Antibodies, integral to the human immune architecture, merited considerable scrutiny in the past century owing to the cardinal role of IgGs in the fight against infectious diseases. In the recently elapsed decade, antibody research has assumed a prominent role in advancing therapeutic interventions. Presently, beyond their contributions in the realm of infectious diseases, recombinant antibodies are quintessential in managing a host of pathological states such as cancer and rheumatoid arthritis.
For therapeutic and endogenous antibodies alike, one critical step towards their identification is the delineation of their constituent amino-acid sequence. At present, this objective is principally achieved by sequencing the B-cell receptor (BCR) repertoire at its nucleotide level. Yet, in the last few years, mass spectrometry (MS) has increasingly surfaced as an alternate and direct method for generating the much-sought-after protein-level sequence information.
Regardless of the establishment of a plethora of mass spectrometric techniques, the analysis of both recombinant and endogenous antibodies persists in confronting distinctive and formidable challenges. These demand an innovative approach that transcends the confines of conventional proteomics workflows. However, this exigency is far from insurmountable and offers promising avenues for novel explorations in the field of antibody research.
Select Service
The Role of Mass Spectrometry in Antibody Discovery
In the pursuit of therapeutic antibody discovery, mass spectrometry occupies a central role, circumnavigating the intricate complexities of antibody structural morphology and sequence heterogeneity. The standard tactic employed by researchers catalogs the systematic screening of B cells sourced from infected individuals. The process entails the isolating peripheral blood mononuclear cells (PBMCs), immortalizing them, and subsequent screening for antigen reactivity to uncover novel neutralizing antibodies. In recent scientific literature, evidence fortifies the effectiveness of such approach in the context of emerging infectious diseases, including Ebola and the Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2).
However, an emergent alternative deserves scrutiny: the direct discovery and characterisation of clinically active, functionally mature antibody clones at the protein level. In the last few years, momentous strides have been made in mass spectrometry-based proteomics, including improvements in sample preparation, mass spectrometry and liquid chromatography instrumentation, coupled with advances in data analysis methodologies. Thus, the leveraging of mass spectrometry techniques for protein level antibody sequencing bears enormous potential.
Currently, researchers engage three primary mass spectrometry-based sequencing strategies to discern antibody sequences (Figure 1). The first strategy, applicable to meticulously purified recombinant antibodies, leverages a variety of proteinases and elegant computational algorithms for comprehensive sequencing. The second strategy conjoins mass spectrometry techniques with genomics and transcriptomics to examine endogenous antibodies, with the sequence databases personalised via whole-genome sequencing or B-cell receptor sequencing; these form the basis for subsequent Bottom-Up Mass Spectrometry (BU-MS) data queries. Finally, the third strategy's versatility allows for the direct determination of comprehensive antibody sequences from selected clones in clinical samples, shunning the necessity for supplemental omics data.
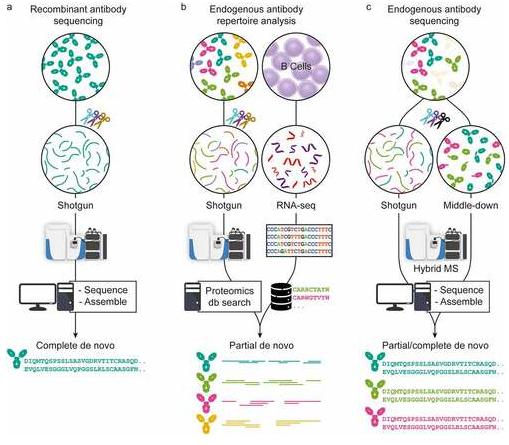 Figure 1 Three approaches in MS-based antibody sequencing
Figure 1 Three approaches in MS-based antibody sequencingMass Spectrometry-Based Monoclonal Antibody Analysis Methods
Proteomics, the large-scale study of proteins, employs numerous peptide and protein-focused mass spectrometry methods for proteomic analysis - certain methods have now been adapted for de novo sequencing of antibodies. Among these, the BU-MS method finds the broadest application in the area of mass spectrometry protein analysis. Enzymatic digestion and mass spectrometric fragmentation process follow specific rules that yield peptides and their fragment ions, producing overlapping peptides to accomplish complete sequence coverage, thereby revealing the antibody sequence as shown in Figure 2. To obtain complete and accurate antibody sequences, it typically requires a collaborative action of sequence-specific proteases and complementary mass spectrometric fragmentation techniques, with a homologous database-assisted search. However, it is worth noting that even in the most extensive databases, there may be no record exactly matching the target sequence. Therefore, the homologous-assisted method needs to score the experimentally determined sequence based on sequence alignment or error-tolerant fragment matching algorithms that extract sub-sequences (i.e., sequence tags) using matched sequences from the homologous database. At present, multiple de novo antibody sequencing software options exist on the market, such as PEAKS AB, capitalizing on data generated by various enzymatic digestions and multiple fragmentation methods, as well as the benefits of homologous antibody sequence databases for a comprehensive de novo sequence prediction of protein samples.
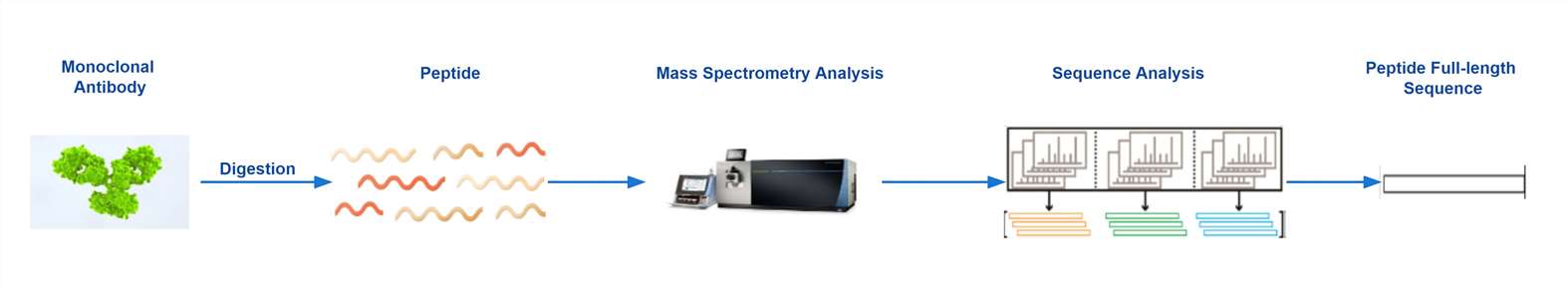 Figure 2 Monoclonal Antibody De novo Sequencing Workflow
Figure 2 Monoclonal Antibody De novo Sequencing WorkflowMass Spectrometry-based Sequencing of Polyclonal Antibodies
The endeavor of sequencing the complete repertoire of serum antibodies poses significant difficulties. One of the major obstacles lies in the restricted availability of samples. Polyclonal antibody samples typically stem from clinical specimens, and therefore, are available only in limited quantities. Moreover, the presence of monoclonal antibodies in plasma hover around a concentration of 1 μg/mL, making the isolation of singular clones excessively demanding, contributing to the complexity of the sequencing process. The majority of software tools are primarily designed for the assembly of individual antibodies, posing limitations for further analysis in instances where there is a presence of multiple similar IgG sequences in the data.
An additional challenge emerges when dealing with the intricate mixtures of endogenous polyclonal antibodies. Sequence signals emanating from highly variable regions often go unnoticed due to dilution effects. Contrarily, sequence signals originating from conservative regions are amplified, due to their ubiquitous presence in every clone, consequently suppressing the unique Complementary Determining Regions (CDRs) signal from each clone.
Notwithstanding these obstacles, progress in mass spectrometry methodologies and the emergence of sophisticated bioinformatics software have enabled the detection and analysis of individual clones from complex antibody mixtures using liquid chromatography-mass spectrometry (LC-MS) for whole proteins. The adoption of hybrid or multi-omics strategies can further bolster the analysis of polyclonal antibodies. For instance, Bondt et al. discerned each donor to possess between 50 to 500 persisting IgG1 Fab clones, and each donor's IgG1 gene library demonstrated a distinct simple repertoire of clones, with most clones numbering only a few hundred. Therefore, they attempted to use lineage templates closely matching light and heavy chains from the International ImMunoGeneTics database (IMGT), without employing antigen-specific capture. Through combining multiple protease digestions and electron-transfer dissociation (ETD) collected Bottom-Up Mass Spectrometry (BU-MS) data, they successively refined these templates, eventually yielding the final mature sequences. This strategy enabled the direct sequencing of the high abundance clones in patient blood, as depicted in Figure 3.
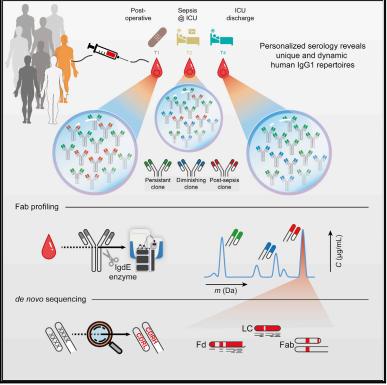 Figure 3 Mass spectrometry-based de novo sequencing of serum antibodies
Figure 3 Mass spectrometry-based de novo sequencing of serum antibodiesThe Development of Therapeutic Antibodies Unveiled through De Novo Sequencing
The field of mass spectrometry-based antibody sequencing has undergone significant advancements over the past decade, providing promising prospects for future development. While the methods for preparation of antibody samples have been evolving since the 1960s, the advent of Sanger sequencing for B cells in 1993 marked a critical milestone in the acquisition of valid sequence data. The rollout of next-generation sequencing technologies circa 2008 paved the way for high-throughput sequencing workflows, thus creating favourable conditions for the evolution of therapeutic antibodies.
Subsequently, the vigorous integration of mass spectrometry-based proteomic technologies has significantly bolstered the evolution of platforms designed for de novo antibody sequencing. Therefore, we have witnessed an extraordinary surge in the number of registered antibody sequences and therapeutics ( Figure 4).
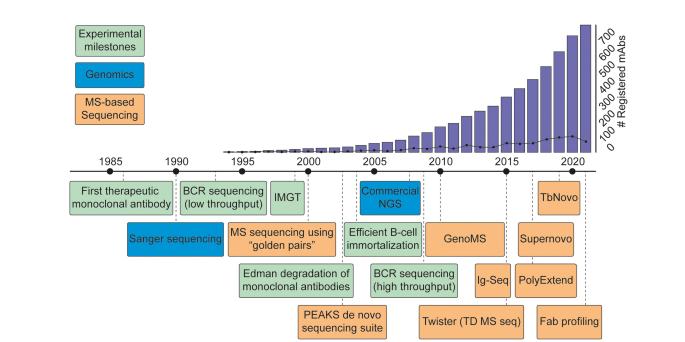 Figure 4 Timeline of key developments paving the way toward MS-based de novo sequencing of serum antibodies
Figure 4 Timeline of key developments paving the way toward MS-based de novo sequencing of serum antibodiesConclusion
This thorough discussion expounds on the influence and evolution of mass spectrometry technology in the sphere of antibody identification, paralleling the emergence of next-generation techniques in mass spectrometry-centric proteomics. The advent of high-fidelity mass spectrometers, coupled with unceasing enhancements in software infrastructure, is progressively mitigating the challenges intrinsic to mass spectrometry-rooted de novo antibody sequencing. The deployment of varied proteinases, multifaceted fragmentation methods, and sequencing approaches reinforced by analogous databases has led to the successful comprehensive sequencing of monoclonal antibodies and their recombinant counterparts. Despite these methodologies not yet being adapted for the direct analysis of native antibody amalgamations within serum, synergizing with alternate protein genomics technologies has enabled partial sequencing of multi-clonal antibodies. The relentless advances in mass spectrometry-oriented strategies hold the promise of continued progress in unraveling the complexities of antibody sequences, thus establishing a robust pathway for more proficient antibody discovery and production.
References
- de Graaf SC, Hoek M, Tamara S, Heck AJR. A perspective toward mass spectrometry-based de novo sequencing of endogenous antibodies. MAbs. 2022 Jan-Dec;14(1):2079449.
- Bondt A, Hoek M, Tamara S, de Graaf B, Peng W, Schulte D, van Rijswijck DMH, den Boer MA, Greisch JF, Varkila MRJ, Snijder J, Cremer OL, Bonten MJM, Heck AJR. Human plasma IgG1 repertoires are simple, unique, and dynamic. Cell Syst. 2021 Dec 15;12(12):1131-1143.