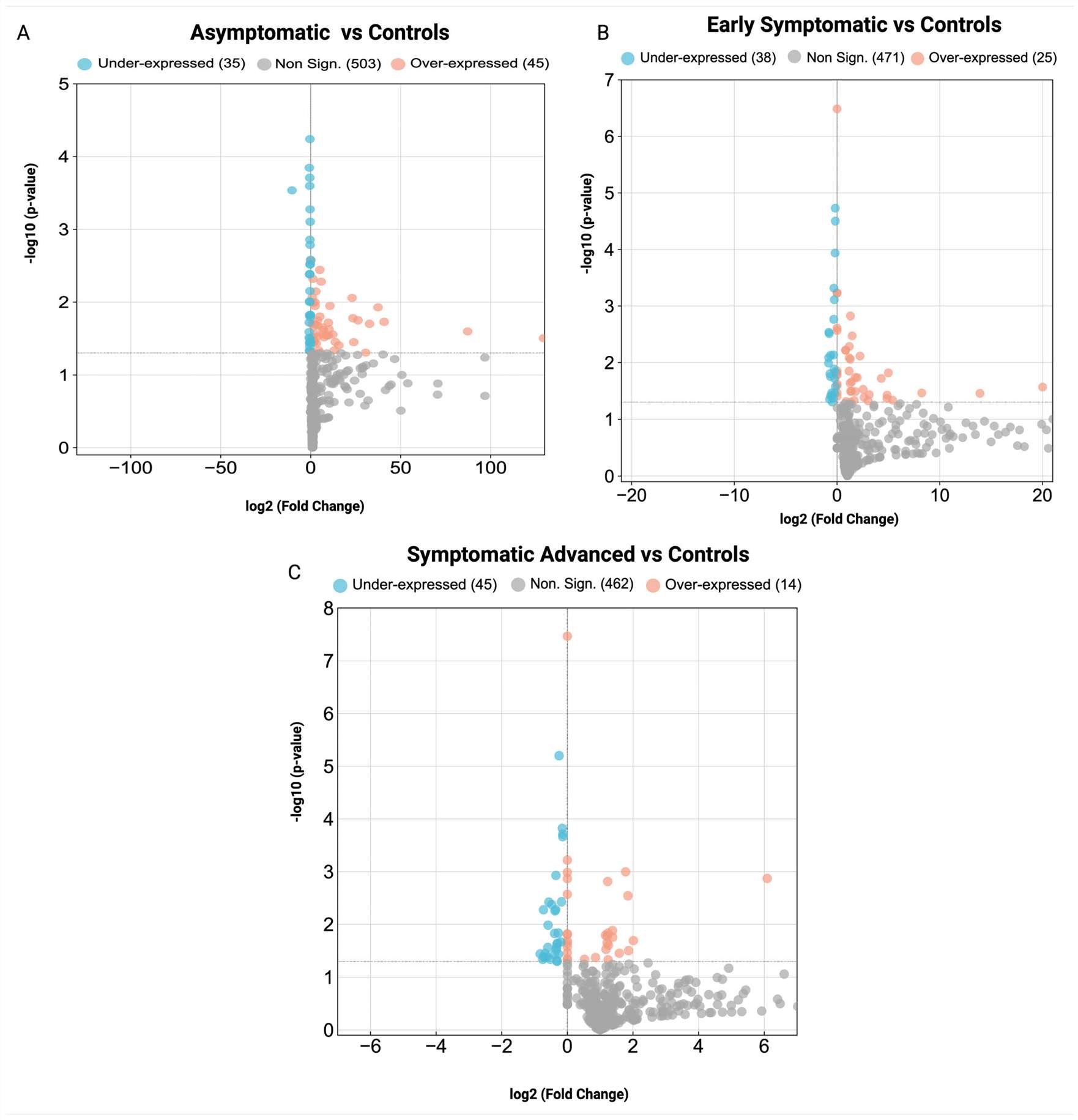
Conclusion & Relevance
This approach enables the robust identification of translatable biomarkers across neurodegenerative disease cohorts, bridging the brain-to-blood gap. By successfully identifying dynamically changing peripheral biomarkers, the platform provides a robust framework for developing non-invasive diagnostic panels and therapeutic monitoring.