- Services
- FAQ
- Demo
- Case Study
- Related Services
- Support Documents
- Inquiry
Understanding and controlling host cell proteins (HCPs) is critical for ensuring the safety, stability, and efficacy of biologic drugs. These proteins, carried over from the expression system during cell lysis or secretion, are among the most common process-related impurities in biopharmaceutical production. Even at trace levels, residual HCPs can:
- Disrupt enzymatic modifications
- Compromise product stability
- Impair purification steps
- Trigger immune responses in patients
While regulatory agencies such as the FDA and EMA have defined acceptable HCP thresholds, low-abundance or unidentified HCPs may still present immunogenic risks. That's why proactive detection and elimination of HCPs is essential throughout the drug development process.
Advanced HCP Detection and Quantification at Creative Proteomics
Creative Proteomics offers sensitive, high-resolution HCP analysis powered by mass spectrometry (MS), in full compliance with ICH Q6B, PharmEu, and USP <1047> guidelines. Our services go beyond generic ELISA by combining broad detection capabilities with the flexibility to validate both generic and process-specific polyclonal antibody coverage.
Our HCP analysis solutions support:
- Quantitative HCP profiling
- Identification of individual HCPs
- Immunoassay validation (e.g., ELISA coverage studies)
- Process optimization based on HCP trends
- Monitoring of high-risk residual proteins
- Custom solutions tailored to your biologics pipeline
Technology Platform and Workflow
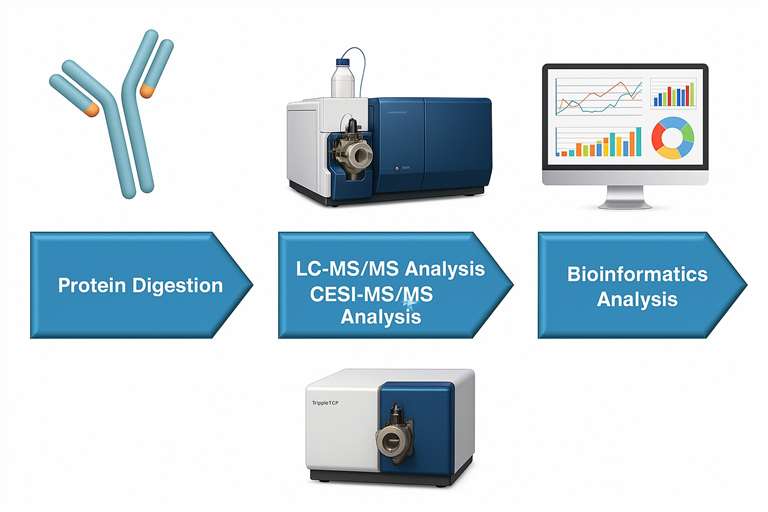
Creative Proteomics integrates several analytical methods—ELISA, 2D electrophoresis, Western blotting, and state-of-the-art mass spectrometry—to deliver accurate, comprehensive HCP characterization. For MS-based analysis, we employ:
- SWATH-MS on a SCIEX TripleTOF system for data-independent acquisition (DIA)
- Two-dimensional liquid chromatography (2D-LC) or capillary electrophoresis-electrospray ionization (CESI) for peptide separation
- CESI-MS, which improves sensitivity by reducing ion suppression and enhancing ionization, enabling the detection of low-abundance HCPs
Why Choose Us for Host Cell Protein Analysis?
- Ultra-sensitive detection: Identify trace-level HCPs with confidence
- Unbiased discovery: Detect known and unknown impurities
- Process optimization support: Pinpoint HCPs affecting downstream purification
- State-of-the-art instrumentation: CESI 8000 Plus, ExionLC™ AD, and TripleTOF 6600
- Rapid reporting
- Custom-tailored analysis: Protocols adapted to your sample type and workflow
- Expert Guidance Every Step of the Way
Our scientists are not only analytical experts—they're problem-solvers. Each project includes detailed technical documentation, tailored recommendations, and a collaborative approach to help you:
- Reduce immunogenicity risks
- Improve manufacturing consistency
- Accelerate regulatory submissions
Platform
CESI 8000 Plus, ExionLC™ AD, and TripleTOF 6600
Application Scenarios
- Impurity profiling
- Process development and downstream purification optimization
- Quality control during biosimilar development
- ELISA cross-validation and gap analysis
- Lot release and in-process monitoring
Deliverables
- Summary report with total HCP content
- Individual HCP identification table
- High-resolution graphs: HCP distribution, intensity heatmaps, purification stage comparisons
- ELISA coverage results (if applicable)
- Methodology and platform specifications
- Regulatory-compliant documentation
Host Cell Protein (HCP) Analysis – Sample Requirements
| Sample Type | Minimum Volume / Amount | Recommended Concentration | Storage & Shipping Conditions |
|---|---|---|---|
| Purified protein (e.g., mAb, fusion protein) | ≥ 100 µg | ≥ 1 mg/mL | -80°C (dry ice shipping) |
| Cell culture supernatant | ≥ 500 µL | N/A | -20°C or below (dry ice or ice packs) |
| Crude cell lysate | ≥ 200 µL | Protein content ≥ 1 mg/mL | -80°C (dry ice shipping) |
| Formulated biologic drug product | ≥ 50 µL or ≥ 100 µg | As supplied | Ship at 4°C or frozen depending on stability |
| ELISA antibody standard (if provided) | ≥ 100 µL |
Frequently Asked Questions (FAQ)
Q: What types of host cell lines can your HCP analysis support?
A: We support a wide range of cell lines, including CHO, HEK293, and E. coli. Custom support for other systems is available upon request.
Q: What is the detection limit for low-abundance HCPs?
A: Our CESI-MS/MS platform can detect HCPs at concentrations as low as sub-ng/mL, enabling reliable trace-level analysis.
Q: Can you validate our ELISA kit's coverage?
A: Yes. We offer ELISA coverage analysis to assess the binding efficiency of your polyclonal antibodies to process-specific HCPs.
Q: Can you help with regulatory submissions?
A: Absolutely. We provide detailed documentation in line with ICH Q6B and other regulatory guidelines to support your submission.
Q: Can you perform just the MS-based analysis if we already use ELISA?
A: Yes, our mass spectrometry platform can serve as a complementary or standalone tool to ELISA, especially when deeper profiling is needed.
Q: How is mass spectrometry improving HCP analysis?
A: Mass spectrometry allows for non-targeted, high-throughput identification and quantification of HCPs at sub-ppm levels, enabling better monitoring and control of impurities that affect drug stability and quality
Q: What are the challenges in HCP analysis?
A: Challenges include detecting low-abundance HCPs amidst high levels of therapeutic proteins, time-consuming sample preparation, lack of standardized assays for all HCPs, and interpreting the risk posed by individual HCPs.
Demo Sample
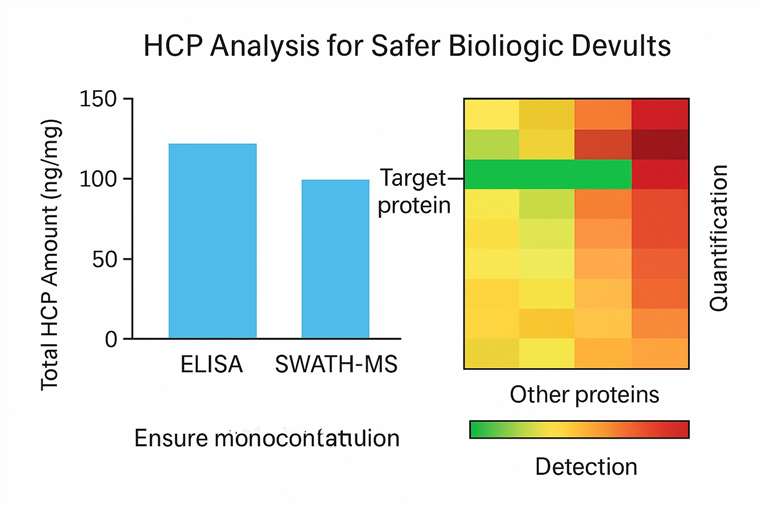
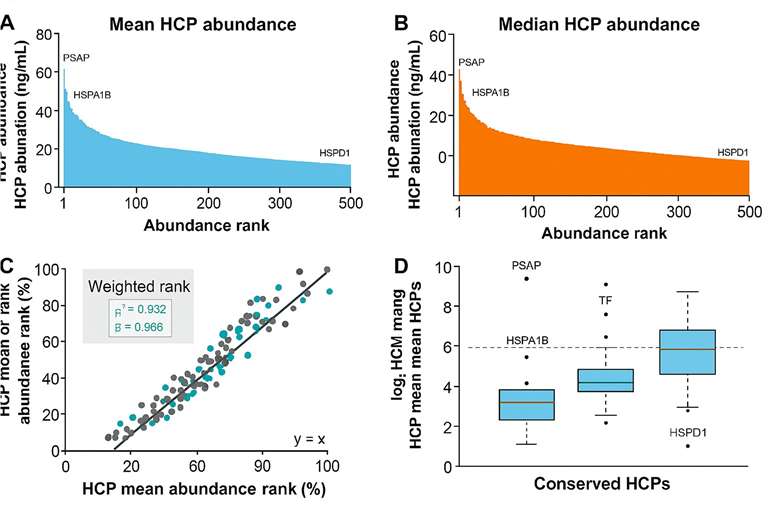
Case Study
SWATH-MS Quantification of HCPs in AAV Vector Purification
Background:
Adeno-associated virus (AAV) vectors are widely used in gene therapy. However, residual HCPs from the host cells can trigger unwanted immune responses. This study assessed HCP profiles across various AAV serotypes during downstream purification.
Technical Method:
Researchers used SWATH-MS (Sequential Window Acquisition of All Theoretical Fragment Ion Spectra) on a TripleTOF mass spectrometer to perform data-independent acquisition. The approach enabled reproducible and high-throughput quantification of HCPs across multiple AAV vector samples.
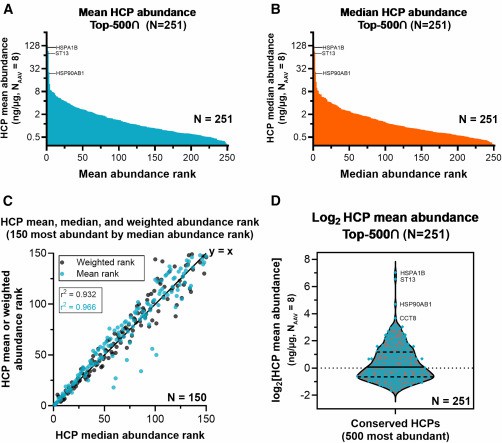 HCP abundance profiling by mean, median, and weighted ranking
HCP abundance profiling by mean, median, and weighted rankingConclusion:
The analysis revealed that high-abundance conserved species (HACS) dominated the residual HCP pool across all AAV serotypes. These results underscore the need to target conserved HCPs for better purification strategies and safer gene therapy products.
References
- Bailey-Kellogg C., Gutierrez A.H., Moise L., et al. CHOPPI: a web tool for the analysis of immunogenicity risk from host cell proteins in CHO-based protein production. Biotechnology and Bioengineering. Vol. 111, No. 11, November, 2014. DOI: 10.1002/bit.25286
- Tscheliessnig A.L., Konrath J., Bates R., et al. Host cell protein analysis in therapeutic protein bioprocessing-methods and application. Biotechnol. J. 2013, 8, 655–670. DOI: 10.1002/biot.201200018
- Valente K.N., Levy N.E., Lee K.H, et al. Applications of proteomic methods for CHO host cell protein characterization in biopharmaceutical manufacturing. Current Opinion in Biotechnology. 2018, 53:144-150. DOI: 10.1016/j.copbio.2018.01.004
Related Services
Support Documents
INFOGRAPHIC