Why Neural-Enriched EV Neuropeptidomics?
In translational neuroscience, the blood-brain barrier (BBB) strictly limits molecular monitoring of the central nervous system (CNS). While direct CSF and biofluid neuropeptidomics offers localized insights, serial sampling in preclinical models or clinical cohorts is highly invasive. Conversely, measuring free neuropeptides in peripheral plasma is severely restricted: CNS-specific signals are drowned out by high-abundance systemic proteins, and free peptides possess plasma half-lives of mere seconds due to ubiquitous circulating peptidases like neprilysin.
Extracellular vesicles (EVs) offer a revolutionary analytical alternative. Brain cells continuously secrete EVs that encapsulate active signaling molecules—including intact neuropeptides and their prohormone precursors—transporting them across the BBB. The robust EV lipid bilayer acts as molecular armor, physically protecting these fragile peptide payloads from enzymatic degradation in the bloodstream.
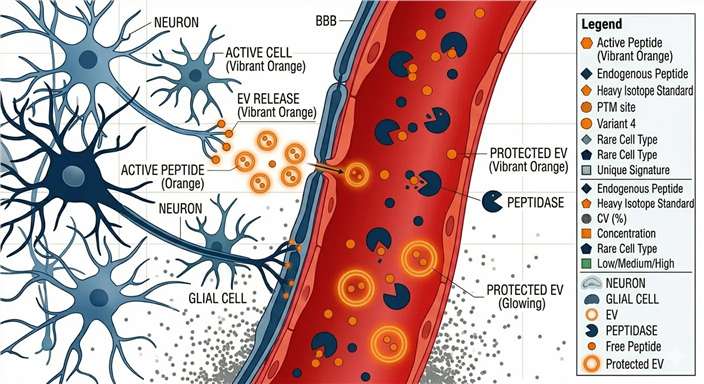
By selectively capturing these CNS-derived vesicles, our service provides a critical research window into the CNS. This strategy drastically improves the probability of detecting neural-associated peptide signals, empowering researchers to track neuromodulation and prohormone processing dynamics from a standard blood draw.
Decision Matrix: Selecting the Right Analytical Approach
When designing your preclinical translational study, comparing available omics platforms ensures you select the workflow that accurately matches your core biological question:
| Assay Goal | Recommended Platform | Why Choose This? |
|---|---|---|
| Discover global systemic changes | Total EV Proteomics | Ideal for oncology or organ-agnostic discovery. Misses low-abundance CNS peptides. |
| Profile high-concentration local fluids | Direct Biofluid Peptidomics | Best for CSF or direct cell culture media where CNS signals are not diluted by systemic blood. |
| Trace CNS signals from peripheral blood | CNS-EV Neuropeptidomics | Specifically enriches neural-associated EV cargo from complex plasma, overcoming the BBB sampling barrier. |
| Validate findings in large cohorts | Targeted PRM/MRM Assays | Transition here after discovery. Provides absolute quantification using heavy stable isotope standards. |
Recommended Research Applications for Preclinical Discovery
This platform is specifically designed to support discovery and preclinical decision support. It is the ideal methodological choice for the following high-value research scenarios:
- Alzheimer’s disease and related dementias exploratory biomarker discovery and progression tracking.
- Parkinson’s disease and synucleinopathies longitudinal neurochemical profiling.
- Neuroinflammation and neuroimmune communication mapping across the gut-brain axis.
- Preclinical pharmacology and CNS-active drug response profiling to confirm target engagement.
- iPSC-neuron, organoid, and translational biofluid signaling studies bridging in vitro models to in vivo relevance.
Study Boundary Statement: This service is intended strictly for research use, exploratory biomarker discovery, and preclinical decision support. It is not designed to establish a clinical diagnosis from a single peripheral sample.
Representative CNS-EV Neuropeptide Families We Detect
Researchers often require confirmation that their specific target of interest can be successfully recovered from peripheral EVs. Our advanced LC-MS/MS analytical pipeline routinely maps the following critical neuropeptide families:
| Neuropeptide Family | Representative Targets in EVs | Biological & Translational Relevance |
|---|---|---|
| Endogenous Opioids | Enkephalins (Met-/Leu-), β-Endorphin, Dynorphins A/B, Nociceptin | Pain modulation, acute/chronic stress response, addiction models |
| Hypothalamic Peptides | Orexin A/B, CRH, TRH, Somatostatin, MCH, GnRH | Sleep/wake cycles, systemic energy metabolism, severe stress |
| Tachykinins | Substance P, Neurokinin A/B, Hemokinin-1 | Neurogenic inflammation, nociception, traumatic brain injury |
| Neuropeptide Y Family | NPY, PYY, Pancreatic Polypeptide (PP), AgRP | Metabolic disorders, feeding behavior, anxiety-like states |
End-to-End Workflow for Neural-Enriched EV Profiling
We process your biofluid samples through a meticulously controlled pipeline designed specifically to preserve vesicle integrity and native peptide boundaries, entirely bypassing the tryptic-digestion bottlenecks of conventional proteomics.
Methodological Considerations for EV Analysis
Neural markers (such as NCAM or L1CAM) are used to enrich CNS-relevant EV subpopulations; therefore, results should be interpreted as neural-enriched or CNS-relevant EV-associated peptide signatures rather than an absolute guarantee of 100% exclusive brain origin.
To maximize biological confidence, our pipeline relies on strict analytical control:
- Advanced Isolation: Combining SEC with affinity capture prevents massive lipoprotein (HDL/VLDL) and soluble protein co-isolation, minimizing mass spectrometry ion suppression.
- Orthogonal QC: Mandatory verification of EV integrity using NTA and TEM prevents the misidentification of random protein aggregates as vesicular cargo.
- Contamination Awareness: Specialized extraction buffers ensure that high-abundance luminal structural proteins do not mask the low-abundance neuropeptidome.
- Rigorous Study Design: We strongly advise robust cohort designs with well-matched controls to ensure biologically meaningful and statistically significant outcomes.
Demo Results: Visualizing CNS Signaling in Peripheral Blood
To ensure the highest biological confidence, we provide multi-dimensional data visualizations that validate the CNS relevance and structural integrity of the detected signals.
NTA & TEM Characterization
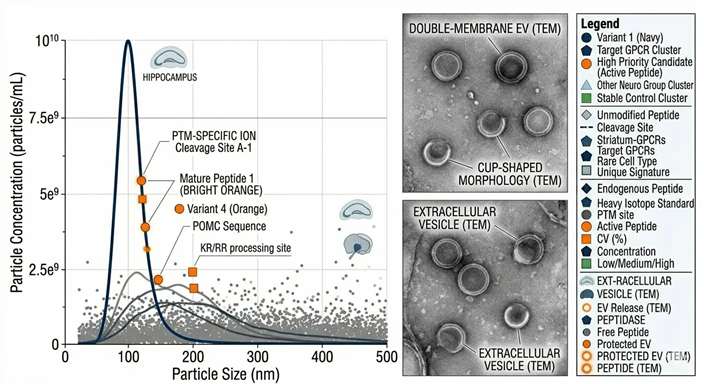
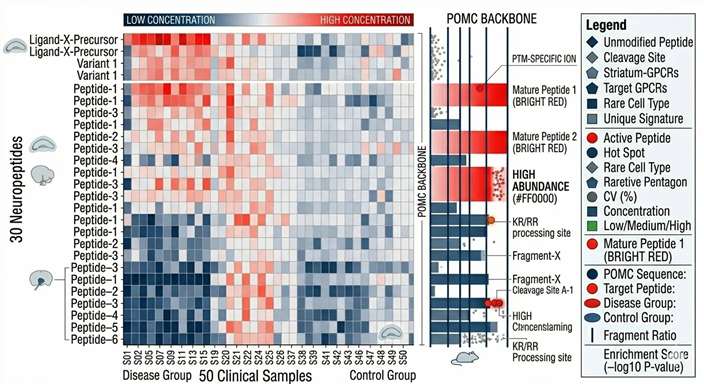
Differential Expression Analysis
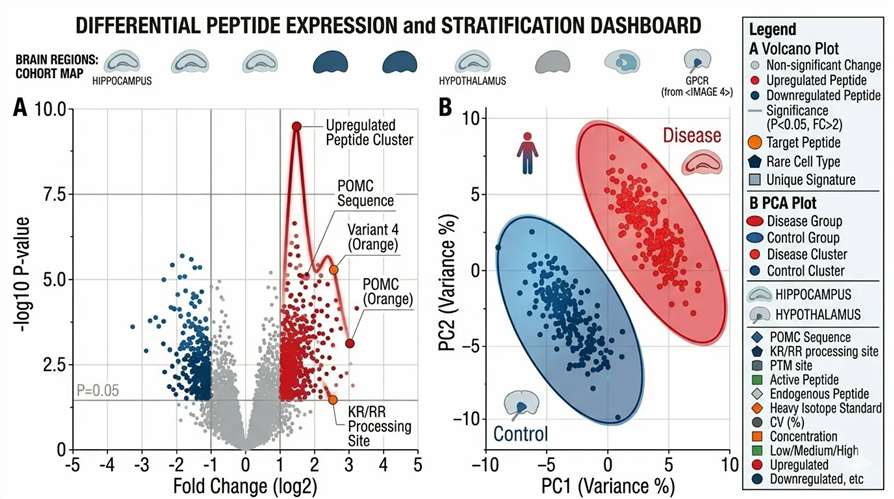
Native PTM MS/MS Mapping
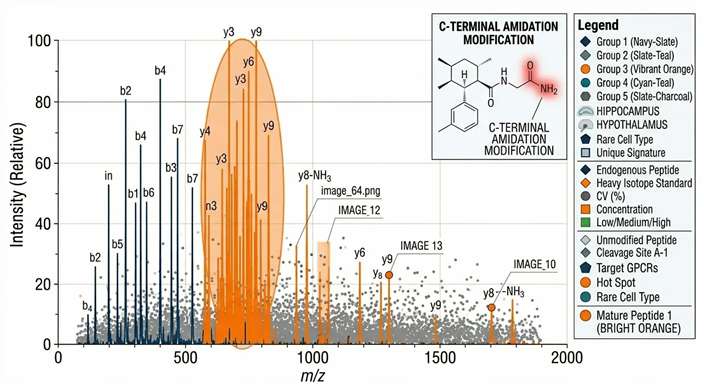
Prohormone Cleavage Dynamics
Typical Deliverables: What You Receive
We deliver actionable, transparent, and publication-ready data assets formatted for immediate integration into your preclinical research pipelines. Your comprehensive computational package includes:
- EV Characterization Summary: NTA size distribution charts and TEM micrographs visually confirming successful vesicle isolation.
- Identified Endogenous Peptide List (CSV): A complete dataset of native peptides, retention times, spectral counts, and strict target-decoy validated FDR confidence scores.
- PTM Annotations: Detailed neuropeptide PTM analysis mapping critical native modifications that dictate bioactivity (e.g., C-terminal amidation, sulfation).
- Relative Quantification Matrix: Formatted raw and normalized MS intensities perfectly aligned across all analyzed samples.
- Group Comparison Statistics: Volcano plots, PCA clustering, and differential expression data highlighting condition-specific EV peptide signatures.
- Pathway & Prohormone Processing Interpretation: Bioinformatic reconstruction of the enzymatic cleavage events found within the EV cargo.
- Target Candidate List: A bioinformatically curated shortlist of high-potential candidates ready for downstream targeted neuropeptide quantification.
- Raw Data Access: Raw mass spectrometry data files (.raw) are readily available upon request.
Cohort-Level Interpretation Note: Due to inter-individual variability in peripheral EV abundance and complex physiological states, we strongly advise cohort-level interpretation with well-matched controls rather than drawing diagnostic inferences from single individual samples.
Sample Requirements & Pre-Analytical Considerations
The success of any EV neuropeptidomics project relies heavily on upstream sample quality. We require the following sample specifications for optimal recovery:
| Sample Type | Minimum Volume | Recommended Tube / Prep | Shipping Condition |
|---|---|---|---|
| Plasma (Preferred) | 1.0 mL | EDTA tubes; Double-spun to yield platelet-poor plasma (PPP) | Dry ice |
| Serum | 1.0 mL | Standard serum separator tubes; Centrifuge promptly | Dry ice |
| CSF | 200 μL | Low-bind tubes; Centrifuge to remove cell debris | Dry ice |
| Cell Culture Media | 10.0 mL | Use serum-free or EV-depleted FBS; Pre-cleared of cells | Dry ice |
To assist in your cohort planning, we recommend adhering to these pre-analytical handling principles:
- Plasma vs. Serum: Plasma is strictly preferred for EV discovery workflows. Clotting cascades physically alter background peptide levels and trigger the release of massive amounts of platelet-derived EVs, masking CNS signals.
- Hemolysis & Platelets: Ensure rigorous double-spin protocols. Platelet contamination can severely distort EV abundance calculations and cargo composition. Discard visibly hemolyzed samples.
- Freeze-Thaw Cycles: Minimize repeated freeze-thaw cycles, which compromise vesicle membrane integrity. Aliquot samples precisely prior to initial freezing at -80°C.
- Phased Study Design: For large cohorts, we recommend a pilot feasibility study (5-10 samples) prior to a full-scale untargeted neuropeptidomics discovery run.
Disclaimer: All services and platforms described are for Research Use Only (RUO). Not for use in diagnostic procedures.