Why HLA Peptidomics Analysis?
In immuno-oncology, infectious disease research, and vaccine development, knowing which genomic mutations exist is only the first step. While in silico prediction algorithms can guess which peptides might bind to Major Histocompatibility Complex (MHC) molecules, these tools inherently suffer from high false-positive rates because they cannot perfectly model the complex intracellular machinery of proteasomal cleavage and peptide transport.
To prove that an antigen is genuinely actionable, it must be physically detected on the cell surface. As a cornerstone of our comprehensive immunopeptidomics service, direct HLA Peptidomics Analysis bypasses prediction limits. By physically enriching MHC-peptide complexes and sequencing the eluted ligands via high-resolution mass spectrometry, we provide absolute, ground-truth evidence of naturally presented epitopes, ensuring your downstream functional assays are driven by biological reality rather than theoretical probability.
Key Application Areas
Our HLA peptidome profiling services provide foundational evidence for diverse translational and basic immunological research pipelines:
What We Offer in HLA Peptidomics Analysis
Creative Proteomics delivers a highly specialized, end-to-end analytical framework designed to comprehensively capture both CD8+ and CD4+ T-cell targets. We employ optimized immunoprecipitation (IP) workflows using pan- or allele-specific antibodies to suit your exact research design.
Types of HLA-Bound and MHC-Associated Peptides We Identify
Our high-sensitivity mass spectrometry platforms are capable of identifying thousands to tens of thousands of unique ligands from a single biological sample, covering diverse genomic origins.
| Peptide Category | Biological Origin | Relevance to Immunology |
|---|---|---|
| Tumor-Specific Antigens (TSAs) | Somatic SNVs, indels, and fusions. | Highly specific "neoantigens"; elite targets for personalized cancer vaccines and TCR-T therapies. |
| Tumor-Associated Antigens (TAAs) | Overexpressed endogenous wild-type proteins. | Shared targets across patient cohorts (e.g., NY-ESO-1, MAGE) for broad-spectrum immunotherapies. |
| Minor Histocompatibility Antigens (MiHAs) | Germline SNPs differing between donor and recipient. | Critical for understanding Graft-versus-Host Disease (GvHD) and Graft-versus-Leukemia (GvL) effects. |
| Pathogen-Derived Peptides | Viral, bacterial, or fungal proteins. | Mapping true targets for infectious disease vaccines and tracking memory T-cell responses. |
| Cryptic / Non-Canonical Peptides | Translated non-coding RNAs or alternative splices. | Uncovering novel, highly immunogenic tumor targets completely missed by DNA-centric approaches. |
| PTM-Modified HLA Ligands | Phosphorylated or citrullinated host proteins. | Discovering inflammation-specific neoantigens driven by aberrant intracellular signaling pathways. |
Methodological Advantages in HLA Peptidomics
Beyond state-of-the-art hardware, our refined immunoaffinity and bioinformatics workflows ensure that we capture the true ligandome while minimizing false discoveries.
Step-by-Step HLA Peptidomics Workflow
Our standardized, quality-controlled wet-lab and dry-lab workflow ensures maximum peptide yield and data integrity from sample receipt to final reporting.
High-Resolution LC-MS/MS Platforms
Achieving deep coverage of the immunopeptidome requires pushing the boundaries of sensitivity and scan speed. We continuously invest in the highest-resolution mass spectrometry platforms to accurately sequence complex endogenous peptide mixtures present at exceptionally low abundances.
- Thermo Fisher Orbitrap™ Series: Utilizing the Orbitrap Eclipse™, Exploris™ 480, and the ultra-high-throughput Astral™ mass spectrometers, providing unparalleled mass accuracy and dynamic range for deep peptidome mining.
- Bruker timsTOF Pro 2 & Ultra: Leveraging 4D-Proteomics™ and PASEF® (Parallel Accumulation-Serial Fragmentation) technology to achieve extreme sensitivity, allowing us to identify low-abundance neoantigens from minimal biopsy inputs.
- Advanced Nano-LC Systems: Coupled with ultimate-performance liquid chromatography (e.g., Thermo Vanquish™ Neo, nanoElute® 2) to ensure flawless peptide separation and reproducible retention times.
Bioinformatics Annotation & Prioritization
While the core of HLA peptidomics is the physical identification of sequences, the resulting data is only actionable when properly contextualized. Bioinformatics is a critical, integrated component of our service.
Once high-confidence peptides are identified, our bioinformatics team performs deep annotation. We analyze peptide length distributions to verify HLA restriction (e.g., sharply peaking at 9-mers for Class I, broadly distributing around 15-mers for Class II). We utilize Gibbs clustering algorithms to extract consensus binding motifs, confirming that the physical peptides match the anchor residue preferences of the sample's specific HLA allotypes.
For complex projects, we offer advanced integration options. We can map identified peptides back to their source proteins for pathway enrichment analysis, or cross-reference the physical MS data with in silico neoantigen prediction and prioritization services to highlight the high-value overlap between predicted immunogenicity and guaranteed natural presentation.
Prediction vs Direct HLA Peptidomics: A Comparison
Understanding when to rely on algorithms versus when to invest in direct mass spectrometry is crucial for efficient pipeline progression.
| Dimension | In Silico HLA Prediction | Direct HLA Peptidomics (LC-MS/MS) |
|---|---|---|
| Core Methodology | Algorithms predicting theoretical binding affinity. | Physical extraction and mass spectrometry sequencing. |
| Evidence Depth | Hypothesis generation; shows what might bind. | Ground-truth evidence; shows what is actually presented. |
| False-Positive Risk | Very High (often >90% false positives). | Extremely Low (Strict <1% FDR control). |
| Complex Peptides (PTMs) | Generally unable to predict PTM presentation. | Directly identifies phosphorylated or modified ligands. |
| Best-Fit Use Case | Broad screening of thousands of mutations early on. | Final validation of targets before vaccine or TCR-T formulation. |
Sample Requirements for HLA Peptidomics
Due to the reliance on extracting intact protein complexes, sample preservation is critical. Formalin-Fixed Paraffin-Embedded (FFPE) tissues are strictly incompatible with this workflow.
| Sample Type | Typical Input Requirement | Preservation & Shipping | Notes |
|---|---|---|---|
| Fresh Frozen Tissue | 100 - 300 mg (≥200 mg for dual class) | Snap-freeze in liquid N2 immediately. Ship on dry ice. | Ideal for tumor biopsy profiling. Do NOT use OCT embedding as polymers interfere with MS. |
| Cultured Cell Lines | 1×10⁸ - 5×10⁸ cells (Up to 10⁹ for Class II) | Wash thoroughly in cold PBS, pellet, and snap-freeze. Ship on dry ice. | Input requirements vary greatly based on baseline HLA expression. IFN-γ pretreatment is often recommended. |
| PBMCs / Primary Immune Cells | ≥ 1×10⁸ viable cells | Snap-freeze dry pellet or ship cryopreserved vials on dry ice. | Used for baseline profiling or characterizing specialized Antigen-Presenting Cells (APCs). |
| Plasma / Serum (sHLA) | 2 - 5 mL | Centrifuge to remove cells/debris, freeze aliquots at -80°C. Ship on dry ice. | Specialized workflow targeting soluble HLA complexes circulating in blood. |
| Optional Metadata | Clinical HLA Typing Report | Digital document transmission. | Highly recommended to enhance bioinformatics motif verification and exact allele assignment. |
Example Results from HLA Peptidomics Analysis
Our comprehensive deliverables include publication-ready visualizations that clearly characterize the discovered immunopeptidome landscape.
Peptide Length Distribution for HLA-I vs HLA-II
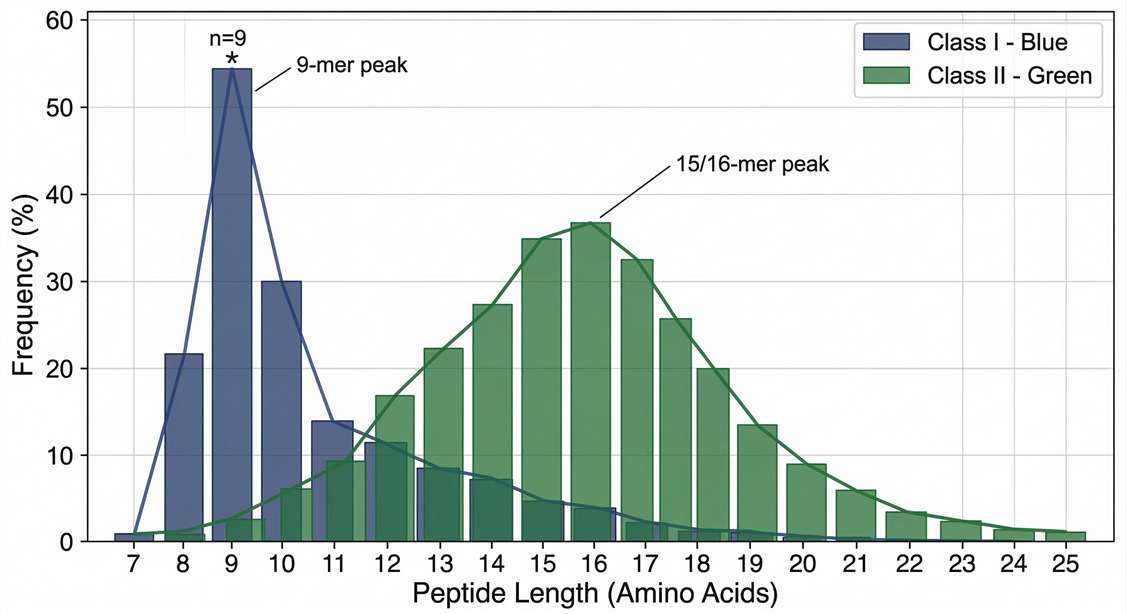
Characteristic peptide length distributions confirming successful and distinct enrichment of classical HLA class I and class II immunopeptidomes.
Motif Logos by HLA Allele
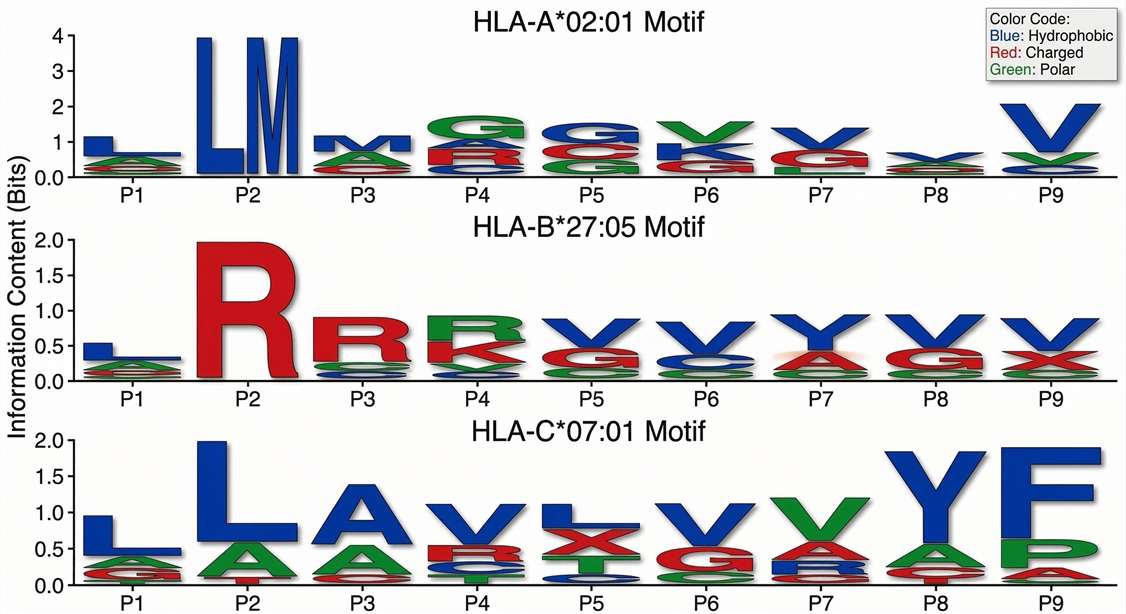
Sequence motif clustering of physically eluted peptides demonstrates strong adherence to the anchor residue preferences of the sample’s specific HLA allotypes.
Source Protein Annotation Summary
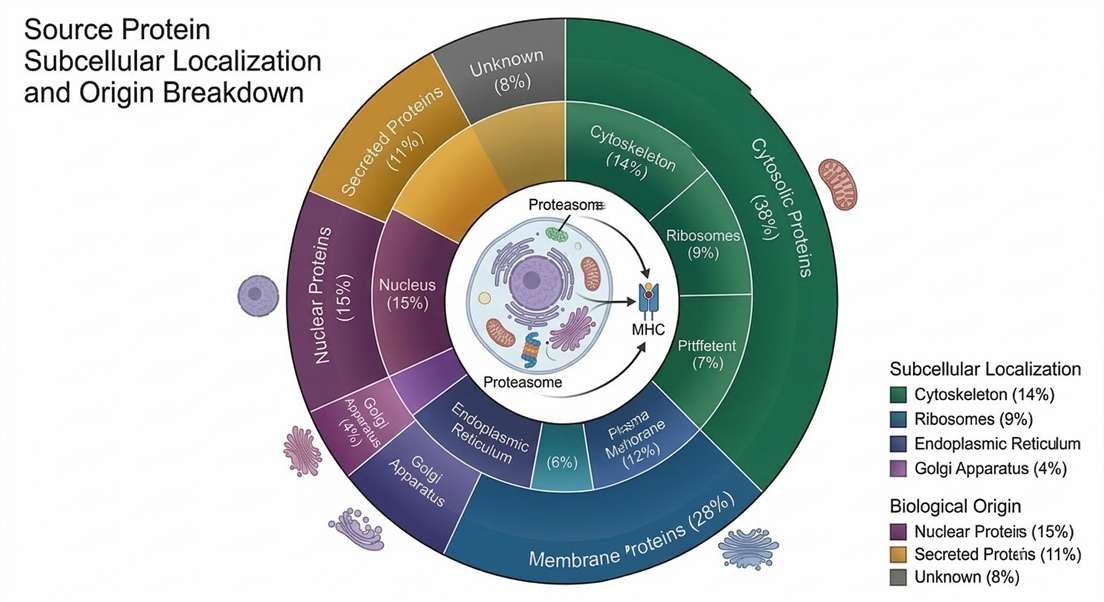
Subcellular localization and pathway mapping of the source proteins contributing to the global naturally presented immunopeptidome.
Representative Annotated MS/MS Spectrum
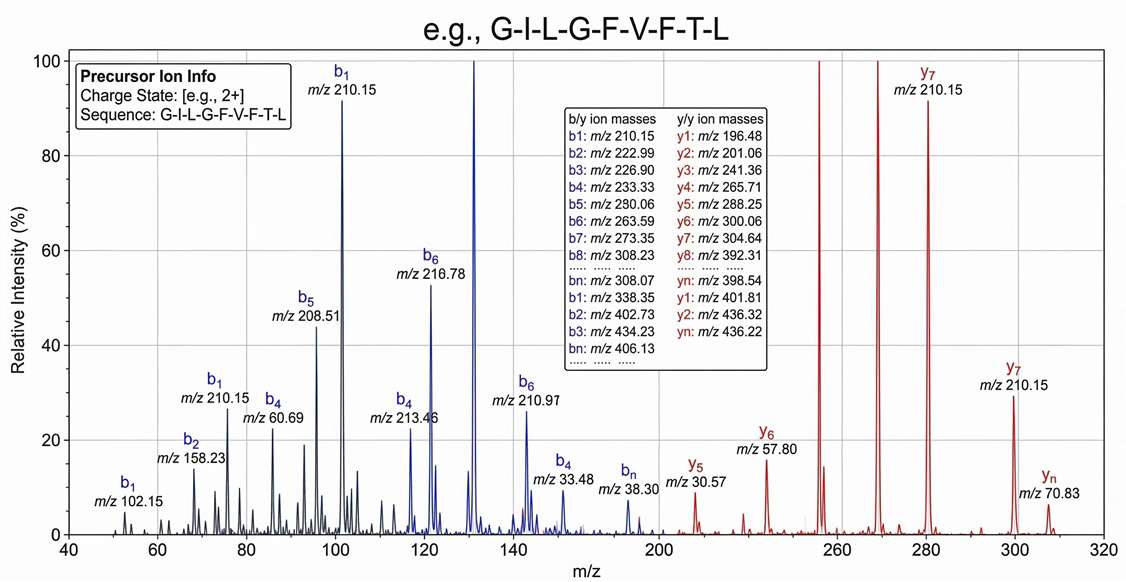
High-resolution tandem mass spectrum providing definitive physical proof of the exact amino acid sequence for a naturally presented target peptide.
Deliverables: What You Will Receive
Our HLA peptidomics service culminates in a highly structured data package, ensuring you have both the raw evidence and the interpreted insights needed to advance your project. Seamless transition paths are available if you wish to push identified targets into our tumor neoantigen discovery and validation services.
- Comprehensive Peptide Identification Matrix: A complete Excel/CSV dataset listing all identified peptides, mapped source proteins, MS intensity values, and confidence scores (FDR).
- Bioinformatics Annotation Report: Graphical visualizations of peptide length distributions, allele-specific binding motifs, and source protein biological pathway analyses.
- Raw MS Data Files: Full access to the original, high-resolution
.rawmass spectrometry files for your own internal archiving and independent verification. - Detailed Methodology Document: A complete, transparent record of the immunoaffinity protocols, LC-MS/MS parameters, and database search logic utilized in your specific project.
References
- Mass spectrometry–based identification of MHC-bound peptides for immunopeptidomics. Nature Protocols, 2019. https://doi.org/10.1038/s41596-019-0133-y
- Sensitive, High-Throughput HLA-I and HLA-II Immunopeptidomics Using Parallel Accumulation-Serial Fragmentation Mass Spectrometry. Molecular & Cellular Proteomics, 2023. https://doi.org/10.1016/j.mcpro.2023.100529
Frequently Asked Questions
Case Study: Flexible and Scalable Immunopeptidomics Pipeline for Epitope Mapping
Journal: Frontiers in Immunology
Published: 2024
Summary
To fully understand the adaptive immune response and advance vaccine design, researchers require precise mapping of the epitopes presented by diverse HLA alleles. This study demonstrates a robust immunopeptidomics pipeline utilizing artificial antigen-presenting cells to systematically identify and distinguish HLA Class I and Class II peptide signatures across multiple antigen and allele combinations, proving the value of direct peptide mapping over prediction-only strategies.
Methods
The research team developed a recombinant antigen-presenting system. This was combined with highly specific HLA peptide immunoaffinity enrichment and advanced LC-MS/MS-based immunopeptidomics analysis. By processing these engineered cells, the researchers could directly capture and sequence the exact peptides naturally processed and presented by specific HLA allotypes from selected model antigens.
Results
The pipeline successfully yielded high-confidence physical data mapping the immunopeptidome. Crucially, the MS analysis revealed distinct, characteristic peptide length distributions—sharply centered around 9-mers for HLA-I and broader 14-16mers for HLA-II—confirming accurate biological presentation.
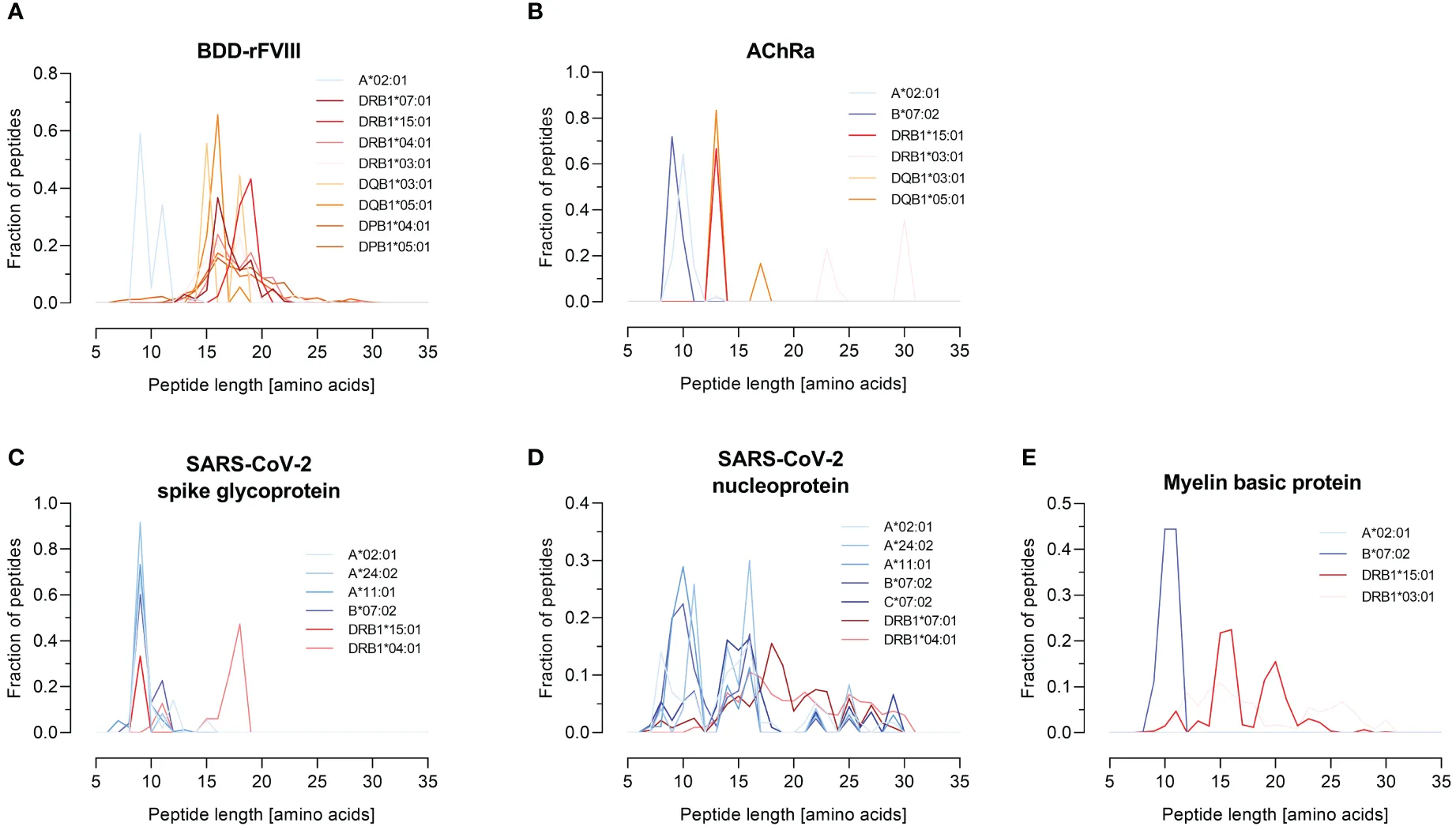
Characteristic peptide length distributions confirming the distinct presentation patterns of classical HLA class I and class II immunopeptidomes.
Furthermore, the study successfully generated allele-specific peptide signatures, demonstrating that direct mass spectrometry can accurately identify the precise epitope regions presented by different HLA variants, a task computational prediction alone often struggles to resolve accurately.
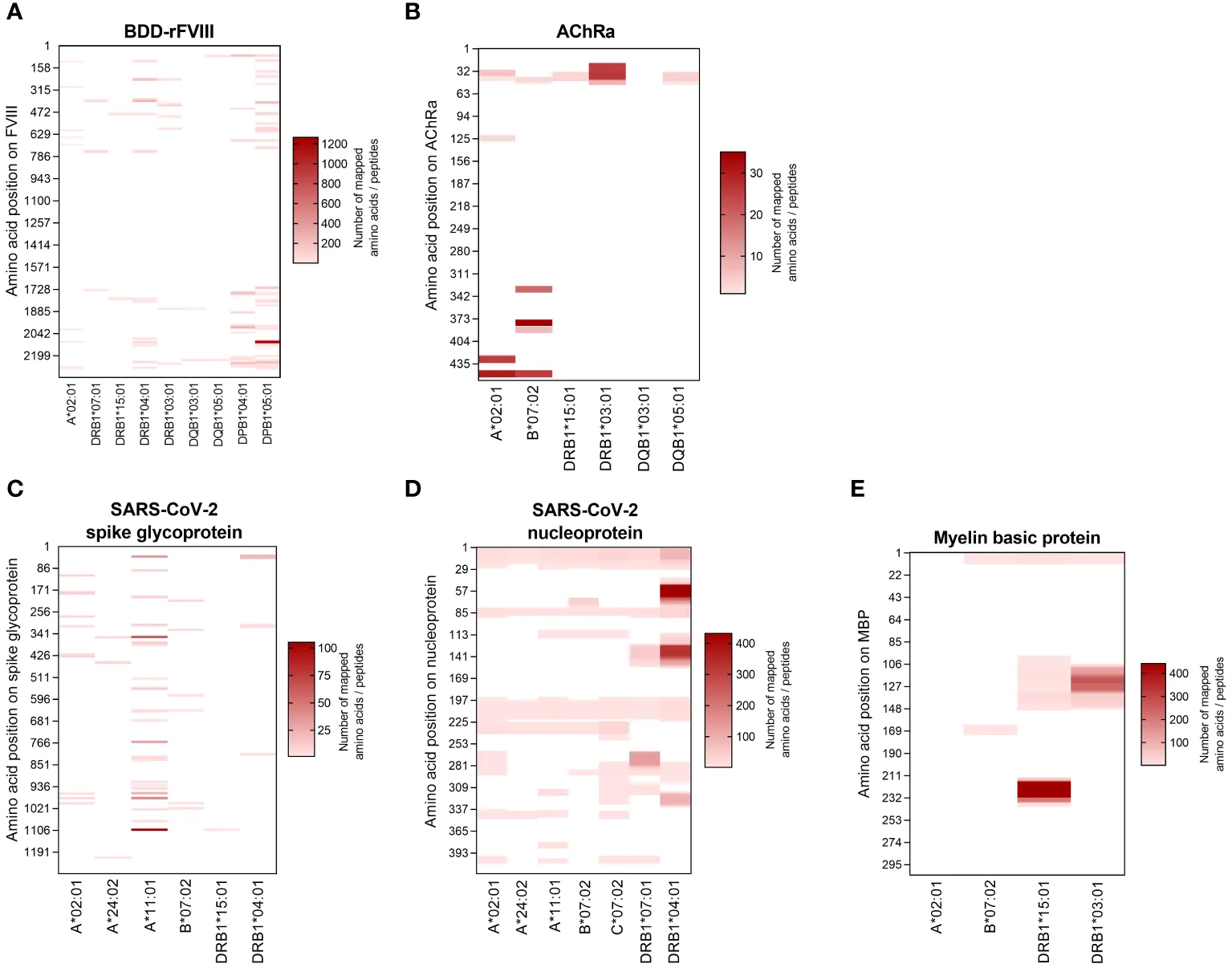
Heatmap visualization of allele-specific peptide signatures mapped across diverse human leukocyte antigen variants using direct mass spectrometry.
Reference
- Weisbrod L, et al. FASTMAP—a flexible and scalable immunopeptidomics pipeline for HLA- and antigen-specific T-cell epitope mapping based on artificial antigen-presenting cells. Frontiers in Immunology, 2024. https://doi.org/10.3389/fimmu.2024.1386160
For Research Use Only. Not for use in diagnostic procedures.