Characterization Analysis of Antibody-based Drugs
Beyond the routine process control and release testing of each batch of antibody-based drugs, it is necessary to conduct deeper analytical investigations within the realm of biochemical and biophysical characterization. The data generated from these analyses actively supports the clinical development of antibody drug formulations, confirmation of process scaling, and can affirm the consistency of the product post-translational modifications or technology transfers. Not every batch of antibody-based drugs requires characterization analysis, but the chosen analytical methods should endeavor to comprehensively represent and affirm the product structure and its variations. It is important to note that some characterization analytical tools are equally applicable for in-process control and release testing. For instance, peptide mapping is versatile, accommodating for primary structure, disulfide bond, or post-translational modification analyses. Simultaneously, it can also fulfill ordinary identification tests to meet the demands of release testing.
The table below lists common characterization analysis methods for antibody drugs and the product structural information they provide. Detailed interpretation of some representative characterization methods is to be discussed in this article.
Table 1: Characterization Methods for Monoclonal Antibodies
| Analysis Method | Provided Structural Information |
| Protein Structure Characterization | |
| N-terminal and C-terminal Sequencing | Confirmation of terminal sequences |
| Peptide Mapping with Mass Spectrometry or Sequencing Analysis | Primary, secondary structure, and post-translational modifications |
| Free Thiol and Disulfide Bond Detection | Confirmation of disulfide bonds |
| Fluorescence Chromatography | Quaternary structure |
| Mass Spectrometry Analysis | Primary structure, post-translational modifications, complete mass |
| Isoelectric Focusing | Charge variants, oxidation, deamidation, sialic acid content |
| Fourier Transform Infrared Spectroscopy (FTIR) | Secondary structure |
| Circular Dichroism (CD) | Secondary and tertiary structure |
| Nuclear Magnetic Resonance (NMR) | Overall integrity confirmation |
| Differential Scanning Calorimetry (DSC) | Structural stability, reflection of correct tertiary structure |
| Aggregate Characterization | |
| Multi-Angle Laser Light Scattering (MALLS) | Detection of aggregates and low molecular weight variants |
| Field-Flow Fractionation (FFF) | Detection of aggregates and low molecular weight variants |
| Analytical Ultracentrifugation (AUC) | Detection of aggregates and low molecular weight variants |
| Glycosylation Characterization | |
| Enzymatic Digestion + LC-MS/MS | N-glycosylation site detection |
| Mass Spectrometry + Fluorescence Detection | Glycoform and oligosaccharide content ratio |
| Activity Characterization | |
| Surface Plasmon Resonance (SPR) | Affinity detection and other molecular interactions |
| Bio-Layer Interferometry (BLI) | Affinity detection and other molecular interactions |
The sequencing of N-terminus and C-terminus confirms that the antibody drugs have the correct N-terminal and C-terminal sequences, which are means of determining the primary structure of the product as well as identifying different forms of antibody drugs. Typically, the standard Edman degradation method is used for N-terminal sequencing, wherein the first 5-10 amino acid residues of heavy and light chains are determined and compared with the anticipated sequences; peptide mapping may also be used to confirm the N-terminal sequence. To confirm the C-terminal amino acid sequence, the antibody is usually enzymatically cleaved into peptides, which then undergo peptide sequencing; this process is often conducted as part of peptide mapping.
Select Service
Learn more
- Application of Mass Spectrometry in Antibody Drug Development: Disulfide Bond Analysis
- Analysis Strategies of Antibody Charge Variants
- Introduction and Characterization of Monoclonal Antibody Drugs
- Application of Mass Spectrometry Technology in Antibody Drug Analysis
- Analytical Methods for Antibody-Drug Conjugates (ADCs)
- Mass Spectrometry-based Antibody De Novo Sequencing: Applications, Advances, and Approaches
- Biological Analysis Methods of Monoclonal Antibody Drugs
Peptide Mapping Analysis
As previously mentioned, during the characterization process of antibody-based therapeutics, it often necessitates protein digestion via proteolytic enzymes, subsequently breaking down the protein scaffold into peptide fragments. Following this step, the resultant peptide fragments are analyzed through reversed phase high-performance liquid chromatography, a technique referred to as peptide mapping analysis.
Through peptide mapping analysis, one can identify degradation products of the protein, such as deamidation of asparagine or oxidation of methionine residues; confirm or verify the connections of disulfide bonds; determine the positions of glycosylation. Also, in conjunction with mass spectrometry, this method can be utilized for protein identification, sequence analysis and so forth. The application of peptide mapping underscores its significant role in analytical aspects of biomolecul.
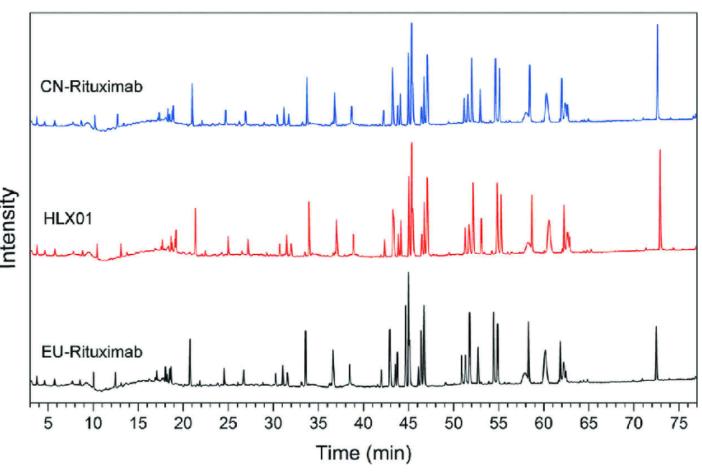 Figure 1: Peptide Mapping Analysis of Representative Monoclonal Antibody Drug (Yanpeng Xu et al,. 2019)
Figure 1: Peptide Mapping Analysis of Representative Monoclonal Antibody Drug (Yanpeng Xu et al,. 2019)Mass Spectrometry Analysis
Mass spectrometry (MS) is a powerful technique that produces charged particles from proteins through various methods, followed by accelerating and separating these proteins through the combined influence of magnetic and electric fields. The acceleration of proteins is directly proportional to their mass-to-charge ratio and these proteins are captured and further analyzed in the detector. The ionization of proteins can be achieved using techniques such as fast atom bombardment (FAB), matrix-assisted laser desorption/ionization (MALDI), and electrospray ionization (ESI). The detectors typically include time-of-flight (TOF) and quadrupole types. Mass spectrometers are generally classified based on the type of ionizer and detector; for example, a MALDI-TOF mass spectrometer.
Mass spectrometry can provide precise molecular weight data for antibody drugs. Furthermore, by inducing fragmentation of the antibody, information related to sequence, such as the molecular weight of the light and heavy chains, can be obtained. The use of tandem mass spectrometers (MS/MS) facilitates the generation of additional antibody fragments. At present, with its high resolution and sensitivity, the MS/MS technology can provide information regarding aspects such as amino acid oxidation, deamidation, and primary structure.
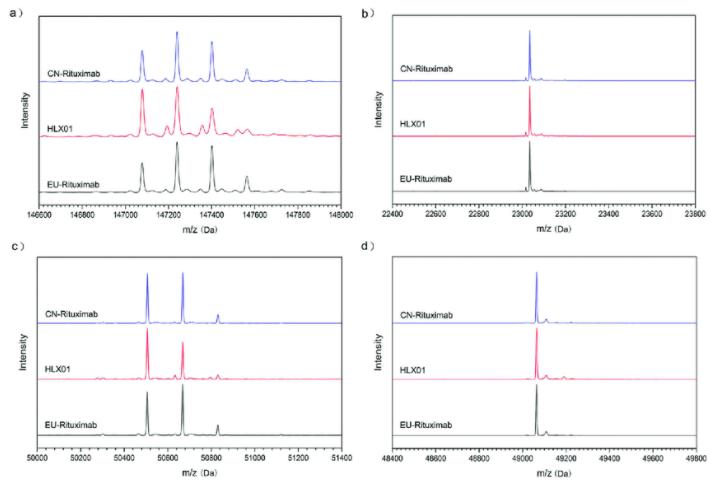 Figure 2: Mass spectrometric analysis of a typical monoclonal antibody drug: he MS spectra for intact mAb (a), reduced LC (b), reduced HC (c) and reduced and deglycosylated HC (d) (Yanpeng Xu et al,. 2019)
Figure 2: Mass spectrometric analysis of a typical monoclonal antibody drug: he MS spectra for intact mAb (a), reduced LC (b), reduced HC (c) and reduced and deglycosylated HC (d) (Yanpeng Xu et al,. 2019)Multi-Attribute Method (MAM)
The MAM is attracting growing attention and is widely applied within the biopharmaceutical sector. This method originates from liquid chromatography-mass spectrometry (LC-MS) and serves as a foundational technique in peptide mapping. The current iteration of MAM is a multi-purpose analytic tool that converges detection, identification, quantification, and quality control (QC) into a comprehensive system. This exhibits the exceptional value of MAM, particularly when dealing with novel molecular entities.
During the manufacturing process of antibody products, enzymatic digestion (typically using trypsin) is used to treat samples, and subsequently, they undergo LC-MS or LC-MS/MS analysis to provide a comprehensive assessment of the quality attributes of the product. Key quality attributes (KQAs) can be singularly quantified when they are recognized as such. Furthermore, MAM encompasses New Peak Detection (NPD) feature for the exploration of degradants or variants of the product.
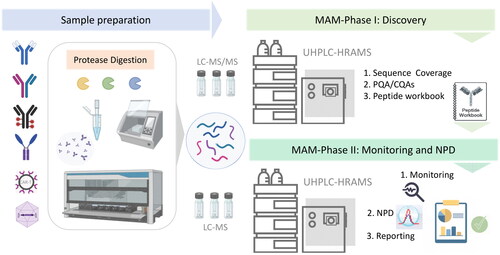 Figure 3: Typical workflow of Multi-Attribute Method (MAM) (Silvia Millán-Martín et al,. 2023)
Figure 3: Typical workflow of Multi-Attribute Method (MAM) (Silvia Millán-Martín et al,. 2023)NMR
Nuclear Magnetic Resonance (NMR) serves as an extraordinarily competent procedure to characterise antibody medications, offering up intricate data sets that aid the elucidation of these drugs' compositional structure. Within the NMR modality, the transitions of atomic nuclei within a magnetic field are observed closely, facilitating a comprehensive understanding of the specific three-dimensional structure, conformational dynamics, as well as converse interactions located within the antibody molecules.
In the realm of conformational dynamics, Nuclear Magnetic Resonance (NMR) stands as a multifaceted tool. Beyond the confines of mere static structures, NMR can illuminate the provisional states of molecules in solution. This is accomplished by monitoring fluctuations in peak intensity and spacial placement, thus allowing for an insightful interpretation of the conformational transitions within antibody therapeutics. These transitions encapsulate both the oscillations of flexible regions and modifications in the overarching protein structure.
Additionally, NMR is instrumental in investigating the interplay between antibodies and their counterparts - be they ligands or disparate molecules. This is of paramount importance for deciphering the binding mechanics of antibodies to their respective targets, gauging the impact of binding affinity, and identifying potential anomalies.
Beyond its realm of traditional usage, NMR also proves critical in uncovering post-translational alterations, encompassing but not limited to glycosylation and methylation. By proffering samples to NMR scrutiny, one can discern specific types and locations of these modifications. Thus, it effectively deepens our comprehension of antibody-based drugs, augmenting our fundamental understanding of their functional and structural dynamics.
Spectroscopic Methods
The investigation of therapeutic antibodies frequently utilizes spectroscopic methods such as circular dichroism (CD), Fourier Transform Infrared Spectroscopy (FTIR), and fluorescence spectroscopy. These techniques hold pivotal roles in the comprehensive characterization analysis of such drugs, due to their ability to facilitate a thorough understanding of the antibodies' structure and function on a molecular level. The in-depth analytical data produced by these methodologies not only implicates the drug's potential efficacy and safety profiles but also provides critical insights during the various developmental and quality control stages in the pharmaceutical industry.
Circular dichroism (CD), a light absorption phenomenon, operates on the principle of differential absorption of left and right polarized light by molecules such as antibodies, primarily in the far-ultraviolet (190-250 nm) or near-ultraviolet regions (230-250 nm). Discrepancies in molecular structures manifest as differential light intensity signals, enabling the analysis of said structures. The far-ultraviolet CD is particularly sensitive to secondary structural alterations, while the near-ultraviolet CD primarily detects changes to tertiary structures. From the standpoint of its non-destructive nature, CD analysis allows for quick data acquisition and provides critical insights under a variety of conditions, including varied pH or temperatures. Nonetheless, a notable limitation is the requirement for relatively higher sample concentration, typically in the range of 0.5-1.0 mg/ml.
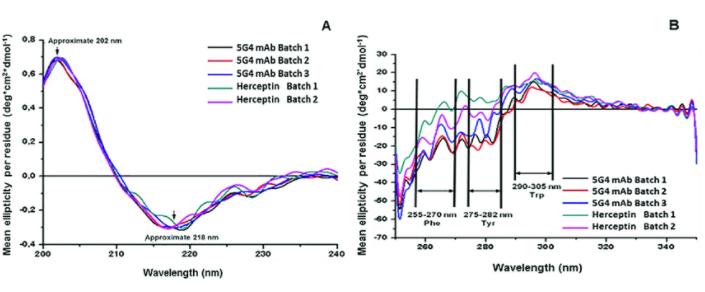 Figure 4: Circular Dichroism (CD) Spectra of Monoclonal Antibody Drug (a) far-UV CD spectrum; (b) near-UV CD spectrum. (Leina Pérez et al,. 2019)
Figure 4: Circular Dichroism (CD) Spectra of Monoclonal Antibody Drug (a) far-UV CD spectrum; (b) near-UV CD spectrum. (Leina Pérez et al,. 2019)Fourier Transform Infrared (FTIR) spectroscopy functions by measuring the interaction between molecular dipoles and incident energy at specific energies and the absorption of incident infrared radiation. This thereby facilitates the analysis of a protein's secondary structure, such as the relative quantities of α-helix or β-sheet.
Fluorescence Spectroscopy operates through the stimulation of proteins using near-ultraviolet to blue laser light wavelengths, triggering the characteristic fluorescence of tryptophan residues in the antibody drugs. The variability in fluorescence spectroscopy can reflect the microenvironment of the tryptophan residues, hence featuring a high sensitivity to tertiary protein structures. Factors such as denaturation, degradation, or aggregation, can trigger changes in the tertiary structure, in turn causing alterations in the maximum fluorescence wavelength or intensity.
Differential Scanning Calorimetry (DSC)
Differential Scanning Calorimetry (DSC) is designed to provide pertinent data regarding the thermal stability of antibody drugs. In these therapeutics, regions folded into thermodynamically most stable structures require more energy compared to less stable areas. The denaturation midpoint temperature (Tm), which is the temperature at which 50% of the molecules in a sample are yet to unfold, is a function of the tertiary structure of the molecules. Notably, multiple denaturation midpoints are typical of antibody drugs.
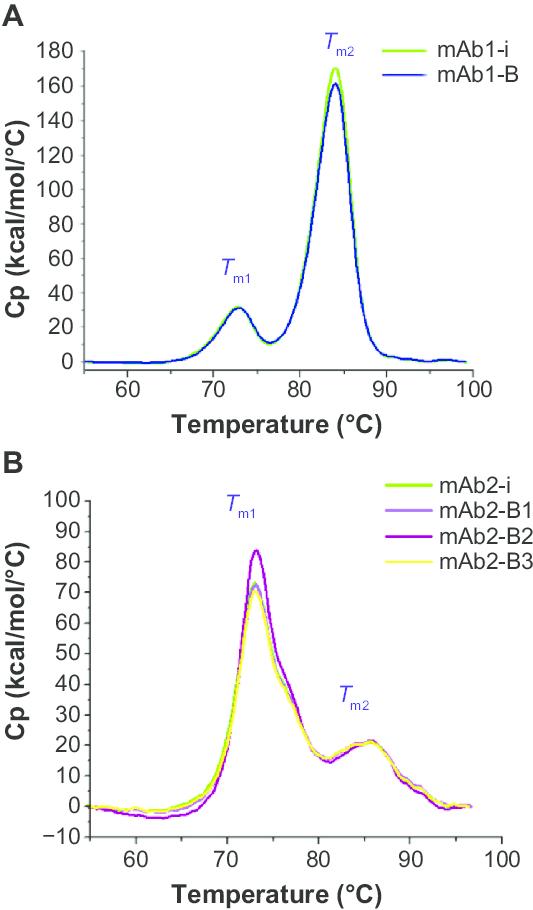 Figure 5: DSC analysis of mAbs. (A) Thermal stability profile of mAb1 and (B) thermal stability profile of mAb2. (Uma Sinhadatta et al,. 2015)
Figure 5: DSC analysis of mAbs. (A) Thermal stability profile of mAb1 and (B) thermal stability profile of mAb2. (Uma Sinhadatta et al,. 2015)Characterization of Aggregates
In the study of aggregates in antibody drugs, techniques like SEC-HPLC or SDS-PAGE are typically employed for process control and release testing. However, to conduct a more comprehensive examination of antibody drugs, sophisticated analytical technologies such as multi-angle laser light scattering (MALLS), analytical ultracentrifugation (AUC), and field flow fractionation (FFF) are also utilized for the characterization of aggregates.
Multi-Angle Laser Light Scattering (MALLS) assesses the scattering behavior of proteins in solution to known light, from which the molecular weight of molecules in solution can be calculated. Analytical Ultracentrifugation (AUC) is a modality that analyzes proteins or their aggregates based on their migration rate through gradient solutions in a centrifugal field, a rate that has a functional relationship with the molecular radius. Field Flow Fractionation (FFF), on the other hand, is a technique that involves the application of a field perpendicular to the sample flow that stratifies solutes in narrow channels, fixing them onto a membrane. The movement of molecules along the flow path has a functional relationship with the molecular weight. This tool exhibits capabilities for analyzing both covalent and non-covalent aggregates simultaneously.
Glycoform Characterization
A crucial component in the characterization analysis of antibody drugs lies in glycoform analysis, including glycosylation sites, glycoforms, and amounts of each oligosaccharide. Peptide mapping is used to identify the glycosylation sites and site occupancy rates within monoclonal antibody products. The specific location of each glycosylation site within the product can be determined through MS/MS or N-terminal sequencing analysis of the peptides released during peptide mapping. The percentage of occupancy at each site can be ascertained by measuring the relative amounts of peptides containing the glycosylation site, in both glycosylated and non-glycosylated states.
To define each glycoform structure, the intact oligosaccharides are first excised from the monoclonal antibody using the enzyme PNGaseF, before HPLC is utilized to separate different oligosaccharide structures for consequent analysis through one or multiple methods.Identification of each oligosaccharide structure can be performed by comparing the elution times of the samples with a series of standard oligosaccharides run under identical chromatographic conditions, or alternatively, the oligosaccharide structures can be delineated through mass spectrometry. By integrating these oligosaccharide structure results with previously determined monosaccharide composition data, the accuracy of glycoforms can be assured.
Functional Characterization
Surface Plasmon Resonance (SPR) technology is an analysis method grounded on optical principles. When light totally reflects at the interface between a prism and metal film, an evanescent wave is produced which penetrates into the optical sparse medium. In the medium, the existence of a plasmon wave forms a resonance phenomenon upon meeting the evanescent wave due to the energy conservation principle. SPR technology can be applied to evaluate drug-related functional activity, including antibody-antigen affinity, binding activity of Fc with FcγRIa, FcγRIIa, FcγRIIb, FcγRIIIa, FcγRIIIb, FcRn and C1q.
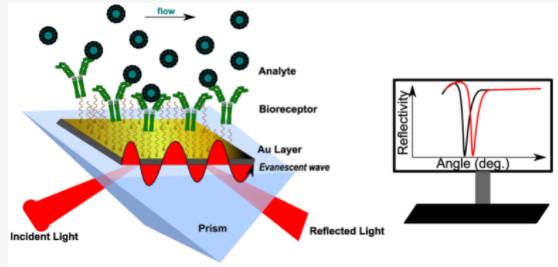 Figure 6: Schematic Principle of Surface Plasmon Resonance (SPR) Technique (Topor C-V, et al,. 2023)
Figure 6: Schematic Principle of Surface Plasmon Resonance (SPR) Technique (Topor C-V, et al,. 2023)Bioluminescence Interferometry (BLI) stands as another effective means of assessing the functional activity of antibody drugs. Compared to Surface Plasmon Resonance (SPR), BLI operates by real-time monitoring of changes in the optical thickness of sensors during molecular binding and dissociation processes, via fiber optic biosensors. As the biological membrane thickness changes, relative displacement occurs in the interference light waves, which are then detected by a spectrometer. This subsequent analysis of spectral changes before and after molecular binding facilitates precise evaluation of the target molecule. Consequently, BLI can also be employed to determine the affinity and kinetic properties of antibody drugs, contributing significantly to the characterization of their functional activity.
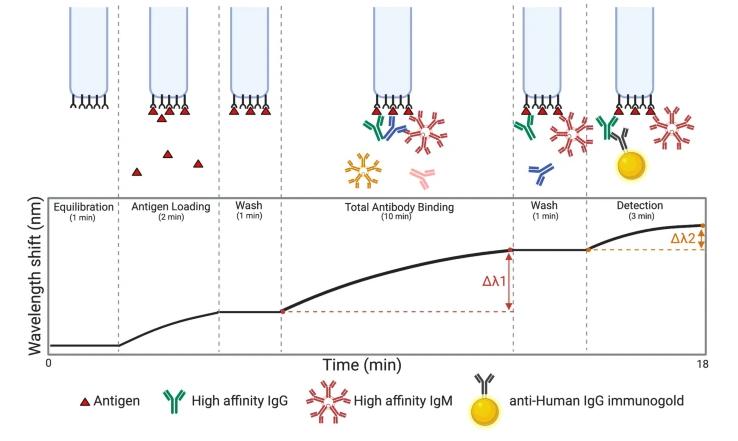 Figure 7: Overview of the BLI (Bio-Layer Interferometry) experiment. (John V. Dzimianski et al,. 2020)
Figure 7: Overview of the BLI (Bio-Layer Interferometry) experiment. (John V. Dzimianski et al,. 2020)Summary
As a category of biologics, antibody drugs demonstrate complex structural characteristics. The heterogeneity of these drugs could result from variations in the manufacturing process or storage conditions. Hence, to guarantee the safety and efficacy of antibody drugs, a systematic and comprehensive characterization analysis is crucial. This article aims to explore and delve into the main methods employed in the characterization analysis of antibody drugs. However, it should be recognized that with the continuous advancement of analytical technology, increasingly accurate and sensitive detection means will emerge, supplementing the existing analytical methods.
References
- Howard L. Levine, Brendan R. Cooney The development of therapeutic monoclonal antibody products, a comprehensive guide to CMC activities from clone to clinic.
- Allen G. Sequencing of proteins and peptides. Second ed. New York: Elsevier Science Publishing Company; c1989. Chapter 6, Determination of peptide sequences; p. 207-93. (Laboratory techniques in biochemistry and molecular biology; vol. 9).
- Dougherty J, et al. Using peptide maps as identity and purity tests for lot release testing of recombinant therapeutic proteins. BioPharm Int. 2003 Apr;16(4):54-8.
- van den Bremer ETJ, Heck AJR. Mass spectrometry in protein conformational analysis and molecular recognition. In: Jiskoot W, Crommelin D, editors. Methods for structural analysis of protein pharmaceuticals. Arlington (VA): American Association of Pharmaceutical Scientists; 2005. p. 435-64.
- Greenfield N. Using circular dichroism spectra to estimate protein secondary structure. Nat Protoc. 2006; 1(6): 2876-2890.
- Jefferis R. Glycosylation as a strategy to improve antibody-based therapeutics. Nat Rev Drug Discov. 2009 Mar;8(3):226-34.