Resolving Media Interference in iPSC Neuron Secretome Profiling
Analyzing the secretome of induced pluripotent stem cell (iPSC)-derived neurons and 3D brain organoids presents unique analytical hurdles that standard proteomics platforms routinely fail to address. The primary obstacle is culture media interference. Supplements essential for neural survival—such as B27, N2, or residual serum—contain highly abundant proteins (e.g., albumin, transferrin) that create a massive mass-spectrometry background signal. This background effectively masks the low-abundance signaling factors, such as neurotrophins, cytokines, and chemokines, which define the true functional secretome.
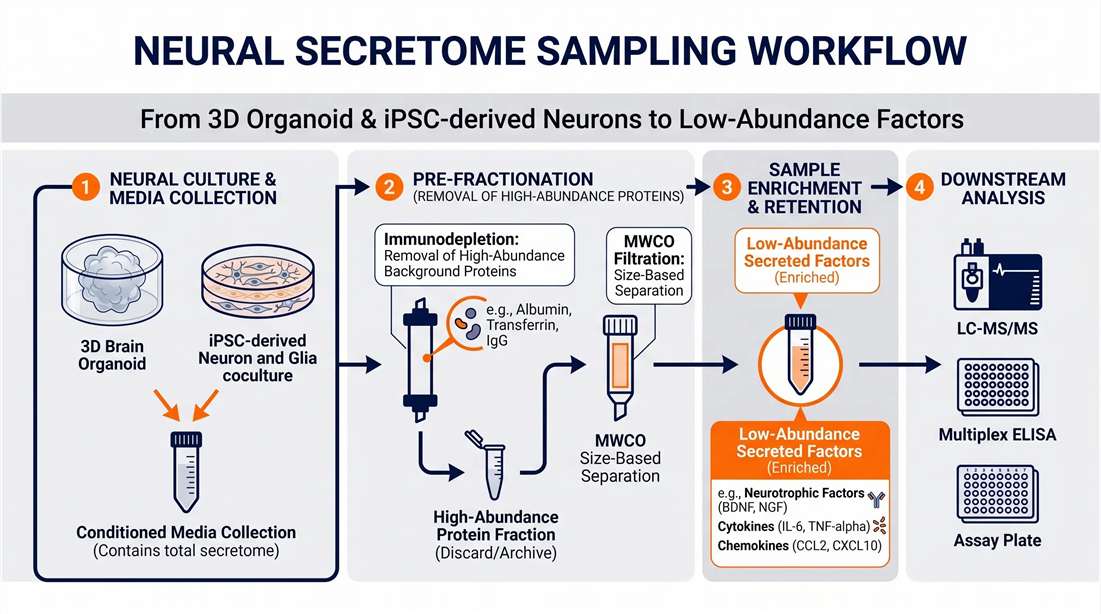
We do not treat neural secretomics as a generic fluid analysis. Our functional neuro-secretomics platform utilizes a hybrid analytical strategy. We work consultatively with your team to optimize short-term serum/supplement-free starvation windows. In our laboratory, we apply specialized molecular weight cut-off (MWCO) ultrafiltration and targeted peptide enrichment protocols, combined with optimized chromatographic elution gradients, to bypass high-abundance background peaks. This methodology allows us to dig deeper into the extracellular space to reveal the biologically relevant signaling molecules driving your neural models.
Strategic Research Applications for Neural Organoid Secretomics
The value of the neural secretome extends far beyond establishing basic protein lists. By capturing the dynamic output of neural cells, we help you map extracellular factors back to specific disease mechanisms and translational outcomes.
How We Distinguish True Secretion from Cell Death-Related Leakage
A critical concern in secretomics is determining whether an identified protein is actively secreted or simply leaked into the media due to cellular necrosis or apoptosis. We address this through a rigorous, multi-tiered Quality Control (QC) strategy:
- Intracellular Marker Filtering: We actively screen for and flag classic intracellular structural or metabolic proteins (e.g., GAPDH, β-Tubulin, Histones). High spectral counts of these markers indicate membrane leakage rather than active secretion.
- Secretion Pathway Prediction: We apply advanced algorithms such as SignalP 6.0 (for classical ER/Golgi-dependent secretion) and SecretomeP (for non-classical secretion) to annotate the active transport likelihood of every identified protein.
- Viability Assay Integration: We strongly recommend clients perform LDH release or parallel viability assays during the collection window to establish a baseline of cell health, which we correlate with our MS data.
- Paired Intracellular Proteome (Optional): By profiling both the intracellular proteome and the secretome, we calculate the Extra/Intracellular abundance ratio to definitively isolate active secretory fluxes from background noise.
Specialized DIA Quantitative Strategies for Brain Organoid Secretomics
Brain organoids and miniaturized iPSC arrays yield notoriously small volumes of conditioned media. While standard proteomics workflows require large starting volumes, our platform is designed for deep coverage of low-input samples.
We utilize Data-Independent Acquisition (DIA) on state-of-the-art high-resolution Orbitrap mass spectrometers (e.g., Exploris/Astral) as our primary quantitative strategy. DIA provides exceptional reproducibility, minimal missing values across biological replicates, and the sensitivity required to handle microliter-scale organoid supernatants. If your discovery phase identifies novel, low-abundance signaling peptides, we seamlessly transition the project into our targeted neuropeptide quantification pipelines for absolute validation.
Mass Spectrometry Platforms for Neural Secretomics
To deliver uncompromised depth on low-input neural models, our facility is equipped with a state-of-the-art analytical stack:
- Mass Spectrometry: High-resolution Orbitrap Exploris and Orbitrap Astral platforms, coupled with nano-LC systems for ultimate sensitivity.
- Data-Independent Acquisition (DIA): Advanced DIA methodologies processed through industry-leading software like Spectronaut or DIA-NN, ensuring deep proteome coverage and robust missing-value imputation.
- Bioinformatics: Beyond SignalP 6.0 and SecretomeP, we utilize R-based custom pipelines, Cytoscape for network visualization, and MaxQuant for baseline discoveries, ensuring your data is structurally ready for high-impact publication.
Technical Comparison: Standard Secretomics vs. Functional Neuro-Secretomics
| Feature | Standard Generic Secretomics | Our Functional Neuro-Secretomics |
|---|---|---|
| Media Background Handling | Assumes pure serum-free; struggles with B27/N2 | Hybrid depletion, MWCO filtration & LC gradient optimization |
| Purity / Leakage Control | Assumes all detected proteins are secreted | SignalP/SecretomeP annotation & intracellular marker QC |
| Bioinformatics Annotation | Generic GO / KEGG pathway enrichment | SynGO, neuroinflammation, and specific neural crosstalk |
| Organoid Compatibility | Requires large pooling (>10 mL) | Optimized for low-input supernatants (≥ 1.0 mL) |
| EV vs. Soluble Distinction | Analyzes total raw media | Modular options for EV-depleted soluble vs. isolated EV profiling |
Step-by-Step Workflow for Neural Secretome Profiling
Typical Results of iPSC Neuron and Organoid Secretomics
We transform raw mass spectrometry data into actionable biological models by integrating neuro-specific bioinformatics directly into our core deliverables.
Protein Identification Depth
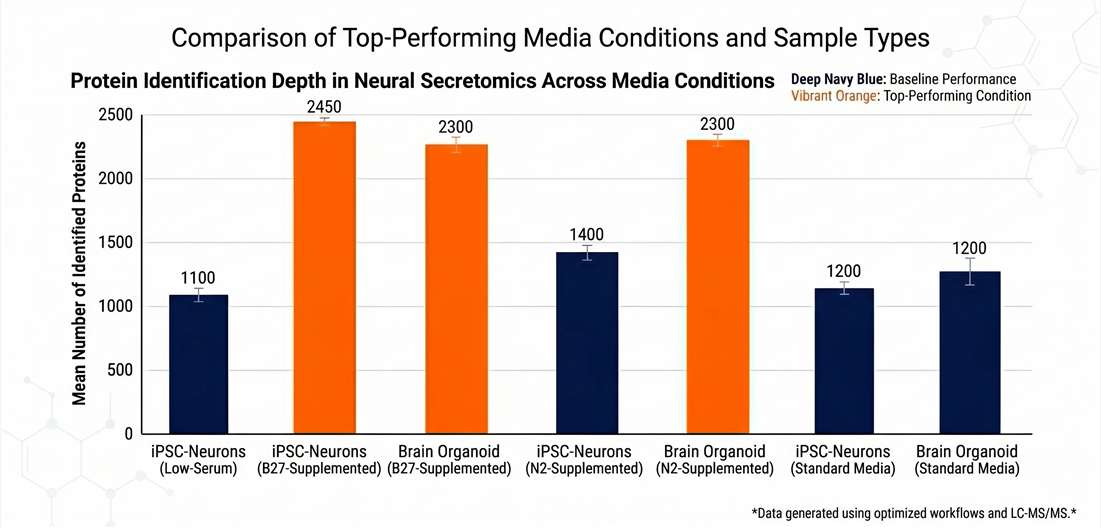
Validating signal recovery from microliter-scale media.
Secretion Purity Analysis
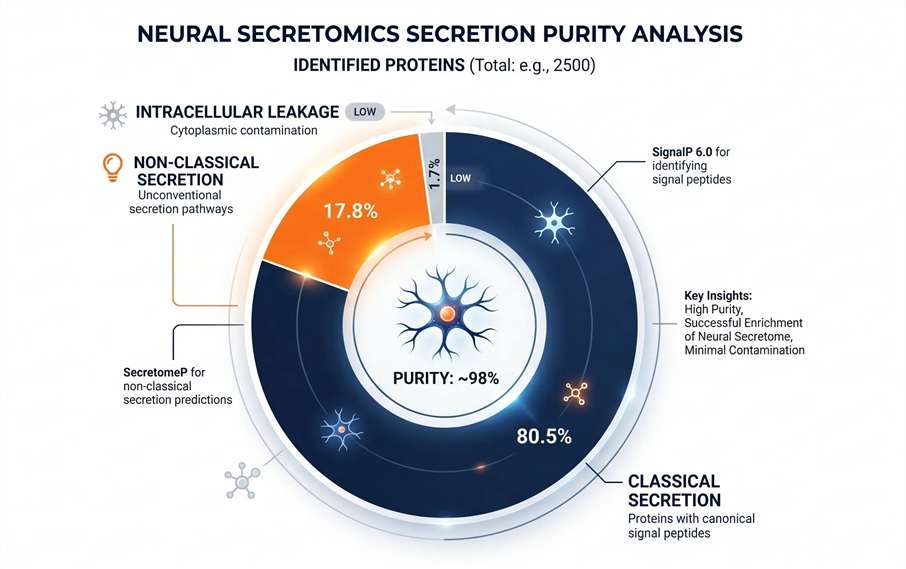
SignalP/SecretomeP modeling to prove data authenticity.
Neuro-Functional Enrichment
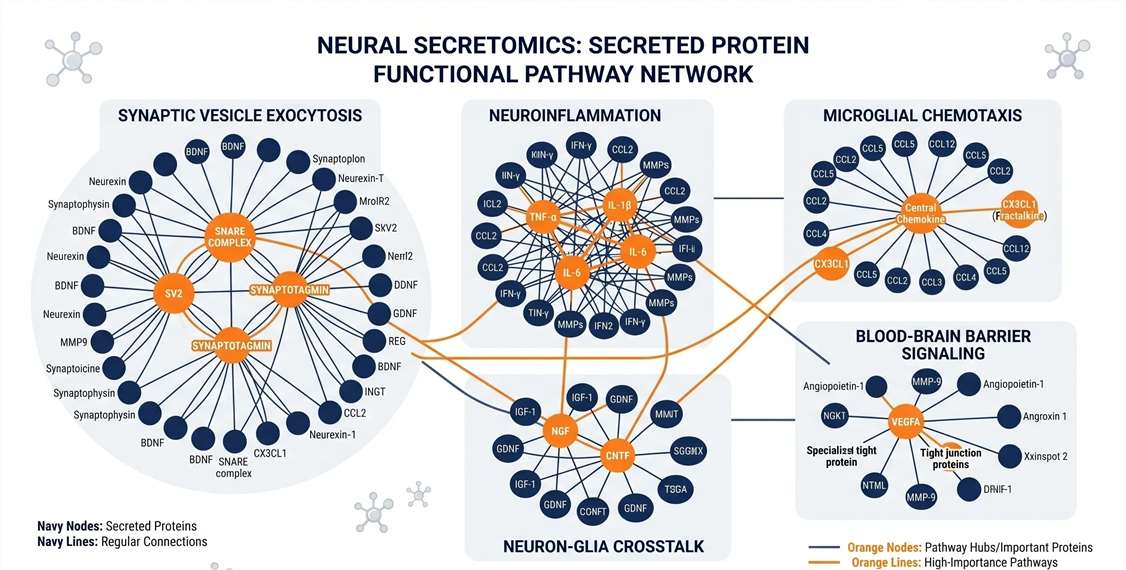
Mapping factors to SynGO and neuroinflammatory terms.
Disease/Drug Response
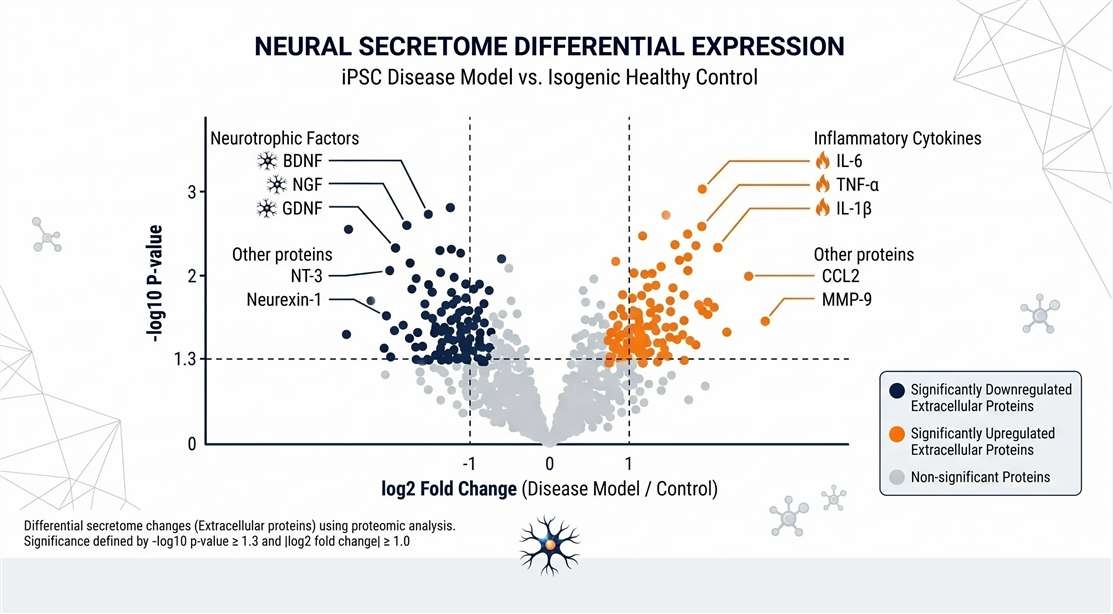
Highlighting functionally relevant extracellular shifts.
Typical Deliverables for Neural Secretome Studies
- Comprehensive identified protein list with raw and normalized abundances.
- Secretion Purity Report: Includes SignalP 6.0 scores and Intracellular Contamination QC summary.
- Neuro-Functional Analysis: Specialized SynGO enrichment plots and neuron-glia crosstalk models.
- Candidate biomarker shortlist and targeted validation recommendations.
- Datasets compatible for integration with untargeted neuropeptidomics or broader neuropeptide profiling studies.
Sample Requirements for iPSC and Organoid Secretomics
Proper design is vital for successful secretomics. We tailor our methods to your specific model.
| Sample Type | Recommended Volume | Recommended Protein | Key Experimental Inputs |
|---|---|---|---|
| 2D iPSC-Neurons | ≥ 1.0 mL | > 10 μg | Cell density, differentiation day, serum-free duration |
| 3D Brain Organoids | ≥ 1.0 mL | > 10 μg | Organoid size/count, media formulation, culture age |
| Co-culture Models | ≥ 2.0 mL | > 15 μg | Cell ratio (e.g., Neuron:Glia), stimulation/drug condition |
| EVs/Exosomes | ≥ 10.0 mL | Case-by-case | Preferred isolation method (e.g., ultracentrifugation) |
Conditioned Media Handling and Shipping Guidelines
The neural secretome is highly susceptible to rapid proteolytic degradation. To ensure the highest fidelity of your extracellular data, we strongly advise the following handling protocols prior to shipping:
- Immediate Preservation: Centrifuge collected media immediately at low speed (e.g., 300 x g for 5 mins) to remove whole cells, followed by a higher speed spin (e.g., 2000 x g for 10 mins) to remove cellular debris.
- Protease Inhibition: Addition of MS-compatible protease/phosphatase inhibitor cocktails is highly recommended immediately after collection.
- Snap Freezing: Aliquot the clarified supernatant into low-protein-binding tubes and snap-freeze in liquid nitrogen.
- Shipping: Ship all samples on dry ice. Our project managers will provide specific shipping logistics and import documentation for international clients.
Methodological Considerations in Brain Organoid Secretomics
Accurate interpretation requires considering the culture environment. Secretome changes are highly sensitive to the collection window; an excessively long window without media exchange can induce cellular stress, inflating intracellular leakage. Additionally, the maturity of a brain organoid dictates its secretory profile—early-stage organoids may secrete factors related to neurogenesis, while older organoids yield profiles rich in synaptic maintenance factors. Our team consults with you to account for these variables during the study design phase.
Disclaimer: All services and analytical platforms described are intended for Research Use Only (RUO). Not for use in diagnostic procedures.